Journal of Cell Science Using fly genetics to dissect the cytoskeletal machinery of neurons during axonal growth and maintenance Andreas Prokop*, Robin Beaven, Yue Qu and Natalia Sa ´ nchez-Soriano* Faculty of Life Sciences, The University of Manchester, Michael Smith Building, Oxford Road, Manchester M13 9PT, UK *Authors for correspondence ([email protected]; [email protected]) Journal of Cell Science 126, 2331–2341 ß 2013. Published by The Company of Biologists Ltd doi: 10.1242/jcs.126912 Summary The extension of long slender axons is a key process of neuronal circuit formation, both during brain development and regeneration. For this, growth cones at the tips of axons are guided towards their correct target cells by signals. Growth cone behaviour downstream of these signals is implemented by their actin and microtubule cytoskeleton. In the first part of this Commentary, we discuss the fundamental roles of the cytoskeleton during axon growth. We present the various classes of actin- and microtubule-binding proteins that regulate the cytoskeleton, and highlight the important gaps in our understanding of how these proteins functionally integrate into the complex machinery that implements growth cone behaviour. Deciphering such machinery requires multidisciplinary approaches, including genetics and the use of simple model organisms. In the second part of this Commentary, we discuss how the application of combinatorial genetics in the versatile genetic model organism Drosophila melanogaster has started to contribute to the understanding of actin and microtubule regulation during axon growth. Using the example of dystonin-linked neuron degeneration, we explain how knowledge acquired by studying axonal growth in flies can also deliver new understanding in other aspects of neuron biology, such as axon maintenance in higher animals and humans. Key words: Growth cone, Cytoskeleton, Axon, Drosophila, Actin, Microtubule, Brain disorders Introduction The cytoskeleton consists of actin, intermediate filaments (IFs) and microtubules (MTs), and has fundamental roles in virtually every function of cells, including cell division, motility, adhesion, signalling, endocytic trafficking and transport, as well as in the regulation of cell and organelle shapes and their distributions (Akhmanova and Stearns, 2013). Naturally, this also applies to the nervous system where the cytoskeleton has essential roles during the development, function, regeneration and degeneration of neurons. Of the more than 200 entries on the OMIM (Online Mendelian Inheritance in Man) database for genes encoding cytoskeleton-associated proteins, 44% have indexed links to human disorders, and 54% of these are linked to nervous system disorders, including neurodevelopmental disorders (e.g. lissencephalies, mental retardations), functional disorders (e.g. deafness) and a wide range of degenerative diseases (see supplementary material Table S1). Therefore, research of the cytoskeleton provides unique opportunities to unravel fundamental mechanisms of the nervous system, both in health and disease. It is currently little understood how cytoskeleton-associated proteins contribute to cellular mechanisms of neurons. For example, the systemic importance of certain MT-binding proteins, such as motor proteins and the structural protein tau, has long been recognised from their involvement in a broad spectrum of neurodegenerative diseases, and we know their principal molecular functions. However, we have yet to explain conclusively how they function within normal or diseased neurons (Hirokawa et al., 2010; Morris et al., 2011). Here, we argue that this is best performed starting with processes for which there is already an extensive body of knowledge about the cytoskeleton, in particular the growth of axons. Axons are slender neuronal extensions, which can be several metres long, and are often arranged into nerves or nerve tracts that provide essential ‘information highways’ in the nervous system. The proper function of nervous systems requires axons to grow and wire up correctly during development or regeneration, and these connections need to be maintained for decades in the ageing body. Here, we review the current concepts for the cellular roles that actin and MTs have in axons and discuss important gaps in our understanding of how the cytoskeleton is regulated to this end. We argue that genetics provides important means to advance this understanding, and explain how studies using the genetic model organism Drosophila melanogaster can make important contributions. We review recent advances with regard to actin and MT regulation during axonal growth in the fly and provide an example of how mechanistic understanding in one context can be used to identify mechanisms underpinning other processes, such as axon maintenance and neurodegeneration. The organisation of the cytoskeleton in axons Mature axons are the longest cellular entities that are produced by animals. They propagate electrical messages, which travel from the neuronal cell bodies towards the synaptic connections with their distant target cells, often several metres away. To maintain This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/3.0), which permits unrestricted use, distribution and reproduction in any medium provided that the original work is properly attributed. Commentary 2331

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
Journ
alof
Cell
Scie
nce
Using fly genetics to dissect the cytoskeletalmachinery of neurons during axonal growth andmaintenance
Andreas Prokop*, Robin Beaven, Yue Qu and Natalia Sanchez-Soriano*Faculty of Life Sciences, The University of Manchester, Michael Smith Building, Oxford Road, Manchester M13 9PT, UK
*Authors for correspondence ([email protected]; [email protected])
Journal of Cell Science 126, 2331–2341� 2013. Published by The Company of Biologists Ltddoi: 10.1242/jcs.126912
SummaryThe extension of long slender axons is a key process of neuronal circuit formation, both during brain development and regeneration. Forthis, growth cones at the tips of axons are guided towards their correct target cells by signals. Growth cone behaviour downstream of
these signals is implemented by their actin and microtubule cytoskeleton. In the first part of this Commentary, we discuss thefundamental roles of the cytoskeleton during axon growth. We present the various classes of actin- and microtubule-binding proteins thatregulate the cytoskeleton, and highlight the important gaps in our understanding of how these proteins functionally integrate into the
complex machinery that implements growth cone behaviour. Deciphering such machinery requires multidisciplinary approaches,including genetics and the use of simple model organisms. In the second part of this Commentary, we discuss how the application ofcombinatorial genetics in the versatile genetic model organism Drosophila melanogaster has started to contribute to the understanding
of actin and microtubule regulation during axon growth. Using the example of dystonin-linked neuron degeneration, we explain howknowledge acquired by studying axonal growth in flies can also deliver new understanding in other aspects of neuron biology, such asaxon maintenance in higher animals and humans.
Key words: Growth cone, Cytoskeleton, Axon, Drosophila, Actin, Microtubule, Brain disorders
IntroductionThe cytoskeleton consists of actin, intermediate filaments (IFs) and
microtubules (MTs), and has fundamental roles in virtually every
function of cells, including cell division, motility, adhesion, signalling,
endocytic trafficking and transport, as well as in the regulation of cell
and organelle shapes and their distributions (Akhmanova and Stearns,
2013). Naturally, this also applies to the nervous system where the
cytoskeleton has essential roles during the development, function,
regeneration and degeneration of neurons. Of the more than 200 entries
on the OMIM (Online Mendelian Inheritance in Man) database for
genes encoding cytoskeleton-associated proteins, 44% have indexed
links to human disorders, and 54% of these are linked to nervous
system disorders, including neurodevelopmental disorders (e.g.
lissencephalies, mental retardations), functional disorders (e.g.
deafness) and a wide range of degenerative diseases (see
supplementary material Table S1). Therefore, research of the
cytoskeleton provides unique opportunities to unravel fundamental
mechanisms of the nervous system, both in health and disease.
It is currently little understood how cytoskeleton-associated
proteins contribute to cellular mechanisms of neurons. For
example, the systemic importance of certain MT-binding
proteins, such as motor proteins and the structural protein tau,
has long been recognised from their involvement in a broad
spectrum of neurodegenerative diseases, and we know their
principal molecular functions. However, we have yet to explain
conclusively how they function within normal or diseased
neurons (Hirokawa et al., 2010; Morris et al., 2011). Here, we
argue that this is best performed starting with processes for which
there is already an extensive body of knowledge about the
cytoskeleton, in particular the growth of axons.
Axons are slender neuronal extensions, which can be several
metres long, and are often arranged into nerves or nerve tracts
that provide essential ‘information highways’ in the nervous
system. The proper function of nervous systems requires axons to
grow and wire up correctly during development or regeneration,
and these connections need to be maintained for decades in the
ageing body. Here, we review the current concepts for the
cellular roles that actin and MTs have in axons and discuss
important gaps in our understanding of how the cytoskeleton is
regulated to this end. We argue that genetics provides important
means to advance this understanding, and explain how studies
using the genetic model organism Drosophila melanogaster can
make important contributions. We review recent advances with
regard to actin and MT regulation during axonal growth in the fly
and provide an example of how mechanistic understanding in one
context can be used to identify mechanisms underpinning other
processes, such as axon maintenance and neurodegeneration.
The organisation of the cytoskeleton in axonsMature axons are the longest cellular entities that are produced by
animals. They propagate electrical messages, which travel from
the neuronal cell bodies towards the synaptic connections with
their distant target cells, often several metres away. To maintain
This is an Open Access article distributed under the terms of the Creative Commons AttributionLicense (http://creativecommons.org/licenses/by/3.0), which permits unrestricted use, distributionand reproduction in any medium provided that the original work is properly attributed.
Commentary 2331
Journ
alof
Cell
Scie
nce
this unusual architecture, shafts of axons contain prominent
bundles of parallel MTs, which provide structural support as well
as the tracks for the vital long-distance cargo transport
(Twelvetrees et al., 2012). Axonal MTs are discontinuous and
are usually not attached to the centrosome (the main microtubule
organizing centre of most cells); their plus ends mostly, but not
entirely, point distally (Fig. 1A) (Ahmad et al., 1999; Baas and
Lin, 2011; Basto et al., 2006; Stiess et al., 2010). The cortex of
axon shafts is lined by short stabilised actin filaments (Fig. 1A),
which are organised into repetitive rings by the scaffolding
protein spectrin and linked to the axonal cell membrane through
ankyrins (Xu et al., 2013). Axons also contain a number of
different IF proteins that can regulate the specific diameters of
different axon classes (Perrot and Eyer, 2009).
Axons are usually maintained for the lifetime of an organism,
i.e. for decades in humans. It is therefore not surprising that axonal
decay is a characteristic feature in the ageing brain (Marner et al.,
2003). Sites of severe disorganisation of axonal MT bundles
(referred to as diverticula) are considered an important trigger of
such axon loss (Adalbert et al., 2009; Fiala et al., 2007). Axonal
MT bundles are believed to be stabilised through structural MT-
binding proteins (MTBPs), such as tau or MAP1B (Chilton and
Gordon-Weeks, 2007). At the same time, there seems to be a
steady and dynamic MT turnover, as suggested by continued MT
polymerisation events in mature axons (Kollins et al., 2009). Such
dynamics might allow for structural plasticity, such as shortening
or extension of the axon shaft (Kuppers-Munther et al., 2004;
Lamoureux et al., 2010; Rajagopalan et al., 2010), pruning and
regrowth of axons or formation of new side branches (Letourneau,
2009), which are required for rewiring processes during learning
and memory formation or the regeneration of damaged axons
(Bradke et al., 2012; Saxena and Caroni, 2007).
Cellular roles of the cytoskeleton during axonaldevelopmentDuring development, axons are initiated as small neuritic
protrusions on the surface of neuronal cell bodies. For this,
MTs in the neuronal cell bodies become stabilised and push
beyond the cell cortex (Brandt, 1998). This is assisted by
dynamic rearrangements of the cortical actin cytoskeleton to
remove barrier function (Flynn et al., 2012; Neukirchen and
Bradke, 2011; Saengsawang et al., 2012) and the formation of
filopodia, which provide membrane protrusions into which MTs
can extend (Dent et al., 2007; Goncalves-Pimentel et al., 2011).
In typical vertebrate neurons, several neurites are formed
simultaneously on the cell body, but only one of these is
selected as the axon. This selection process (referred to as neuron
polarisation) involves signalling cascades leading to the targeted
Filopodia
Engorgingfilopodium
Engorging filopodium
Cortical actinCortical actin rings
F-actinlattice
F-actinbundle
F-actinarc
LamellipodiumSoma
Axon
CentrosomeMT polymerisation
F-actin backflowRepulsive cue
MT-destabi-lising drug
MT-stabilising drugElectric field MTF-actin
latticeF-actinbundle
StabilisedMT
Attractive cue
oror
or or
Growthcone
Nucleus
A
B
E F-act– F F-act– C Splayed MT– D F-act–
New axon segment
Key
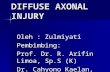
Fig. 1. Principal organisation and roles of actin and
MTs in growing axons. (A) The figure shows the
principal distribution and organisation of MTs and F-
actin networks in the soma, axon and growth cone of a
neuron that is exposed to an attractive guidance cue.
The tip of the axonal MT bundle forms the central zone
of the growth cone from where splayed MTs emanate
into the actin-rich peripheral zone (Dent et al., 2011).
(B) Directed axon extension (growth cone turning).
Splayed MTs become preferentially stabilised in a
certain direction through guidance cues [or
experimentally through unilateral exposure to
MT-(de-)stabilising drugs or the application of an
electrical field]. Additional MTs follow in that
direction, leading to engorgement of membrane
structures in that area and, eventually, the establishment
of a new axon segment (Buck and Zheng, 2002;
Lowery and Van Vactor, 2009; McCaig et al., 2002;
Sabry et al., 1991; Tanaka and Kirschner, 1995;
Tanaka and Kirschner, 1991). (C) Growth cone turning
requires splayed MTs: low doses of MT-stabilising
drugs suppress the occurrence of splayed, side-stepping
MTs (splayed MT2). Growth cones that are treated this
way fail to show normal turning behaviour at repulsive
borders and stall instead (Challacombe et al., 1997;
Williamson et al., 1996). (D,E) Growth cone turning
requires F-actin. When F-actin is destabilised through
drugs (F-act2), MTs tend to be bundled and growth
cones fail to turn at repulsive borders (Challacombe et
al., 1996); they also no longer respond to unidirectional
application of MT-stabilising drugs (Buck and Zheng,
2002). (F) Growth cone turning towards the cathode in
electrical fields (galvanotropism) does not require F-
actin (McCaig, 1989); this turning could be mediated
by MTBPs, which are attracted to the cathode through
positive net-charge of their MT-binding domains, thus
dragging the MTs along with them.
Journal of Cell Science 126 (11)2332
Journ
alof
Cell
Scie
nce
localisation of cell polarity factors (Barnes and Polleux, 2009)and also depends on the specific stabilisation of MTs in the
selected neurite (Brandt, 1998; Chen et al., 2013).
Once established, axons grow along stereotypic paths to theirtarget cells, a process that is implemented by prominent hand-shaped growth cones at the axon tips. Growth cones are guided
towards their specific target areas and cells through precisespatio-temporal patterns of chemical signals that are arrangedalong their paths (Tessier-Lavigne and Goodman, 1996). Many
guidance cues and their receptors have been identified and shownto mediate repulsion or attraction (Araujo and Tear, 2003; Huberet al., 2003), thus controlling growth cone behaviours, such as
growth velocity, pausing, turning, retraction or collapse. Thesemorphogenetic movements downstream of guidance signallingare implemented by the prominent actin and MT cytoskeleton ofgrowth cones (Fan et al., 1993), which is arranged in a particular
spatial geometry.
The geometry of growth cones can be subdivided into an actin-rich peripheral zone and a MT-rich central zone (Fig. 1A), and
the area in which MTs and F-actin prominently overlap is oftenreferred to as the transition zone. The central zone of growthcones comprises the tips of the axonal MT bundles, and from here
single MTs splay out into the peripheral zone. The peripheralzone consists of actin-rich membrane protrusions, which includefinger-like filopodia (containing parallel F-actin bundles) andveil-like lamellipodia (containing F-actin lattices; Fig. 1A).
Filopodia and lamellipodia sense guidance cues by presentingtheir receptors on their surfaces. They undergo constant shapechanges, driven by continuous assembly of actin filaments at the
leading edges, the retrograde flow of these filaments, and theirdepolymerisation and recycling further back in the transitionzone (Box 1). This actin dynamics allow filopodia to reach out
into the growth cone environment and act as signalling andadhesion sensors (Chien et al., 1993; Davenport et al., 1993;Dwivedy et al., 2007; Mattila and Lappalainen, 2008). They
allow lamellipodia to act as dynamic sites of substrate adhesionand myosin-II-driven contraction and force generation, which areimportant in influencing MT behaviours during guided axonextension (Box 2).
In current models of growth cone guidance (Fig. 1B) (Dentet al., 2011; Lowery and Van Vactor, 2009), external signalstrigger the directional stabilisation of single MTs in the
peripheral zone. These stabilised MTs are then joined by theextension of the bulk of MTs from the central domain (a processreferred to as engorgement), thus giving rise to a new axon
segment. This elongation step is completed when lamellipodiaand filopodia relocate distally to the newly formed axon tip(Fig. 1B). In this model, MTs are the essential implementers ofaxon extension, whereas F-actin is required for the directionality
of this extension (through influencing MT behaviours; Box 2).Accordingly, blocking MT dynamics inhibits axon growth(Letourneau and Ressler, 1984; Sanchez-Soriano et al., 2010;
Tanaka et al., 1995), whereas the pharmacological or geneticinhibition of actin polymerisation, either in cultured neurons or in
vivo, does not usually suppress axonal growth per se, but
abrogates growth cone turning and pathfinding (Fig. 1D,E) (Buckand Zheng, 2002; Challacombe et al., 1996; Chien et al., 1993;Kaufmann et al., 1998; Marsh and Letourneau, 1984; Sanchez-
Soriano et al., 2010). This said, a directional movement of growthcones does not always require F-actin (Fig. 1F), as somesignalling mechanisms can target MTs directly. For example,
GSK3-mediated phosphorylation of the MTBPs APC1, MAP1B
and/or CLASP is known to directly regulate MT stability in
growth cones (Hur et al., 2011; Lucas et al., 1998; Purro et al.,
2008).
Box 1. ABPs and MTBPs as essential regulators ofcytoskeletal dynamics
MTs are stiff hollow tubes typically formed by 13 protofilaments
(filamentous polymers of a- and b-tubulin heterodimers) that grow
and shrink primarily at their plus ends. Actin filaments are more
flexible double-helical chains of actin monomers that maintain their
equilibrium mainly by growth at plus ends and shrinkage at minus
ends (Fletcher and Mullins, 2010; Hawkins et al., 2010; Stricker
et al., 2010). Both have in common that they are polymers of
repetitive building blocks (black helmet-like symbols in the figure)
that are assembled in a polar fashion. Their dynamics are
regulated in comparable ways through the following different
classes of actin-binding proteins (ABPs) and MT-binding proteins
(MTBPs) as illustrated in the box figure.
(1) New actin filaments or MTs are generated through nucleation
factors that catalyse the transition from mono- or oligomers to
polymers; (2) plus-end dynamics, such as (de-)polymerisation,
stabilisation, directionality and targeting is regulated by plus-end-
binding proteins; (3) nucleation or (de-)polymerisation processes
are further regulated by proteins that bind actin or tubulin
monomers or oligomers and determine their availability, for
example, SCAR or N-WASP activates Arp2/3 nucleation, APC1
cooperates with mDia1 in nucleation, stathmin sequesters tubulin,
profilin enhances actin polymerisation by ENAH (Bear and Gertler,
2009; Okada et al., 2010; Pollitt and Insall, 2009; Steinmetz,
2007); (4) proteins that bind along actin filaments or MTs can
stabilise them against depolymerisation, cross-link them into
bundles and/or networks, and/or link them to other cytoskeletal
components, organelles or the cortex; (5) minus-end-binding
proteins regulate stability versus (de-)polymerisation; (6,7) plus-
and minus-end-directed motor proteins mediate filament
contraction, the sliding along other cellular structures or
filaments, or the transport of cargo (C in the figure) along F-actin
or MTs; (8) different classes of proteins can sever or actively
depolymerise F-actin or MTs, for example, cofilin depolymerises or
severs pointed ends of F-actin, type 13 kinesins depolymerise MTs
from the plus end, and katanin or spastin sever MT shafts (Pak
et al., 2008; Conde and Caceres, 2009); (9) post-translational
modifications (PTM) influence F-actin or MT stability and the
interaction with certain ABPs and MTBPs (Fukushima et al., 2009;
Janke and Bulinski, 2011; Terman and Kashina, 2013).
91
2
6
7
85
4
3
33
C
CPTM
Nucleation
G-actin orα-tubulin–β-tubulin
Polymer/network regulation
Key
Cytoskeletal machinery of neurons 2333
Journ
alof
Cell
Scie
nce
The importance and challenge of understandingcytoskeletal machineryClearly, the cytoskeleton has essential roles during axonal
growth, but we still do not understand how it is regulated to
perform these functions. Different classes of actin-binding
proteins (ABPs) and MTBPs are known to regulate cytoskeletal
dynamics (Box 2); many of these are expressed in neurons
(Table 1), and we often have a reasonable understanding of their
molecular mechanisms, at least in vitro. However, we do not
understand how these individual regulators functionally integrate
into the common cytoskeletal machinery that ultimately
implements neuronal behaviours. This deficit also leaves us
with little means to explain the cellular aberrations caused
by mutations of genes encoding ABPs or MTBPs that have
been linked to brain disorders, including disorders of
neurodevelopment and axonal growth (supplementary material
Table S1) (Nugent et al., 2012; Tischfield et al., 2011).
The need to understand cytoskeletal regulation at the cellular
level in neurons and non-neuronal cells alike is a current problem(Fletcher and Mullins, 2010; Insall and Machesky, 2009). This isclearly reflected by the number of review articles that attempt to
propose models explaining the functional integration ofcytoskeletal regulators (e.g. Conde and Caceres, 2009; Dentet al., 2011; Gallo, 2013; Lowery and Van Vactor, 2009; Paket al., 2008). For example, there are mechanistic models aiming
to explain how different classes of ABPs functionally combineduring filopodia formation (Mattila and Lappalainen, 2008) andmodels based on end-binding (EB) proteins that propose how an
increasing number of MT-plus-end-associating proteins canregulate MT dynamics (Akhmanova and Steinmetz, 2008;Akhmanova and Steinmetz, 2010) (see also Fig. 2). Currently,
such models depend on knowledge gained from a broad rangeof different neuronal systems, and they therefore remainhypothetical and are difficult to test experimentally.
In this Commentary, we promote strategic considerations that
will help to overcome some of the current shortcomings inexplaining cytoskeletal machinery. We believe that strongeremphasis must be given to studying the various parts of
cytoskeletal machinery consistently in the same neuronsystems, so that their functional links and interfaces can bedirectly assessed. These neuronal systems should not only be
amenable to biochemical, biophysical and cell biologicalapproaches, but also to the use of combinatorial genetics,therefore allowing a systematic assessment of different genemanipulations or combinations of mutations in the same cells.
Such a strategy will help us to integrate the knowledge we haveabout single cytoskeletal regulators so that we can develophigher-order concepts about the governance of the cytoskeletal
machinery during axon growth or in the context of braindisorders.
Drosophila as a model to study cytoskeletal machineryduring axonal growthOne organism in which combinatorial genetics has been andcontinues to be most successfully applied to the study of complex
biological problems is the fruit fly Drosophila melanogaster
(Keller, 1996; Roote and Prokop, 2013). Work in Drosophila islow cost, impressively fast, and particularly amenable to geneticmanipulations and unbiased genetic screens (Sanchez-Soriano
et al., 2007). Accordingly, Drosophila research has been anincubator for new ideas and concepts in neurobiology, and workin the fly revealed many fundamental molecular and cellular
mechanisms that have turned out to be conserved in higheranimals (Bellen et al., 2010). Work in Drosophila has madeimportant contributions to the understanding of axon growth,
including the discovery and analysis of proteins involved inpathfinding, and the discovery of mechanisms underpinningpioneer guidance, nerve branching, target cell recognition and
axonal transport (Araujo and Tear, 2003; Astigarraga et al., 2010;Brochtrup and Hummel, 2011; Landgraf and Thor, 2006;Sanchez-Soriano and Prokop, 2005; Sanchez-Soriano et al.,2007; Zala et al., 2013).
To study the cytoskeletal machinery during axon growth, flyneurons offer a number of useful features. First, most of theDrosophila ABPs and MTBPs display a high level of sequence
conservation with their mammalian counterparts, and they areusually expressed in the nervous system (Table 1) (Sanchez-Soriano et al., 2007). Second, cytoskeletal regulators tend to
Box 2. How F-actin networks influence MTdynamics in growth cones
N F-actin networks generate membrane protrusions on all sides
of growth cones that can be invaded by MTs in any direction.
This facilitates lateral extension and stabilisation of MTs as an
important prerequisite for growth cone turning (see also
Fig. 1B).
N F-actin networks can influence MT behaviours mechanically
(Lee and Suter, 2008; Schaefer et al., 2002) (see also Fig. 1A).
Thus, MT extension is antagonised by F-actin backflow or
through the formation of transverse bundles (arcs) that form
‘road-blocks’. MT extension can be promoted and guided
through the formation of radial F-actin bundles as tracks for MT
elongation. MTs can be pushed laterally or medially through
actomyosin contractions. Finally, MT extension can be
channelled into certain directions through coordinated loss of
F-actin from local spots within lamellipodia.
N Actin networks form scaffolds that serve as a platform for ABPs.
Some of these ABPs influence MT behaviours through mediating
actin–MT linkage. This linkage can be direct through ABPs that
also contain MT-binding domains [e.g. MAP1B, spectraplakins,
coronin 7 (POD1 in Drosophila), cytoplasmic-linker-associated
proteins (CLASPs) or adenomatous polyposis coli (APC)
(Bouquet et al., 2007; Moseley et al., 2007; Rothenberg et al.,
2003; Sanchez-Soriano et al., 2009; Tsvetkov et al., 2007)]. Other
ABPs are involved in indirect crosstalk with MTs by interacting
with MTBPs. Reported examples are the ABP drebrin that is
linked to the MTBP EB3, the ABP PPP1R9A (better known as
spinophilin) that binds to DCX, IQGAP that interacts with CLIP-
170, or functional interactions between the actin-binding motor
protein myosin II and the MT-associated dynein motor protein
complex (Bielas et al., 2007; Geraldo et al., 2008; Myers et al.,
2006; Swiech et al., 2011).
N Some ABPs can tether actin networks to transmembrane
receptors when they are engaged with their extracellular
ligands. This has been demonstrated for different classes of
transmembrane receptors (Giannone et al., 2009; Moore et al.,
2009; Moore et al., 2010). Such linkage of F-actin to the
extracellular environment generates mechanical forces across
the membrane, which can trigger intracellular signalling events
(referred to as the ‘clutch’ mechanism) that then can influence
actin and MT dynamics (Suter and Forscher, 2000; Suter and
Miller, 2011).
Journal of Cell Science 126 (11)2334
Journ
alof
Cell
Scie
nce
display minimal redundancy in the fly genome. For example,
eliminating the function of Ena/VASP requires a triple knockout
in mouse, whereas only a single gene needs to be removed in
Drosophila (Goncalves-Pimentel et al., 2011; Kwiatkowski et al.,
2007). Third, mutant alleles or other genetic tools are often
readily available (McQuilton et al., 2012), and they can usually
be combined in the same animal in a short time (Goncalves-
Pimentel et al., 2011; Koper et al., 2012). Finally,
experimentation is fast; axon extension can be studied in fly
embryos after only half a day of development, and in primary
neurons after only 6 hours in culture (Prokop et al., 2012;
Sanchez-Soriano et al., 2007).
To study cytoskeletal regulation during axonal growth, genetic
manipulations of cytoskeletal regulators can be combined with a
broad spectrum of morphological readouts for axon growth in
vivo (Sanchez-Soriano et al., 2007), including live imaging of
cytoskeletal dynamics (Mattie et al., 2010; Pawson et al., 2008).
This repertoire can be further expanded when making
complementary studies in Drosophila primary neurons, which
give access to quantitative measurements of filopodia and
lamellipodia, retrograde F-actin flow, the organisation,
stabilisation and polymerisation rates of MTs, as well as axonal
growth rates (Prokop et al., 2012; Sanchez-Soriano et al., 2010).
Such readouts are crucial to unravel how ABPs and MTBPs
regulate the cytoskeleton in cells. Notably, cytoskeletal dynamics
in fly neurons are similar to those of other animal neuron
systems, indicating that the underlying regulation is fundamental
and well preserved (Sanchez-Soriano et al., 2010). This said,
there will be limitations when using neurons of this little insect
because some aspects are different in vertebrate neurons;
for example, the axon length and life time, the absence
of neurofilaments and myelination, and their unipolar
organisation, including the spike-initiating zone, which is
located at the base of dendrites away from the soma and not
near the cell body as is the axon initial segment in vertebrates
(Grubb and Burrone, 2010; Sanchez-Soriano et al., 2005). In the
following sections, we will provide examples of work on axonal
growth in the fly that has contributed new understanding of
cytoskeletal machinery.
Use of the fly to unravel new ABP functionsduring axonal growthWithin a few years, work in Drosophila growth cones has
identified at least four new ABPs as actin regulators in neurons,including Shot, DAAM, Psidin and Canoe. Shot is the only
Drosophila member of the spectraplakin family of actin- and
MT-binding proteins with two close homologues in mammals,
MACF1 (also known as ACF7) and dystonin (DST, also known
as BPAG1) (Suozzi et al., 2012). Both Shot and ACF7 promote
filopodia formation (Sanchez-Soriano et al., 2009). In Shot, this
function is relevant for axon guidance at the CNS midline, and it
requires binding of the two EF-hand domains to the putative
translational regulator Extra bases (Exba, also known as eIF5C)
(Fig. 2A) (Lee et al., 2007; Sanchez-Soriano et al., 2009), thus
potentially contributing to local translation events in growth
cones (Van Horck and Holt, 2008). DAAM (DAAM1 in
mammals) is one of four Drosophila formins (Diaphanous,
DAAM, Fhos and CG32138) reported to be present in the
nervous system (Pawson et al., 2008; Prokop et al., 2011;
Sanchez-Soriano et al., 2007). DAAM has now been shown to be
the essential formin in growth cones of embryonic neurons,
where it promotes filopodia formation and axonal pathfinding in
Table 1. Examples of evolutionary conserved MTBPs and ABPs in human and fly
Cytoskeleton regulator class MTBPs ABPs
Nucleators c-tubulin ring complex (cTub23C and other components);doublecortin* (DCX-EMAP)
Arp2/3 complex (Arpc1, Arp3, etc.); formins(DAAM, Fhos, CG32138, dia)
Plus-end-binding proteins MAPFRE/EB* (eb1); MT-plus-end-tracking proteins (+TIPs);doublecortin* (DCX-EMAP); CKAP5/CHTOG/XMAP215*(msps)
ENAH/VASP* (ena); formins (DAAM, Fhos,CG32138, dia); capping proteins* (cpa, cpb);adducins* (hts)
Mono- or oligomer-binding proteins Stathmin/SCG10* (stai); DPYSL2/CRMP2* (CRMP); CKAP5/CHTOG/XMAP215* (msps); doublecortin* (DCX-EMAP);tub-specific chaperone E* (tbce); CLASP* (chb)
WASF/SCAR/WAVE* (SCAR); WASL/N-WASP*(WASp); APC* (Apc); b4-thymosin/TMSB4X*(cib)a; profilin* (chic); twinfilin* (twf)
Shaft stabilisers and cross-linkers MAPT* (tau); MAP2* (tau); MAP1B* (futsch); MACF1*;dystonin* (shot)
Fascin* (sn); a-actinin* (Actn); plastin/fimbrin*(Fim); filamin* (cher); tropomyosin* (Tm1);moesin*, ezrin*, radixin* (Moe); a/b-spectrin* (a/b-Spec); drebrin* (CG10083); coronin* (coro,pod1); MLLT4/Afadin-6* (cno); CTTN*(Cortactin); PPP1R9A* (spinophilin)b
Minus-end stabilisers c-tubulin ring complex (cTub23C and other components);CAMSAP1* (Patronin)
Tropomodulin* (tmod)b; Arp2/3 complex (Arpc1,Arp66B, etc.)
Plus-end motors Type 1 kinesins (Khc, Klc, Pat1); type 2 kinesins (Klp64D,Klp68D, Kif3C, Kap3); type 3 kinesins (unc-104, Khc-73)
Myosin XVA* (Myo10A); myosin VA* (didum);myosin II (zip, sqh)
Minus-end motors cytoplasmic dynein/dynactin (Dhc64C, ctp, sw, Gl, etc.); type13 kinesins/KIF2C/MCAK* (Klp10A, Klp59C, Klp59D)
Myosin VI* (jar)
Severers, de-polymerisers katanin* (Kat60); spastin* (spas); type 13 kinesins/KIF2C/MCAK* (Klp10A, Klp59C, Klp59D)
Cofilin* (tsr); gelsolin* (Gel)
Post-translational modifications(enzymes not listed)c
GTP hydrolysis; acetylation; (poly-) glycylation; (poly-)glutamylation; de-tyrosination; de-glutamylation (D2 tubulin)
ATP hydrolysis; potentially acetylation, arginylation,phosphorylation, etc.
Some proteins fulfil more than one function and are repeated in different positions; terms highlighted with an asterisk are listed in OMIM, other terms use otherdescriptors for protein complexes or classes. Terms in brackets are the orthologous Drosophila genes with known functions and/or prominent expression in thenervous system, as indicated in FlyBase (McQuilton et al., 2012).
aCib represents a triple-repeat of B4-thymosin.bNot declared as orthologues in FlyBase in spite of partial sequence homologies.cAs described by Janke and Bulinski, 2011; Terman and Kashina, 2013.
Cytoskeletal machinery of neurons 2335
Journ
alof
Cell
Scie
nce
the central nervous system (CNS) (Goncalves-Pimentel et al.,
2011; Matusek et al., 2008). Psidin (known as NAA25 in
mammals) is the auxiliary subunit of the NatB acetylation
complex required for protein acetylation (Stephan et al., 2012). In
addition, it has been shown that Psidin can act outside this
complex as an ABP that regulates actin dynamics both in neurons
and in non-neuronal cells of Drosophila (Kim et al., 2011;
Stephan et al., 2012). These studies show that Psidin antagonises
F-actin stabilisation mediated by Tropomyosin 1 in lamellipodia,
which is required for the pathfinding of olfactory neurons in the
fly brain. Canoe and its mammalian homologue MLLT4 (also
known as afadin, AFD6) are PDZ-domain-containing ABPs that
localise to adherens junctions and have been shown to interact
with profilin (Boettner et al., 2000; Lorger and Moelling, 2006;
Sawyer et al., 2009). In Canoe-deficient embryos, axonal
pathfinding at the CNS midline is impaired, and this aberration
correlates with two growth cone phenotypes: an increase in
filopodia numbers and a failure to localise Robo proteins
(receptors that are important regulators of midline pathfinding)
to filopodia (Slovakova et al., 2012). In this scenario, Canoe
EB1Exba /eIF5C
Tau
DA
AM
Arp
2/3
Enab
led
Prof
ilin
CpA
+BPs
isin
Tm1
Shot
Shot
MAP1BGRDC-tail
MtLSEFSR-rodplakinCH
EB1MTsExba/eIF5CF-actin
Axon extension
FilopodiaLamellipodia
Cortex
Other MTs
ApcDyneinType 13 kinesinsCLIP190
CLASP
XMAPDCX STMN
Ssp2
Actin bindingMT binding
EB1 binding (+TIP)Tubulin binding
Other binding
Actin regulatingOther regulatingprocessesSub-machinery
2 Actin dynamics
1 MT plus-end dynamics3 MTshaft
stability
MT growth
α-tubulin– F-actin
B
A
Key
β-tubulin
Interactions
Fig. 2. Roles of Shot in MT regulation at the interface with other classes of ABPs and MTBPs. (A) The functional domains of Shot (below the structure) and
their reported molecular interactions (above the structure). CH, calponin homology domains; plakin, plakin-like domain; SR-rod, spectrin repeat rod; EF, EF-hand
motifs; GRD, Gas2-related domain; MtLS, microtubule tip localisation sequence; C-tail, C-terminal tail. (B) A close-up of a growth cone showing a single MT and
three different aspects of the cytoskeletal machinery (emboxed and numbered), all of which are being studied in fly neurons. (1) Regulation of MT plus-end
dynamics; it includes end-binding proteins (here EB1) which directly bind MT plus ends and recruit MT-plus-end-tracking proteins (+TIPs) (blue arrows; see
Akhmanova and Steinmetz, 2008; Etienne-Manneville, 2010). Shot has to compete with other +TIPs for binding to a limited pool of EB1; when bound to EB1, it
links MT plus ends to F-actin (via its N-terminal CH domains) and guides polymerising MT plus ends in the direction of axon growth. The proper localisation of
Shot at MT plus ends also requires direct association with MTs through its GRD and C-terminal tail. CLASPs and XMAP bind tubulin and promote MT
polymerisation; CLASPs are +TIPs whereas XMAP215 (XMAP) binds MT plus-ends either directly or through +TIPs: Small spindles 2 (Ssp2) in Drosophila and
Slains in mammals (not shown; Al-Bassam and Chang, 2011; Currie et al., 2011; van der Vaart et al., 2012; Li et al., 2011; Lowery et al., 2010). Stathmins
(STMN) sequester tubulin (Manna et al., 2009), and doublecortin (DCX) binds MT plus ends independently (Bechstedt and Brouhard, 2012). (2) Actin dynamics.
The Shot–actin linkage is influenced by actin regulators (red stippled arrows) that coordinate the dynamics and structure of F-actin networks (cortex, lamellipodia,
filopodia; Pak et al., 2008; Xu et al., 2013). In the context of filopodia formation, Shot contributes to the regulation of F-actin through binding of Exba/eIF5C at its
EF-hand motifs (Lee et al., 2007; Sanchez-Soriano et al., 2009). (3) MT shaft stability. In addition, Shot binds along MT shafts and stabilises them independent of
its linkage to F-actin (Sanchez-Soriano et al., 2009; Alves-Silva et al., 2012). Other MT shaft binders are Tau which protects MTs against the MT-severing protein
Katanin in axons (not shown; Qiang et al., 2006), and mammalian MAP1B can cross-link MTs as well as mediate actin-MT linkage during axon growth (Bouquet
et al., 2007; Riederer, 2007). Tau, MAP1B and Shot (in an EB1-independent mode) influence MT polymerisation kinetics through yet unknown mechanisms
(stippled grey arrow; Alves-Silva et al., 2012; Feinstein and Wilson, 2005; Tymanskyj et al., 2012).
Journal of Cell Science 126 (11)2336
Journ
alof
Cell
Scie
nce
might regulate the number of filopodia directly through its
function as an ABP, or indirectly through acting as a scaffold forRobo receptors, since loss of Robo has been previously shownto lead to an increase in the number of filopodia (Murray and
Whitington, 1999).
Use of the fly to study the functional integrationof ABPs during filopodia regulationDrosophila growth cones have been used to carry out loss-of-function analyses of other ABP-encoding genes, including Arpc1
(ARP2), Arp3 (ARP3), capping protein A (CAPZA), capping
protein B (CAPZB), chickadee (profilin), enabled (ENAH, also
known as VASP), singed (fascin) and tropomyosin 1 (TPM1)(human homologues are given in the brackets) (Goncalves-Pimentel et al., 2011; Kraft et al., 2006; Sanchez-Soriano et al.,
2010). All these data and tools can now be combined tounderstand the functional networks of ABPs, as is best illustratedfor studies of filopodia formation. Filopodia are prominent actin-
based structures, and data from numerous studies in a wide rangeof cellular models have led to the formulation of two compositemodels of filopodia formation (Mattila and Lappalainen, 2008).In the ‘convergent elongation model’, Arp2/3 generates actin
filaments in lamellipodia, and ENAH/VASP aggregates their plusends to give rise to elongating filopodial bundles. In the ‘de novo
nucleation model’, formin molecules aggregate to generate
clusters of new actin filaments, which then directly elongateinto parallel filopodial bundles.
Drosophila neurons were used to test whether these two
models co-exist in neurons. Single-mutant analyses of the factorsessential in these models (i.e. Enabled, the formin DAAM anddifferent Arp2/3 complex components) all caused a partial butsignificant reduction in filopodia numbers, in agreement with the
potential co-existence of both models (Goncalves-Pimentel et al.,2011; Matusek et al., 2008). When loss of the Arp2/3 componentSop2 was combined with functional loss of DAAM in the same
neurons (DAAM2/2 Sop22/2 double-mutant neurons), filopodiawere completely absent and F-actin levels were severely depleted(Goncalves-Pimentel et al., 2011). To our knowledge, a genetic
condition that depletes F-actin to this degree has not beenreported for other eukaryotic systems. It indicates that Arp2/3 andDAAM are the prevailing nucleators in embryonic Drosophila
neurons. The fact that the two nucleators enhance each others’
phenotypes indicates that they work in distinct complementaryfunctional pathways during filopodia formation. This isconsistent with the biochemical evidence that Arp2/3 and
formins nucleate actin filaments through different molecularmechanisms (Chesarone and Goode, 2009; Mattila andLappalainen, 2008). However, it does not yet resolve whether
these two nucleators contribute to two entirely separate processesof filopodia formation.
To make this distinction, genetic interaction studies usingpartial loss of gene functions are very helpful. For example, in
heterozygous gene constellation (i.e. one mutant and one normalgene copy), the levels of the proteins they encode tend to be onlymoderately reduced, and, typically, this reduction is not sufficient
to cause a phenotype. Accordingly, neurons that are heterozygousfor DAAM, Sop2 or ena show normal filopodia numbers(Goncalves-Pimentel et al., 2011). However, any combinations
of these heterozygous conditions (transheterozygous mutantneurons: DAAM2/+ Sop22/+ or DAAM2/+ ena2/+ or Sop22/+
ena2/+) become functionally rate-limiting and lead to a strong
reduction in filopodia numbers (Goncalves-Pimentel et al., 2011).
These findings are inconsistent with regard to whether twocompletely separate modes of filopodia formation exist, but dosuggest that the pathways downstream of the two nucleators
converge at some point. Therefore, these authors suggested amore general model of convergent elongation, in which not onlyArp2/3-derived but also DAAM-derived actin filaments can beused by Ena to be transformed into filopodial actin bundles
(Goncalves-Pimentel et al., 2011). These examples illustrate howsystematic genetic analyses used in Drosophila neurons cancontribute to the formulation of new mechanistic models of
cytoskeletal regulation.
Use of the fly to study mechanisms of actin–MTlinkage during axonal growth and maintenanceActin–MT crosstalk is an important aspect of cytoskeletalregulation in growing axons, but its mechanisms are littleunderstood (Box 2). Promising contributions have been made by
work on the above mentioned spectraplakins Shot and ACF7.Besides filopodia-associated phenotypes, we have shown thatloss of Shot in fly neurons and loss of the mouse homologueACF7 in mouse neurons result in another conserved phenotype –
disorganised axonal MT bundles that is coupled with reducedaxon growth (Sanchez-Soriano et al., 2009). Detailed work onShot has shown that these functions depend on actin–MT linkage.
Structure–function analyses have revealed that functions of Shotin MT organisation depend on three simultaneous molecularinteractions (Fig. 2): (1) binding of its N-terminal calponin
homology domains to F-actin, (2) association to MTs throughtwo C-terminal domains (the Gas2-related domain and thepositively charged C-terminal tail), and (iii) binding of C-
terminal tail to EB1 (end-binding protein 1) at MT plus ends(Alves-Silva et al., 2012; Bottenberg et al., 2009; Lee andKolodziej, 2002; Sanchez-Soriano et al., 2009). These findingssuggest a model in which Shot binds to the plus ends of
polymerising MTs and guides them along actin structures in thedirection of axon growth (Alves-Silva et al., 2012). This model isfurther supported by the observation that frequent off-track
extensions of MTs are seen during live imaging of shot mutantneurons; this can explain the observed MT disorganisation anddiminished axon growth (off-track MTs are less likely to
contribute to axon extension). The guidance model is alsosupported by shot-like MT-disorganisation phenotypes observedin EB1-deficient, as well as in shot2/+ eb12/+ transheterozygous
mutant neurons (Alves-Silva et al., 2012), and, furthermore, isconsistent with proposed roles of ACF7 in guiding MT extensionalong actin stress fibres towards focal adhesions in non-neuronalcells (Kodama et al., 2004).
Notably, the MT guidance model offers a promising molecularplatform for further research. On the one hand, it can be used tounderstand the roles of other cytoskeletal regulators during MT
guidance, for example of ABPs or the various proteins that bindand regulate MT plus ends (see details in Fig. 2). On the otherhand, the MT guidance model suggests a potential mechanismcontributing to the maintenance of axons in the mature nervous
system, as will be explained in the following.
The second mammalian homologue of Shot, dystonin, isstrongly expressed in peripheral neural ganglia (Leung et al.,
2001), which, in dystonin mutant mice, undergo a severeneurodegeneration at postnatal stages that is referred to asHSAN (hereditary sensory autonomic neuropathy) (Duchen et al.,
Cytoskeletal machinery of neurons 2337
Journ
alof
Cell
Scie
nce
1963; Duchen et al., 1964). The underlying mechanisms remain
inconclusive and were proposed to be associated with roles ofdystonin in intermediate filament organisation and axonaltransport, and in ER or Golgi dysfunction (Ferrier et al., 2013;
Young and Kothary, 2007). We propose instead that HSANpathology might be primarily caused by defects in MT guidanceand stabilisation on the basis of the following argument. AxonalMT bundles are believed to be stabilised through structural
MTBPs, such as tau or MAP1B (Fig. 2) (Chilton and Gordon-Weeks, 2007). At the same time, there appears to be a steadyand dynamic MT turnover, as suggested by continued MT
polymerisation events in mature axons (Kollins et al., 2009).Spectraplakins can regulate both functions through twofunctionally conserved domains at their C-terminus: the Gas2-
related domain can stabilise MTs against depolymerisation,whereas the C-terminal tail mediates the aforementioned bindingto EB1 required for MT guidance. These domains jointly mediate
a robust association along the shaft and at the tip of MTs that isrequired for both MT guidance and stabilisation (Fig. 2) (Alves-Silva et al., 2012; Honnappa et al., 2009; Sun et al., 2001).Absence of either domain in Shot causes severe MT
disorganisation and loss of MT stability in axons of developingfly neurons (Alves-Silva et al., 2012; Sanchez-Soriano et al.,2009) and, in addition, mature sensory neurons of dystonin
mutant mice, which are prone to degeneration display severe MTdisorganisation and loss of MT stability (Ferrier et al., 2013;Yang et al., 1999). Consistently, a recently reported frame-shift
mutation in the human dystonin gene functionally deletes theGas2-related domain and the C-terminal tail and causes a severeform of HSAN that mirrors the degenerative pathology of dystonin
mutant mice (Edvardson et al., 2012). These data suggest thatspectraplakins perform similar MT-regulating functions during thedevelopment of fly neurons and the maintenance of sensorymammalian neurons. Severe MT disorganisation and instability
caused by absence of spectraplakins can, therefore, be expected toaffect vital functions such as axonal transport and impactnegatively on neuronal survival (Sheng and Cai, 2012).
This example shows how functional studies of cytoskeletalregulators during axon growth in the fly can provide new ideasand testable hypotheses for research on other aspects of neuronalcytoskeleton, even in other animals.
Conclusions and future perspectivesIn this Commentary, we have discussed how systematic andcombinatorial genetic studies of different ABPs and MTBPs in
the neuronal system of the fly can improve our understandingof cytoskeletal regulation during a biological process such asaxon growth. Considering the high degree of evolutionary
conservation of actin, tubulin and their regulators, theenormous similarities of cytoskeletal dynamics in fly andvertebrate neurons, and the similarities of mutant phenotypes
reported so far, we feel that insights gained in the fly system willbe informative for cytoskeletal research in mammalian or othervertebrate neurons.
Understanding the cytoskeletal regulation that underpins
cellular behaviours will have important implications andapplications. For example, we still have little understanding ofhow signalling mechanisms instruct the cytoskeleton to influence
growth cone behaviours (Huber et al., 2003). Knowing how thevarious ABPs and MTBPs that are targeted by particularsignalling events contribute to the cellular cytoskeletal
machinery will greatly help to close this knowledge gap.
Understanding these cellular roles of cytoskeletal regulators isalso the first step on the long path to developing mathematicalmodels that can describe the cytoskeletal dynamics that are at the
basis of different cellular processes (Oelz et al., 2008), and it willfacilitate the introduction of systems approaches for cell biologyresearch (Liberali and Pelkmans, 2012). Furthermore, theprincipal strategies described here for axon growth can also be
applied to the investigation of the roles and regulation of thecytoskeleton in other relevant areas, such as synapse formation,axonal pruning and Wallerian degeneration, axon regeneration
and neurodegeneration. Notably, for all of these processes,suitable Drosophila models have already been established, thushopefully paving the way for rapid progress (Bossing et al., 2012;
Fang and Bonini, 2012; Gistelinck et al., 2012; Goellner andAberle, 2012; Jaiswal et al., 2012).
AcknowledgementsWe would like to thank Tom Millard, Laura Anne Lowery, PhillipGordon-Weeks, Joszef Mihaly and Erik Dent for their very helpfuland constructive comments on this manuscript.
FundingAuthors on this Commentary were funded by the Biotechnology andBiological Sciences Research Council (BBSRC) [grant numbers BB/D526561/1, BB/I002448/1]. Deposited in PMC for immediate release.
Supplementary material available online at
http://jcs.biologists.org/lookup/suppl/doi:10.1242/jcs.126912/-/DC1
ReferencesAdalbert, R., Nogradi, A., Babetto, E., Janeckova, L., Walker, S. A.,
Kerschensteiner, M., Misgeld, T. and Coleman, M. P. (2009). Severely dystrophicaxons at amyloid plaques remain continuous and connected to viable cell bodies.Brain 132, 402-416.
Ahmad, F. J., Yu, W., McNally, F. J. and Baas, P. W. (1999). An essential role forkatanin in severing microtubules in the neuron. J. Cell Biol. 145, 305-315.
Akhmanova, A. and Stearns, T. (2013). Cell architecture: putting the building blockstogether. Curr. Opin. Cell Biol. 25, 3-5.
Akhmanova, A. and Steinmetz, M. O. (2008). Tracking the ends: a dynamic proteinnetwork controls the fate of microtubule tips. Nat. Rev. Mol. Cell Biol. 9, 309-322.
Akhmanova, A. and Steinmetz, M. O. (2010). Microtubule +TIPs at a glance. J. Cell
Sci. 123, 3415-3419.
Al-Bassam, J. and Chang, F. (2011). Regulation of microtubule dynamics by TOG-domain proteins XMAP215/Dis1 and CLASP. Trends Cell Biol. 21, 604-614.
Alves-Silva, J., Sanchez-Soriano, N., Beaven, R., Klein, M., Parkin, J., Millard,T. H., Bellen, H. J., Venken, K. J. T., Ballestrem, C., Kammerer, R. A. et al.
(2012). Spectraplakins promote microtubule-mediated axonal growth by functioningas structural microtubule-associated proteins and EB1-dependent +TIPs (tipinteracting proteins). J. Neurosci. 32, 9143-9158.
Araujo, S. J. and Tear, G. (2003). Axon guidance mechanisms and molecules: lessonsfrom invertebrates. Nat. Rev. Neurosci. 4, 910-922.
Astigarraga, S., Hofmeyer, K. and Treisman, J. E. (2010). Missed connections:photoreceptor axon seeks target neuron for synaptogenesis. Curr. Opin. Genet. Dev.
20, 400-407.
Baas, P. W. and Lin, S. (2011). Hooks and comets: The story of microtubule polarityorientation in the neuron. Dev. Neurobiol. 71, 403-418.
Barnes, A. P. and Polleux, F. (2009). Establishment of axon-dendrite polarity indeveloping neurons. Annu. Rev. Neurosci. 32, 347-381.
Basto, R., Lau, J., Vinogradova, T., Gardiol, A., Woods, C. G., Khodjakov, A. andRaff, J. W. (2006). Flies without centrioles. Cell 125, 1375-1386.
Bear, J. E. and Gertler, F. B. (2009). Ena/VASP: towards resolving a pointedcontroversy at the barbed end. J. Cell Sci. 122, 1947-1953.
Bechstedt, S. and Brouhard, G. J. (2012). Doublecortin recognizes the 13-protofilament microtubule cooperatively and tracks microtubule ends. Dev. Cell 23,181-192.
Bellen, H. J., Tong, C. and Tsuda, H. (2010). 100 years of Drosophila research and itsimpact on vertebrate neuroscience: a history lesson for the future. Nat. Rev. Neurosci.
11, 514-522.
Bielas, S. L., Serneo, F. F., Chechlacz, M., Deerinck, T. J., Perkins, G. A., Allen,
P. B., Ellisman, M. H. and Gleeson, J. G. (2007). Spinophilin facilitatesdephosphorylation of doublecortin by PP1 to mediate microtubule bundling at theaxonal wrist. Cell 129, 579-591.
Journal of Cell Science 126 (11)2338
Journ
alof
Cell
Scie
nce
Boettner, B., Govek, E. E., Cross, J. and Van Aelst, L. (2000). The junctionalmultidomain protein AF-6 is a binding partner of the Rap1A GTPase and associateswith the actin cytoskeletal regulator profilin. Proc. Natl. Acad. Sci. USA 97, 9064-9069.
Bossing, T., Barros, C. S., Fischer, B., Russell, S. and Shepherd, D. (2012).Disruption of microtubule integrity initiates mitosis during CNS repair. Dev. Cell 23,433-440.
Bottenberg, W., Sanchez-Soriano, N., Alves-Silva, J., Hahn, I., Mende, M. and
Prokop, A. (2009). Context-specific requirements of functional domains of theSpectraplakin Short stop in vivo. Mech. Dev. 126, 489-502.
Bouquet, C., Ravaille-Veron, M., Propst, F. and Nothias, F. (2007). MAP1Bcoordinates microtubule and actin filament remodeling in adult mouse Schwann celltips and DRG neuron growth cones. Mol. Cell. Neurosci. 36, 235-247.
Bradke, F., Fawcett, J. W. and Spira, M. E. (2012). Assembly of a new growth coneafter axotomy: the precursor to axon regeneration. Nat. Rev. Neurosci. 13, 183-193.
Brandt, R. (1998). Cytoskeletal mechanisms of axon outgrowth and pathfinding. Cell
Tissue Res. 292, 181-189.
Brochtrup, A. and Hummel, T. (2011). Olfactory map formation in the Drosophila
brain: genetic specificity and neuronal variability. Curr. Opin. Neurobiol. 21, 85-92.
Buck, K. B. and Zheng, J. Q. (2002). Growth cone turning induced by direct localmodification of microtubule dynamics. J. Neurosci. 22, 9358-9367.
Challacombe, J. F., Snow, D. M. and Letourneau, P. C. (1996). Actin filamentbundles are required for microtubule reorientation during growth cone turning toavoid an inhibitory guidance cue. J. Cell Sci. 109, 2031-2040.
Challacombe, J. F., Snow, D. M. and Letourneau, P. C. (1997). Dynamic microtubuleends are required for growth cone turning to avoid an inhibitory guidance cue.J. Neurosci. 17, 3085-3095.
Chen, S., Chen, J., Shi, H., Wei, M., Castaneda-Castellanos, D. R., Bultje, R. S., Pei,
X., Kriegstein, A. R., Zhang, M. and Shi, S.-H. (2013). Regulation of microtubulestability and organization by mammalian Par3 in specifying neuronal polarity. Dev.
Cell 24, 26-40.
Chesarone, M. A. and Goode, B. L. (2009). Actin nucleation and elongation factors:mechanisms and interplay. Curr. Opin. Cell Biol. 21, 28-37.
Chien, C. B., Rosenthal, D. E., Harris, W. A. and Holt, C. E. (1993). Navigationalerrors made by growth cones without filopodia in the embryonic Xenopus brain.Neuron 11, 237-251.
Chilton, J. and Gordon-Weeks, P. (2007). Role of microtubules and MAPs duringneuritogenesis. In Intracellular Mechanisms for Neuritogenesis (ed. I. de Curtis), pp.57-88. New York, NY: Springer.
Conde, C. and Caceres, A. (2009). Microtubule assembly, organization and dynamicsin axons and dendrites. Nat. Rev. Neurosci. 10, 319-332.
Currie, J. D., Stewman, S., Schimizzi, G., Slep, K. C., Ma, A. and Rogers, S. L.
(2011). The microtubule lattice and plus-end association of Drosophila Mini spindlesis spatially regulated to fine-tune microtubule dynamics. Mol. Biol. Cell 22, 4343-4361.
Davenport, R. W., Dou, P., Rehder, V. and Kater, S. B. (1993). A sensory role forneuronal growth cone filopodia. Nature 361, 721-724.
Dent, E. W., Kwiatkowski, A. V., Mebane, L. M., Philippar, U., Barzik, M.,
Rubinson, D. A., Gupton, S., Van Veen, J. E., Furman, C., Zhang, J. et al. (2007).Filopodia are required for cortical neurite initiation. Nat. Cell Biol. 9, 1347-1359.
Dent, E. W., Gupton, S. L. and Gertler, F. B. (2011). The growth cone cytoskeleton inaxon outgrowth and guidance. Cold Spring Harb. Perspect. Biol. 3, 3.
Duchen, L. W., Falconer, D. S. and Strich, S. J. (1963). Dystonia musculorum. Ahereditary neuropathy of mice affecting mainly sensory pathways. J. Physiol. 165, 7P-9P.
Duchen, L. W., Strich, S. J. and Falconer, D. S. (1964). Clinical and pathologicalstudies of a hereditary neuropathy in mice (dystonia musculorum). Brain 87, 367-378.
Dwivedy, A., Gertler, F. B., Miller, J., Holt, C. E. and Lebrand, C. (2007). Ena/VASP function in retinal axons is required for terminal arborization but not pathwaynavigation. Development 134, 2137-2146.
Edvardson, S., Cinnamon, Y., Jalas, C., Shaag, A., Maayan, C., Axelrod, F. B. and
Elpeleg, O. (2012). Hereditary sensory autonomic neuropathy caused by a mutationin dystonin. Ann. Neurol. 71, 569-572.
Etienne-Manneville, S. (2010). From signaling pathways to microtubule dynamics: thekey players. Curr. Opin. Cell Biol. 22, 104-111.
Fan, J., Mansfield, S. G., Redmond, T., Gordon-Weeks, P. R. and Raper, J. A.
(1993). The organization of F-actin and microtubules in growth cones exposed to abrain-derived collapsing factor. J. Cell Biol. 121, 867-878.
Fang, Y. and Bonini, N. M. (2012). Axon degeneration and regeneration: insights fromDrosophila models of nerve injury. Annu. Rev. Cell Dev. Biol. 28, 575-597.
Feinstein, S. C. and Wilson, L. (2005). Inability of tau to properly regulate neuronalmicrotubule dynamics: a loss-of-function mechanism by which tau might mediateneuronal cell death. Biochim. Biophys. Acta 1739, 268-279.
Ferrier, A., Boyer, J. G. and Kothary, R. (2013). Cellular and molecular biology ofneuronal dystonin. Int. Rev. Cell Mol. Biol. 300, 85-120.
Fiala, J. C., Feinberg, M., Peters, A. and Barbas, H. (2007). Mitochondrialdegeneration in dystrophic neurites of senile plaques may lead to extracellulardeposition of fine filaments. Brain Struct. Funct. 212, 195-207.
Fletcher, D. A. and Mullins, R. D. (2010). Cell mechanics and the cytoskeleton. Nature
463, 485-492.
Flynn, K. C., Hellal, F., Neukirchen, D., Jacob, S., Tahirovic, S., Dupraz, S., Stern,
S., Garvalov, B. K., Gurniak, C., Shaw, A. E. et al. (2012). ADF/cofilin-mediated
actin retrograde flow directs neurite formation in the developing brain. Neuron 76,1091-1107.
Fukushima, N., Furuta, D., Hidaka, Y., Moriyama, R. and Tsujiuchi, T. (2009).Post-translational modifications of tubulin in the nervous system. J. Neurochem. 109,683-693.
Gallo, G. (2013). Mechanisms underlying the initiation and dynamics of neuronalfilopodia: from neurite formation to synaptogenesis. Int. Rev. Cell Mol. Biol. 301, 95-156.
Geraldo, S., Khanzada, U. K., Parsons, M., Chilton, J. K. and Gordon-Weeks, P. R.
(2008). Targeting of the F-actin-binding protein drebrin by the microtubule plus-tipprotein EB3 is required for neuritogenesis. Nat. Cell Biol. 10, 1181-1189.
Giannone, G., Mege, R. M. and Thoumine, O. (2009). Multi-level molecular clutchesin motile cell processes. Trends Cell Biol. 19, 475-486.
Gistelinck, M., Lambert, J. C., Callaerts, P., Dermaut, B. and Dourlen, P. (2012).Drosophila models of tauopathies: what have we learned? Int. J. Alzheimers Dis.
2012, 970980.
Goellner, B. and Aberle, H. (2012). The synaptic cytoskeleton in development anddisease. Dev. Neurobiol. 72, 111-125.
Goncalves-Pimentel, C., Gombos, R., Mihaly, J., Sanchez-Soriano, N. and Prokop,A. (2011). Dissecting regulatory networks of filopodia formation in a Drosophila
growth cone model. PLoS ONE 6, e18340.
Grubb, M. S. and Burrone, J. (2010). Building and maintaining the axon initialsegment. Curr. Opin. Neurobiol. 20, 481-488.
Hawkins, T., Mirigian, M., Selcuk Yasar, M. and Ross, J. L. (2010). Mechanics ofmicrotubules. J. Biomech. 43, 23-30.
Hirokawa, N., Niwa, S. and Tanaka, Y. (2010). Molecular motors in neurons: transportmechanisms and roles in brain function, development, and disease. Neuron 68, 610-638.
Honnappa, S., Gouveia, S. M., Weisbrich, A., Damberger, F. F., Bhavesh, N. S.,
Jawhari, H., Grigoriev, I., van Rijssel, F. J., Buey, R. M., Lawera, A. et al. (2009).An EB1-binding motif acts as a microtubule tip localization signal. Cell 138, 366-376.
Huber, A. B., Kolodkin, A. L., Ginty, D. D. and Cloutier, J. F. (2003). Signaling atthe growth cone: ligand-receptor complexes and the control of axon growth andguidance. Annu. Rev. Neurosci. 26, 509-563.
Hur, E. M., Saijilafu, Lee, B. D, Kim, S. J, Xu, W. L. and Zhou, F. Q. (2011). GSK3controls axon growth via CLASP-mediated regulation of growth cone microtubules.Genes Dev. 25, 1968-1981.
Insall, R. H. and Machesky, L. M. (2009). Actin dynamics at the leading edge: fromsimple machinery to complex networks. Dev. Cell 17, 310-322.
Jaiswal, M., Sandoval, H., Zhang, K., Bayat, V. and Bellen, H. J. (2012). Probingmechanisms that underlie human neurodegenerative diseases in Drosophila. Annu.
Rev. Genet. 46, 371-396.
Janke, C. and Bulinski, J. C. (2011). Post-translational regulation of the microtubulecytoskeleton: mechanisms and functions. Nat. Rev. Mol. Cell Biol. 12, 773-786.
Kaufmann, N., Wills, Z. P. and Van Vactor, D. (1998). Drosophila Rac1 controlsmotor axon guidance. Development 125, 453-461.
Keller, E. F. (1996). Drosophila embryos as transitional objects: the work of DonaldPoulson and Christiane Nusslein-Volhard. Hist. Stud. Phys. Biol. Sci. 26, 313-346.
Kim, J. H., Cho, A., Yin, H., Schafer, D. A., Mouneimne, G., Simpson, K. J.,Nguyen, K. V., Brugge, J. S. and Montell, D. J. (2011). Psidin, a conserved proteinthat regulates protrusion dynamics and cell migration. Genes Dev. 25, 730-741.
Kodama, A., Lechler, T. and Fuchs, E. (2004). Coordinating cytoskeletal tracks topolarize cellular movements. J. Cell Biol. 167, 203-207.
Kollins, K. M., Bell, R. L., Butts, M. and Withers, G. S. (2009). Dendrites differ fromaxons in patterns of microtubule stability and polymerization during development.Neural Dev. 4, 26.
Koper, A., Schenck, A. and Prokop, A. (2012). Analysis of adhesion molecules andbasement membrane contributions to synaptic adhesion at the Drosophila embryonicNMJ. PLoS ONE 7, e36339.
Kraft, R., Escobar, M. M., Narro, M. L., Kurtis, J. L., Efrat, A., Barnard, K. and
Restifo, L. L. (2006). Phenotypes of Drosophila brain neurons in primary culturereveal a role for fascin in neurite shape and trajectory. J. Neurosci. 26, 8734-8747.
Kuppers-Munther, B., Letzkus, J. J., Luer, K., Technau, G., Schmidt, H. and
Prokop, A. (2004). A new culturing strategy optimises Drosophila primary cellcultures for structural and functional analyses. Dev. Biol. 269, 459-478.
Kwiatkowski, A. V., Rubinson, D. A., Dent, E. W., Edward van Veen, J., Leslie,
J. D., Zhang, J., Mebane, L. M., Philippar, U., Pinheiro, E. M., Burds, A. A. et al.
(2007). Ena/VASP Is Required for neuritogenesis in the developing cortex. Neuron
56, 441-455.
Lamoureux, P., Heidemann, S. R., Martzke, N. R. and Miller, K. E. (2010). Growthand elongation within and along the axon. Dev. Neurobiol. 70, 135-149.
Landgraf, M. and Thor, S. (2006). Development of Drosophila motoneurons:specification and morphology. Semin. Cell Dev. Biol. 17, 3-11.
Lee, S. and Kolodziej, P. A. (2002). Short Stop provides an essential link between F-actin and microtubules during axon extension. Development 129, 1195-1204.
Lee, A. C. and Suter, D. M. (2008). Quantitative analysis of microtubule dynamicsduring adhesion-mediated growth cone guidance. Dev. Neurobiol. 68, 1363-1377.
Lee, S., Nahm, M., Lee, M., Kwon, M., Kim, E., Zadeh, A. D., Cao, H., Kim, H. J.,Lee, Z. H., Oh, S. B. et al. (2007). The F-actin-microtubule crosslinker Shot is aplatform for Krasavietz-mediated translational regulation of midline axon repulsion.Development 134, 1767-1777.
Letourneau, P. C. (2009). Actin in axons: stable scaffolds and dynamic filaments.Results Probl. Cell Differ. 48, 265-290.
Cytoskeletal machinery of neurons 2339
Journ
alof
Cell
Scie
nce
Letourneau, P. C. and Ressler, A. H. (1984). Inhibition of neurite initiation and growth
by taxol. J. Cell Biol. 98, 1355-1362.
Leung, C. L., Zheng, M., Prater, S. M. and Liem, R. K. (2001). The BPAG1 locus:
Alternative splicing produces multiple isoforms with distinct cytoskeletal linker
domains, including predominant isoforms in neurons and muscles. J. Cell Biol. 154,
691-697.
Li, W., Miki, T., Watanabe, T., Kakeno, M., Sugiyama, I., Kaibuchi, K. and
Goshima, G. (2011). EB1 promotes microtubule dynamics by recruiting Sentin in
Drosophila cells. J. Cell Biol. 193, 973-983.
Liberali, P. and Pelkmans, L. (2012). Towards quantitative cell biology. Nat. Cell Biol.
14, 1233.
Lorger, M. and Moelling, K. (2006). Regulation of epithelial wound closure and
intercellular adhesion by interaction of AF6 with actin cytoskeleton. J. Cell Sci. 119,
3385-3398.
Lowery, L. A. and Van Vactor, D. (2009). The trip of the tip: understanding the growth
cone machinery. Nat. Rev. Mol. Cell Biol. 10, 332-343.
Lowery, L. A., Lee, H., Lu, C., Murphy, R., Obar, R. A., Zhai, B., Schedl, M., Van
Vactor, D. and Zhan, Y. (2010). Parallel genetic and proteomic screens identify
Msps as a CLASP-Abl pathway interactor in Drosophila. Genetics 185, 1311-
1325.
Lucas, F. R., Goold, R. G., Gordon-Weeks, P. R. and Salinas, P. C. (1998). Inhibition
of GSK-3beta leading to the loss of phosphorylated MAP-1B is an early event in
axonal remodelling induced by WNT-7a or lithium. J. Cell Sci. 111, 1351-1361.
Manna, T., Thrower, D. A., Honnappa, S., Steinmetz, M. O. and Wilson, L. (2009).
Regulation of microtubule dynamic instability in vitro by differentially phosphorylated
stathmin. J. Biol. Chem. 284, 15640-15649.
Marner, L., Nyengaard, J. R., Tang, Y. and Pakkenberg, B. (2003). Marked loss of
myelinated nerve fibers in the human brain with age. J. Comp. Neurol. 462, 144-152.
Marsh, L. and Letourneau, P. C. (1984). Growth of neurites without filopodial or
lamellipodial activity in the presence of cytochalasin B. J. Cell Biol. 99, 2041-2047.
Mattie, F. J., Stackpole, M. M., Stone, M. C., Clippard, J. R., Rudnick, D. A., Qiu,
Y., Tao, J., Allender, D. L., Parmar, M. and Rolls, M. M. (2010). Directed
microtubule growth, +TIPs, and kinesin-2 are required for uniform microtubule
polarity in dendrites. Curr. Biol. 20, 2169-2177.
Mattila, P. K. and Lappalainen, P. (2008). Filopodia: molecular architecture and
cellular functions. Nat. Rev. Mol. Cell Biol. 9, 446-454.
Matusek, T., Gombos, R., Szecsenyi, A., Sanchez-Soriano, N., Czibula, A., Pataki,
C., Gedai, A., Prokop, A., Rasko, I. and Mihaly, J. (2008). Formin proteins of the
DAAM subfamily play a role during axon growth. J. Neurosci. 28, 13310-13319.
McCaig, C. D. (1989). Nerve growth in the absence of growth cone filopodia and the
effects of a small applied electric field. J Cell Sci 93, 715-721.
McCaig, C. D., Rajnicek, A. M., Song, B. and Zhao, M. (2002). Has electrical growth
cone guidance found its potential? Trends Neurosci. 25, 354-359.
McQuilton, P., St Pierre, S. E., Thurmond, J.; FlyBase Consortium (2012). FlyBase
101—the basics of navigating FlyBase. Nucleic Acids Res. 40 Database issue, D706-
D714.
Moore, S. W., Biais, N. and Sheetz, M. P. (2009). Traction on immobilized netrin-1 is
sufficient to reorient axons. Science 325, 166.
Moore, S. W., Roca-Cusachs, P. and Sheetz, M. P. (2010). Stretchy proteins on
stretchy substrates: the important elements of integrin-mediated rigidity sensing. Dev.
Cell 19, 194-206.
Morris, M., Maeda, S., Vossel, K. and Mucke, L. (2011). The many faces of tau.
Neuron 70, 410-426.
Moseley, J. B., Bartolini, F., Okada, K., Wen, Y., Gundersen, G. G. and Goode,
B. L. (2007). Regulated binding of adenomatous polyposis coli protein to actin.
J. Biol. Chem. 282, 12661-12668.
Murray, M. J. and Whitington, P. M. (1999). Effects of Roundabout on growth cone
dynamics, filopodial length, and growth cone morphology at the midline and
throughout the neuropile. J. Neurosci. 19, 7901-7912.
Myers, K. A., Tint, I., Nadar, C. V., He, Y., Black, M. M. and Baas, P. W. (2006).
Antagonistic forces generated by cytoplasmic dynein and myosin-II during growth
cone turning and axonal retraction. Traffic 7, 1333-1351.
Neukirchen, D. and Bradke, F. (2011). Neuronal polarization and the cytoskeleton.
Semin. Cell Dev. Biol. 22, 825-833.
Nugent, A. A., Kolpak, A. L. and Engle, E. C. (2012). Human disorders of axon
guidance. Curr. Opin. Neurobiol. 22, 837-843.
Oelz, D., Schmeiser, C. and Small, J. V. (2008). Modeling of the actin-cytoskeleton in
symmetric lamellipodial fragments. Cell Adh. Migr. 2, 117-126.
Okada, K., Bartolini, F., Deaconescu, A. M., Moseley, J. B., Dogic, Z., Grigorieff,
N., Gundersen, G. G. and Goode, B. L. (2010). Adenomatous polyposis coli protein
nucleates actin assembly and synergizes with the formin mDia1. J. Cell Biol. 189,
1087-1096.
Pak, C. W., Flynn, K. C. and Bamburg, J. R. (2008). Actin-binding proteins take the
reins in growth cones. Nat. Rev. Neurosci. 9, 136-147.
Pawson, C., Eaton, B. A. and Davis, G. W. (2008). Formin-dependent synaptic growth:
evidence that Dlar signals via Diaphanous to modulate synaptic actin and dynamic
pioneer microtubules. J. Neurosci. 28, 11111-11123.
Perrot, R. and Eyer, J. (2009). Neuronal intermediate filaments and neurodegenerative
disorders. Brain Res. Bull. 80, 282-295.
Pollitt, A. Y. and Insall, R. H. (2009). WASP and SCAR/WAVE proteins: the drivers
of actin assembly. J. Cell Sci. 122, 2575-2578.
Prokop, A., Sanchez-Soriano, N., Goncalves-Pimentel, C., Molnar, I., Kalmar,
T. and Mihaly, J. (2011). DAAM family members leading a novel path into forminresearch. Commun. Integr. Biol. 4, 538-542.
Prokop, A., Kuppers-Munther, B. and Sanchez-Soriano, N. (2012). Using primaryneuron cultures of Drosophila to analyse neuronal circuit formation and function. InThe Making and Un-Making Of Neuronal Circuits in Drosophila, (ed. B. A. Hassan),pp. 225-247. New York, NY: Springer Science & Business Media.
Purro, S. A., Ciani, L., Hoyos-Flight, M., Stamatakou, E., Siomou, E. and Salinas,P. C. (2008). Wnt regulates axon behavior through changes in microtubule growthdirectionality: a new role for adenomatous polyposis coli. J. Neurosci. 28, 8644-8654.
Qiang, L., Yu, W., Andreadis, A., Luo, M. and Baas, P. W. (2006). Tau protectsmicrotubules in the axon from severing by katanin. J. Neurosci. 26, 3120-3129.
Rajagopalan, J., Tofangchi, A. and A Saif, M. T. (2010). Drosophila neurons activelyregulate axonal tension in vivo. Biophys. J. 99, 3208-3215.
Riederer, B. M. (2007). Microtubule-associated protein 1B, a growth-associated andphosphorylated scaffold protein. Brain Res. Bull. 71, 541-558.
Roote, J. and Prokop, A. (2013). How to design a genetic mating scheme: a basictraining package for Drosophila genetics. G3 (Bethesda) 3, 353-358.
Rothenberg, M. E., Rogers, S. L., Vale, R. D., Jan, L. Y. and Jan, Y. N. (2003).Drosophila pod-1 crosslinks both actin and microtubules and controls the targeting ofaxons. Neuron 39, 779-791.
Sabry, J. H., O’Connor, T. P., Evans, L., Toroian-Raymond, A., Kirschner, M. and
Bentley, D. (1991). Microtubule behavior during guidance of pioneer neuron growthcones in situ. J. Cell Biol. 115, 381-395.
Saengsawang, W., Mitok, K., Viesselmann, C., Pietila, L., Lumbard, D. C., Corey,
S. J. and Dent, E. W. (2012). The F-BAR protein CIP4 inhibits neurite formation byproducing lamellipodial protrusions. Curr. Biol. 22, 494-501.
Sanchez-Soriano, N. and Prokop, A. (2005). The influence of pioneer neurons on agrowing motor nerve in Drosophila requires the neural cell adhesion moleculehomolog FasciclinII. J. Neurosci. 25, 78-87.
Sanchez-Soriano, N., Bottenberg, W., Fiala, A., Haessler, U., Kerassoviti, A., Knust,
E., Lohr, R. and Prokop, A. (2005). Are dendrites in Drosophila homologous tovertebrate dendrites? Dev. Biol. 288, 126-138.
Sanchez-Soriano, N., Tear, G., Whitington, P. and Prokop, A. (2007). Drosophila asa genetic and cellular model for studies on axonal growth. Neural Dev. 2, 9.
Sanchez-Soriano, N., Travis, M., Dajas-Bailador, F., Goncalves-Pimentel, C.,
Whitmarsh, A. J. and Prokop, A. (2009). Mouse ACF7 and Drosophila short stopmodulate filopodia formation and microtubule organisation during neuronal growth.J. Cell Sci. 122, 2534-2542.
Sanchez-Soriano, N., Goncalves-Pimentel, C., Beaven, R., Haessler, U., Ofner-Ziegenfuss, L., Ballestrem, C. and Prokop, A. (2010). Drosophila growth cones: agenetically tractable platform for the analysis of axonal growth dynamics. Dev.
Neurobiol. 70, 58-71.
Sawyer, J. K., Harris, N. J., Slep, K. C., Gaul, U. and Peifer, M. (2009). TheDrosophila afadin homologue Canoe regulates linkage of the actin cytoskeleton toadherens junctions during apical constriction. J. Cell Biol. 186, 57-73.
Saxena, S. and Caroni, P. (2007). Mechanisms of axon degeneration: fromdevelopment to disease. Prog. Neurobiol. 83, 174-191.
Schaefer, A. W., Kabir, N. and Forscher, P. (2002). Filopodia and actin arcs guide theassembly and transport of two populations of microtubules with unique dynamicparameters in neuronal growth cones. J. Cell Biol. 158, 139-152.
Sheng, Z. H. and Cai, Q. (2012). Mitochondrial transport in neurons: impact onsynaptic homeostasis and neurodegeneration. Nat. Rev. Neurosci. 13, 77-93.
Slovakova, J., Speicher, S., Sanchez-Soriano, N., Prokop, A. and Carmena, A.(2012). The actin-binding protein Canoe/AF-6 forms a complex with Robo and isrequired for Slit-Robo signaling during axon pathfinding at the CNS midline.J. Neurosci. 32, 10035-10044.
Steinmetz, M. O. (2007). Structure and thermodynamics of the tubulin-stathmininteraction. J. Struct. Biol. 158, 137-147.
Stephan, D., Sanchez-Soriano, N., Loschek, L. F., Gerhards, R., Gutmann, S.,Storchova, Z., Prokop, A. and Kadow, I. C. (2012). Drosophila Psidin regulatesolfactory neuron number and axon targeting through two distinct molecularmechanisms. J. Neurosci. 32, 16080-16094.
Stiess, M., Maghelli, N., Kapitein, L. C., Gomis-Ruth, S., Wilsch-Brauninger, M.,Hoogenraad, C. C., Tolic-Nørrelykke, I. M. and Bradke, F. (2010). Axonextension occurs independently of centrosomal microtubule nucleation. Science 327,704-707.
Stricker, J., Falzone, T. and Gardel, M. L. (2010). Mechanics of the F-actincytoskeleton. J. Biomech. 43, 9-14.
Sun, D., Leung, C. L. and Liem, R. K. (2001). Characterization of the microtubulebinding domain of microtubule actin crosslinking factor (MACF): identification of anovel group of microtubule associated proteins. J. Cell Sci. 114, 161-172.
Suozzi, K. C., Wu, X. and Fuchs, E. (2012). Spectraplakins: master orchestrators ofcytoskeletal dynamics. J. Cell Biol. 197, 465-475.
Suter, D. M. and Forscher, P. (2000). Substrate-cytoskeletal coupling as a mechanismfor the regulation of growth cone motility and guidance. J. Neurobiol. 44, 97-113.
Suter, D. M. and Miller, K. E. (2011). The emerging role of forces in axonalelongation. Prog. Neurobiol. 94, 91-101.
Swiech, L., Blazejczyk, M., Urbanska, M., Pietruszka, P., Dortland, B. R., Malik, A.
R., Wulf, P. S., Hoogenraad, C. C. and Jaworski, J. (2011). CLIP-170 and IQGAP1cooperatively regulate dendrite morphology. J. Neurosci. 31, 4555-4568.
Tanaka, E. M. and Kirschner, M. W. (1991). Microtubule behavior in the growthcones of living neurons during axon elongation. J. Cell Biol. 115, 345-363.
Journal of Cell Science 126 (11)2340
Journ
alof
Cell
Scie
nce
Tanaka, E. and Kirschner, M. W. (1995). The role of microtubules in growth coneturning at substrate boundaries. J. Cell Biol. 128, 127-137.
Tanaka, E., Ho, T. and Kirschner, M. W. (1995). The role of microtubule dynamics ingrowth cone motility and axonal growth. J. Cell Biol. 128, 139-155.
Terman, J. R. and Kashina, A. (2013). Post-translational modification and regulationof actin. Curr. Opin. Cell Biol. 25, 30-38.
Tessier-Lavigne, M. and Goodman, C. S. (1996). The molecular biology of axonguidance. Science 274, 1123-1133.
Tischfield, M. A., Cederquist, G. Y., Gupta, M. L., Jr and Engle, E. C. (2011).Phenotypic spectrum of the tubulin-related disorders and functional implications ofdisease-causing mutations. Curr. Opin. Genet. Dev. 21, 286-294.
Tsvetkov, A. S., Samsonov, A., Akhmanova, A., Galjart, N. and Popov, S. V. (2007).Microtubule-binding proteins CLASP1 and CLASP2 interact with actin filaments.Cell Motil. Cytoskeleton 64, 519-530.
Twelvetrees, A., Hendricks, A. G. and Holzbaur, E. L. F. (2012). SnapShot: axonaltransport. Cell 149, 950-950.e1.
Tymanskyj, S. R., Scales, T. M. and Gordon-Weeks, P. R. (2012). MAP1B enhancesmicrotubule assembly rates and axon extension rates in developing neurons. Mol.
Cell. Neurosci. 49, 110-119.
van der Vaart, B., Franker, M. A., Kuijpers, M., Hua, S., Bouchet, B. P., Jiang, K.,
Grigoriev, I., Hoogenraad, C. C. and Akhmanova, A. (2012). Microtubule plus-end
tracking proteins SLAIN1/2 and ch-TOG promote axonal development. J. Neurosci.
32, 14722-14728.
Van Horck, F. P. and Holt, C. E. (2008). A cytoskeletal platform for local translation in
axons. Sci. Signal. 1, pe11.
Williamson, T., Gordon-Weeks, P. R., Schachner, M. and Taylor, J. (1996).
Microtubule reorganization is obligatory for growth cone turning. Proc. Natl. Acad.
Sci. USA 93, 15221-15226.
Xu, K., Zhong, G. and Zhuang, X. (2013). Actin, Spectrin, and associated proteins
form a periodic cytoskeletal structure in axons. Science 339, 452-456.
Yang, Y., Bauer, C., Strasser, G., Wollman, R., Julien, J. P. and Fuchs, E. (1999).
Integrators of the cytoskeleton that stabilize microtubules. Cell 98, 229-238.
Young, K. G. and Kothary, R. (2007). Dystonin/Bpag1—a link to what? Cell Motil.
Cytoskeleton 64, 897-905.
Zala, D., Hinckelmann, M.-V., Yu, H., Lyra da Cunha, M. M., Liot, G., Cordelieres,
F. P., Marco, S. and Saudou, F. (2013). Vesicular glycolysis provides on-board
energy for fast axonal transport. Cell 152, 479-491.
Cytoskeletal machinery of neurons 2341
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 1
OMIM entries of cytoskeletal genes
HUMAN NEURONAL DISORDERS
1. ACTB 7 (ACTIN, BETA; 102630) - Baraitser-Winter syndrome 2 243310 (including microcephaly, intellectual disability, seizures, and hearing loss); Dystonia, juvenile-onset 607371 (with neurodegenerative traits)
2. ACTG1 (ACTIN, GAMMA 1; 102560) - Baraitser-Winter syndrome 2 (microcephaly, intellectual disability, hearing loss, anterior-to-posterior gradient lissencephaly of the pachygyria or pachygyria-band type); 614583; Deafness, autosomal dominant 20/26 604717
3. APC (ADENOMATOUS POLYPOSIS OF THE COLON; 611731) - adenomatous polyposis coli 175100; Colorectal cancer, somatic 114500; Desmoid disease, hereditary 135290; Gastric cancer, somatic 613659; Hepatoblastoma, somatic 114550 ; Brain tumor-polyposis syndrome 2
4. CEP290 (CENTROSOMAL PROTEIN 290kDa; 610142) - localises to cilia and centrosome (ATP-binding, CC, tropomyosin-like domains); Leber congenital amaurosis 10 611755 (retinal degeneration and impaired olfaction)
5. CLIP2/CLIP115 (CAP-GLY domain containing linker protein 2; 603432) - Williams-Beuren syndrome, aortic stenosis with mental retardation (WBS; 194050) - supported by a mouse model.
6. DCTN1 (DYNACTIN 1; 601143) - Perry syndrome 168605; unusual neuropsychiatric disorder with mental depression and late onset of Parkinsonism; Neuropathy, distal hereditary motor, type VIIB 607641; susceptibility to Amyotrophic lateral sclerosis (neurodegenerative disorder) 105400
7. DCX (DOUBLECORTIN; 300121) - tubulin-binding, regulator of tubulin nucleation and polymerisation; Lissencephaly, X-linked 300067; X-linked subcortical laminal heteropia/double cortex (mild lissencephaly) 300067
8. DES (DESMIN; 125660) - type III intermediate filament; neurogenic Scapuloperoneal syndrome, Kaeser type (some individuals: mild demyelinating polyneuropathy and electromyography demonstrated acute and chronic denervation in both proximal and distal muscles) 181400
9. DIAPH3 (DIAPHANOUS, DROSOPHILA, HOMOLOG OF, 3; 614567) - Auditory neuropathy, autosomal dominant, 1 609129
10. DMD (DYSTROPHIN; 300377) - Becker muscular dystrophy 300376; Cardiomyopathy, dilated, 3B 302045; Duchenne muscular dystrophy accompanied by mental retardation 310200
11. DST/BPAG1/BP240 (dystonin; 113810) - Neuropathy, hereditary sensory and autonomic, type VI 614653
12. DYNC1H1 (CYTOPLASMIC DYNEIN HEAVY CHAIN; 600112) - Charcot-Marie-Tooth disease, axonal, type 20 614228; Mental retardation, autosomal dominant 13 614563; Spinal muscular atrophy, lower extremity, autosomal dominant 158600
13. FSCN2 (FASCIN, SEA URCHIN, HOMOLOG OF, 2; 607643) - Retinitis pigmentosa (retinal degeneration) 30 607921
14. ELP1 (IKBKAP; 603722) - tubulin acetylase; Dysautonomia, familial 223900
15. ESPN (ESPIN / ECTOPLASMIC SPECIALIZATION PROTEIN; 606351) - multifunctional actin-bundling protein; Deafness, autosomal recessive 36 609006; Deafness, neurosensory, without vestibular involvement, autosomal dominant
16. FLNA (FILAMIN A, ALPHA) - actin-binding; many skeletal disorders; periventricular Heterotopia (neuronal migration defect) 300049 and 300537; Intestinal pseudoobstruction, neuronal (including abnormal intermediate signal in the peritrigonal white matter) 300048
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 2
17. GAN (GAN GENE; 605379) - putative cytoskeleton binding; Giant axonal neuropathy-1
256850
18. GFAP (GLIAL FIBRILLARY ACIDIC PROTEIN; 137780) - intermediate-filament (IF) protein that is highly specific for astroglia; Alexander disease (megalencephaly in infancy accompanied by progressive spasticity and dementia with most patients dying by 10 years) 203450
19. GSN (GELSOLIN; 137350) - Amyloidosis, Finnish type (including cranial and peripheral neuropathies) 105120
20. HDAC4 (HISTONE DEACETYLASE 4; 605314) - deacetylase, likely to act on tubulin; Brachydacytly-mental retardation syndrome 600430
21. INF2 (INVERTED FORMIN; 610982) - Charcot-Marie-Tooth disease, dominant intermediate E 614455; focal segmental glomerulosclerosis (renal disease) 613237
22. KATNAL2 (KATANIN P60 ATPASE-CONTAINING SUBUNIT A-LIKE 2; ) - autism spectrum diseases (reference)
23. KIAA0196 (STRUMPELLIN; 610657) - potential cytoskeletal regulator; Spastic paraplegia 8, autosomal dominant 603563
24. KIF1A (KINESIN FAMILY MEMBER 1A; 601255) - Mental retardation, autosomal dominant 9 614255; Neuropathy, hereditary sensory, type IIC 614213; Spastic paraplegia 30, autosomal recessive 610357
25. KIF1B (KINESIN FAMILY MEMBER 1B; 605995) - Charcot-Marie-Tooth disease, type 2A1 (hereditary motor and sensory neuropathies) 118210; Pheochromocytoma 171300; susceptibility to Neuroblastoma 1 256700; increased risk of nonsyndromic paraganglioma
26. KIF5A KINESIN or NKHC (KINESIN FAMILY MEMBER 5B, HEAVY CHAIN, NEURON-SPECIFIC; 602821) - Spastic paraplegia 10, autosomal dominant 604187
27. KIF7 (KINESIN FAMILY MEMBER 7; 611254) - Acrocallosal syndrome/Joubert syndrome 12 (mental retardation syndrome with brain abnormalities) 200990
28. KIF11 or KNSL1 or KHC (KINESIN FAMILY MEMBER 11, KINESIN-LIKE 1; KINESIN HEAVY CHAIN; 148760) - Microcephaly with or without chorioretinopathy, lymphedema, or mental retardation 152950
29. LIS1 (LISSENCEPHALY 1; 607432; official name: PLATELET-ACTIVATING FACTOR ACETYLHYDROLASE, ISOFORM 1B, ALPHA SUBUNIT; PAFAH1B1; 601545) - lissencephaly 1 607432; subcortical laminar heterotopia 607432
30. MAPT (MICROTUBULE-ASSOCIATED PROTEIN TAU; 157140) - Frontotemporal Dementia with or without parkinsonism (600274); Pick disease 172700; Supranuclear palsy, progressive 601104; Supranuclear palsy, progressive atypical 260540
31. MYH9 (MYOSIN IIA HEAVY CHAIN 9; 160775) - Deafness, autosomal dominant 17 603622; Epstein syndrome 153650; Fechtner syndrome 153640: Macrothrombocytopenia and progressive sensorineural deafness 600208; May-Hegglin anomaly 155100; Sebastian syndrome 605249
32. MYO1A (MYOSIN IA; 607841) - Deafness, autosomal dominant 48 607841
33. MYO5A (MYOSIN VA; MYO5; MYOXIN; MYOSIN HEAVY CHAIN 12; 160777) - Griscelli syndrome, type 1 214450 (hypomelanosis with a primary neurologic deficit including motor development delay and mental retardation)
34. MYO6 (MYOSIN VI, 600970) - nonsyndromic deafness (hearing loss without related signs and symptoms affecting other parts of the body) called DFNB37, likely disrupting stereocilia
35. MYO7A (MYOSIN VIIA; 276903) - Deafness, autosomal dominant 11 601317; Deafness, autosomal recessive 2 600060; Usher syndrome, type 1B (hearing loss or deafness and progressive vision loss) 276900
36. MYO15A (MYOSIN XVA; 602666) - Deafness, autosomal recessive 3 600316
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 3
37. NDE1 (NUDE NUCLEAR DISTRIBUTION E HOMOLOG 1; 609449) - part of a multiprotein
complex that regulates dynein function; Lissencephaly 4 (with microcephaly) 614019
38. NEFH (NEUROFILAMENT, HEAVY POLYPEPTIDE; 162230) - susceptibility to Amyotrophic lateral sclerosis 105400
39. NEFL (NEUROFILAMENT, LIGHT POLYPEPTIDE; 162280) - Charcot-Marie-Tooth disease, type 1F 607734; Charcot-Marie-Tooth disease, type 2E 607684
40. PRPH (PERIPHERIN; 170710) - type III intermediate filament; susceptibility to Amyotrophic lateral sclerosis (neurodegenerative disorder characterized by the death of motor neurons) 105400
41. RDX (RADIXIN; 179410) - Deafness, autosomal recessive 24 611022 (autosomal recessive sensorineural deafness)
42. SPAST (SPASTIN; 604277) - Spastic paraplegia 4, autosomal dominant 182601
43. TUBA1A (ALPHA 1 TUBULIN) - Lissencephaly 3 611603
44. TUBA8 (ALPHA 8 TUBULIN) - Polymicrogyria with optic nerve hypoplasia 613180
45. TUBB3 (ß3 TUBULIN; 602661) - Fibrosis of extraocular muscles, congenital, 3A (600638); Cortical dysplasia, complex, with other brain malformations (CDCBM) 614039, a more severe disorder of aberrant neuronal migration and disturbed axonal guidance resulting in mental retardation.
46. TUBB2B (ß2 TUBULIN; 612850) - asymmetric Polymicrogyria 610031; malformation of cortical development characterized by an excessive number of small gyri with abnormal lamination
47. TRIOBP (TRIO and F-actin binding protein; 609761) - organization of actin filaments; Deafness, autosomal recessive 28 609823
48. SPAST (SPASTIN; 604277) - MT severer; Spastic paraplegia 4, autosomal dominant 182601
49. SPTAN1 (SPECTRIN, ALPHA, NONERYTHROCYTIC 1; 182810) - Epileptic encephalopathy, early infantile, 5 613477
50. SPTBN2 (SPECTRIN, BETA, NONERYTHROCYTIC, 2; 604985) - Spinocerebellar ataxia 5 600224
51. TUBGCP6 (TUBULIN-GAMMA COMPLEX-ASSOCIATED PROTEIN 6; 610053) - Microcephaly and chorioretinopathy with or without mental retardation 251270
ANIMAL MODELS OF NEURONAL DISORDERS (where no human brain disorder reported)
52. CFL1 (COFILIN 1; 601442) - deacetylase, likely to act on tubulin; mouse model with morphological abnormalities of cortex
53. DBN1 (DREBRIN E; 126660) - age-related significant decrease in postsynaptic drebrin in a transgenic mouse model
54. DPYSL2 or DRP2 or CRMP2 / COLLAPSIN RESPONSE MEDIATOR PROTEIN 2 (DIHYDROPYRIMIDINASE-LIKE 2; 602463) - tubulin dimer binding - rat model for inflammatory and neuropathic hypersensitivity
55. HDAC1 (HISTONE DEACETYLASE 1; 601241) - mouse model: Deletion of both Hdac1 and Hdac2 led to major abnormalities of cortical, hippocampal, and cerebellar development, whereas deletion of either individually had no apparent effect on neuronal development.
56. SPTBN4 (SPECTRIN, BETA, NONERYTHROCYTIC, 4; QUIVERING; 606214) - autosomal recessive mouse mutation 'quivering' (qv) produces progressive ataxia with hindlimb paralysis, deafness, and tremor
57. STMN1 (STATHMIN 1; 151442) - increased amounts of MTs in brain and behavioural deficits in mouse models (memory, parental care, fear)
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 4
58. TBCE (TUBULIN-SPECIFIC CHAPERONE E; 604934) - Hypoparathyroidism-retardation-
dysmorphism syndrome (skeleton) 241410; Kenny-Caffey syndrome-1 (skeleton) 244460; mouse model: progressive caudiocranial motor axon degeneration from the age of 2 weeks
59. UTRN (UTROPHIN; 128240) - mice deficient for both dystrophin and utrophin pheoncopy DMD in humans, including severe progressive muscular dystrophy resulting in premature death, ultrastructural myotendinous and neuromuscular junction abnormalities
NON-NEURONAL HUMAN DISORDERS
60. ACTA1 (ACTIN, ALPHA 1, skeletal muscle; 102610) - different myopathies: Myopathy, actin, congenital, with excess of thin myofilaments 161800; Myopathy, congenital, with fiber-type disproportion 1 255310; Myopathy, nemaline, 3 161800
61. ACTA2 (ACTIN, ALPHA 2, smooth muscle, aorta; 102620) - Aortic aneurysm, familial thoracic 6 611788; Moyamoya disease 5 614042; Multisystemic smooth muscle dysfunction syndrome 613834
62. ACTC1 (ACTIN, ALPHA, CARDIAC MUSCLE; 102540) - Atrial septal defect 5 612794; Cardiomyopathy, dilated, 1R 613424; Cardiomyopathy, familial hypertrophic, 11 612098; Left ventricular noncompaction 4 613424
63. ACTN2 (ACTININ, ALPHA-2; 102573) - Cardiomyopathy, dilated, 1AA 612158
64. ACTN4 (ACTININ, ALPHA-4; 604638) - Glomerulosclerosis, focal segmental, 1 603278
65. {Hypertension, essential, salt-sensitive} 145500
66. ANK2 (ANKYRIN 2, neuronal) - heart disorders
67. BFSP2 (BEADED FILAMENT STRUCTURAL PROTEIN 2; 603212) - intermediate filament; Cataract, autosomal dominant, multiple types 1 611597; Cataract, congenital 604219; Cataract, juvenile-onset 604219
68. CFL2 (COFILIN 2; 601443) - Nemaline myopathy 7 (NEM7; 610687)
69. CTNNB1 (CATENIN, BETA-1; 116806) - Hepatocellular carcinoma 114550; Ovarian cancer 167000; Pilomatricoma 132600
70. DNAAF3 (DYNEIN, AXONEMAL, ASSEMBLY FACTOR 3; 614566) - Ciliary dyskinesia, primary, 2 606763
71. DNAH5 (DYNEIN, AXONEMAL, HEAVY CHAIN 5) - Ciliary dyskinesia, primary, 3, with or without situs inversus 608644 (chronic respiratory tract infections, abnormally positioned internal organs, infertility)
72. DNAH11 (DYNEIN, AXONEMAL, HEAVY CHAIN 11; 603339) - Ciliary dyskinesia, primary, 7, with or without situs inversus 611884
73. DNAI1 (DYNEIN, AXONEMAL, INTERMEDIATE CHAIN 1; 604366) - Ciliary dyskinesia, primary, 1, with or without situs inversus 244400
74. DNAI2 (DYNEIN, AXONEMAL, INTERMEDIATE CHAIN 2; 605483) - Ciliary dyskinesia, primary, 9, with or without situs inversus 612444
75. DSP (DESMOPLAKIN; 125647) - Arrhythmogenic right ventricular dysplasia 8 607450; Dilated cardiomyopathy with woolly hair and keratoderma 605676; Epidermolysis bullosa, lethal acantholytic 609638; Keratosis palmoplantaris striata II 612908; Skin fragility-woolly hair syndrome 607655
76. DYNC2H1 (DYNEIN, CYTOPLASMIC 2, HEAVY CHAIN 1; 603297) - skeletal and early developmental defects: Asphyxiating thoracic dystrophy 3 613091; Short rib-polydactyly syndrome, type II, digenic 263520; Short rib-polydactyly syndrome, type III 263510
77. FLNB (filamin B, beta; 603381) - actin-binding; skeletal disorders: Atelosteogenesis, type I 108720; Atelosteogenesis, type III 108721; Boomerang dysplasia 112310; Larsen syndrome 150250; Spondylocarpotarsal synostosis syndrome 272460
78. FLNC (FILAMIN C; FLN2; 102565) - Myopathy, distal, 4 614065; Myopathy, myofibrillar, 5 609524)
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 5
79. GSN (GELSOLIN; 137350) - Amyloidosis, Finnish type 105120 (lattice corneal dystrophy type
II)
80. JUP (JUNCTION PLAKOGLOBIN, PKGB, GAMMA CATENIN, DESMOPLAKIN III; 173325) - Arrhythmogenic right ventricular dysplasia 12 611528; Naxos disease 601214
81. KIF21A (KINESIN FAMILY MEMBER 21A) - Fibrosis of extraocular muscles, congenital, 1 135700; Fibrosis of extraocular muscles, congenital, 3B 135700
82. KIF22 or KNSL4 or OBP (KINESIN FAMILY MEMBER 22, KINESIN-LIKE 4, ORIP-BINDING PROTEIN; 603213) - Spondyloepimetaphyseal dysplasia with joint laxity, type 2 603546; skeletal disorder without intellectual impairment
83. MYBPC1 (MYOSIN BINDING PROTEIN C, slow type; 160794) - Arthrogryposis, distal, type 1B 614335
84. MYH2/2A/S2A (MYOSIN, HEAVY CHAIN 2/2A/S2A, SKELETAL MUSCLE, ADULT; 160740) - Inclusion body myopathy-3 605637
85. MYH3 (MYOSIN, HEAVY CHAIN 3, skeletal muscle, embryonic; 160720) - Arthrogryposis, distal, type 2B (muscular) 601680
86. MYH7 (MYOSIN, HEAVY CHAIN 7, cardiac muscle, beta; 160760) - myopathies
87. MYH11 or SMMHC (MYOSIN, HEAVY CHAIN 11, SMOOTH MUSCLE; SMOOTH MUSCLE MYOSIN HEAVY CHAIN; 160745) - Aortic aneurysm, familial thoracic 4 132900
88. MYL2 or RLC (MYOSIN, LIGHT CHAIN 2, REGULATORY, CARDIAC, SLOW; REGULATORY LIGHT CHAIN OF MYOSIN; 160781) - Cardiomyopathy, familial hypertrophic, 10 608758
89. MYL3 or MLC1SB/1V (MYOSIN, LIGHT CHAIN 3, ALKALI, VENTRICULAR, SKELETAL, SLOW; MYOSIN, LIGHT CHAIN 1, SLOW, B/VENTRICULAR; 160790) - Cardiomyopathy, familial hypertrophic, 8 608751
90. NEM3 (NEMALINE MYOPATHY 3; 161800) - Myopathy, nemaline, 3 161800; Myopathy, actin, congenital, with excess of thin myofilaments 161800
91. NF2 (NEUROFIBROMIN 2, merlin/Moesin-Ezrin-Radixin-Like Protein; 607379) - Meningioma, NF2-related, somatic 607174; Neurofibromatosis, type 2 101000; Schwannomatosis 162091
92. PCNT (PERICENTRIN; 605925) - Microcephalic osteodysplastic primordial dwarfism, type II 210720
93. PLEC1 (PLECTIN 1; 601282) - Epidermolysis bullosa simplex with pyloric atresia 612138; Epidermolysis bullosa simplex, Ogna type 131950; Muscular dystrophy with epidermolysis bullosa simplex 226670; Muscular dystrophy, limb-girdle, type 2Q 613723
94. PKP1 (PLAKOPHILIN 1; 601975) - Ectodermal dysplasia/skin fragility syndrome 604536
95. PKP2 (PLAKOPHILIN 2; 602861) - Arrhythmogenic right ventricular dysplasia 9 609040
96. SPTA1 (SPECTRIN, ALPHA, ERYTHROCYTIC 1; 182860) - Elliptocytosis-2 130600; Pyropoikilocytosis 266140; Spherocytosis, type 3 270970
97. TNNI2 (TROPONIN I, FAST-TWITCH SKELETAL MUSCLE ISOFORM; 191043) - binds to actin in thin myofilaments; Arthrogryposis multiplex congenital, distal, type 2B 601680
98. TNNI3 (TROPONIN I, CARDIAC; 191044) - binds to actin in thin myofilaments; Cardiomyopathy, dilated, 1FF 613286; Cardiomyopathy, dilated, 2A 611880; Cardiomyopathy, familial hypertrophic, 7 613690; Cardiomyopathy, familial restrictive 115210
99. TPM1 (TROPOMYOSIN 1; 191010) - Cardiomyopathy, dilated, 1Y 611878; Cardiomyopathy, familial hypertrophic, 3 115196
100. TPM2 (TROPOMYOSIN 2; 190990) - Arthrogryposis multiplex congenita, distal, type 1/2B 108120 601680; Nemaline myopathy 609285
101. TPM3 (TROPOMYOSIN 3; 191030) - CAP myopathy 609284; Myopathy, congenital, with fiber-type disproportion 255310; Nemaline myopathy 1, autosomal dominant 609284
102. VCL (VINCULIN; 193065) - Cardiomyopathy, dilated, 1W 611407; Cardiomyopathy, familial hypertrophic, 15 613255
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 6
103. WAS(P) (WISKOTT-ALDRICH SYNDROME PROTEIN; 300392) - various immuodeficiencies:
Neutropenia, severe congenital, X-linked 300299; Thrombocytopenia, X-linked 313900; Thrombocytopenia, X-linked, intermittent 313900; Wiskott-Aldrich syndrome 301000
NO REPORTED HUMAN DISORDER LINKS
104. ACTL6B (ACTIN-LIKE 6B; ACTIN-RELATED PROTEIN, ALPHA; 612458)
105. ACTIN, KAPPA (611266)
106. ACTN1 (ACTININ, ALPHA-1; 102575)
107. ACTN3 (ACTININ, ALPHA-3; 102574) - potential link to Sprinting performance
108. ACTR2 (ARP2; 604221) - Arp2/3 complex component
109. ACTR3 (ARP3; 604222) - Arp2/3 complex component
110. ADD2 (ADDUCIN 2; 102681)
111. ADD3 (ADDUCIN 3; 601568)
112. ARPC1A (ARP2/3 COMPLEX, SUBUNIT 1A; SOP2L; 604220)
113. ARPC1B (ARP2/3 COMPLEX, SUBUNIT 1B; ARC41; 604223)
114. ARPC2 (ARP2/3 COMPLEX, SUBUNIT 2; ARC34; 604224)
115. ARPC3 (ARP2/3 COMPLEX, SUBUNIT 3; ARC21; 604225)
116. ARPC4 (ARP2/3 COMPLEX, SUBUNIT 4; ARC20; 604226)
117. ARPC5 (ARP2/3 COMPLEX, SUBUNIT 5; ARC16; 604227)
118. CAMSAP1 (CALMODULIN-REGULATED SPECTRIN-ASSOCIATED PROTEIN 1; Ssp4; 613774) - MT minus end stabiliser
119. CAPG (CAPPING PROTEIN, GELSOLIN-LIKE; 153615)
120. CAPZA1 (CAPPING PROTEIN, ALPHA-1; 601580)
121. CAPZA2 (CAPPING PROTEIN, ALPHA-2; 601571)
122. CAPZA3 (CAPPING PROTEIN, ALPHA-3; 608722)
123. CAPZB (CAPPING PROTEIN, BETA; 601572)
124. CKAP5 (CYTOSKELETON-ASSOCIATED PROTEIN 5, CHTOG, XMAP215; 611142)
125. CLASP1 (CYTOPLASMIC LINKER-ASSOCIATED PROTEIN 1; 605852)
126. CLASP2 (605853)
127. CLIP1/CLIP170/RESTIN/RSN (179838) - weak links to Hodgkin disease/lymphoma
128. CORO1A (CORONIN 1A; 605000) - actin binding; mouse models with lymphocyte migration defects
129. CORO1B (CORONIN 1B; 609849) - actin binding
130. CORO1C (CORONIN 1C; 605269) - actin binding
131. CORO2A (CORONIN 2A; 602159) - actin binding
132. CORO2B (CORONIN 2B; 609849) - actin binding
133. CORO7 (CORONIN 7; 611668) - actin binding
134. CTTN (CORTACTIN; 164765) - actin binder and regulator
135. DAAM1 (DISHEVELLED-ASSOCIATED ACTIVATOR OF MORPHOGENESIS 1; 606626)
136. DAAM2 (DISHEVELLED-ASSOCIATED ACTIVATOR OF MORPHOGENESIS 2; 606627)
137. DBNL (DREBRIN-LIKE; 610106)
138. DNAH6 (DYNEIN, AXONEMAL, HEAVY CHAIN 6; 603336)
139. DNAH9 (DYNEIN, AXONEMAL, HEAVY CHAIN 9; 603330)
140. DNAL1 (DYNEIN, AXONEMAL, LIGHT CHAIN 1; 610062) - Ciliary dyskinesia, primary, 16 614017
141. DNALI1 (DYNEIN, AXONEMAL, LIGHT INTERMEDIATE POLYPEPTIDE 1; 602135)
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 7
142. DSTN (DESTRIN/ACTIN DEPOLYMERIZING FACTOR; ADF; 609114) - mouse model with
irregular thickening of the corneal epithelium
143. DYNC1I2 (DYNEIN, CYTOPLASMIC 1, INTERMEDIATE CHAIN 2; 603331)
144. DYNC1I1 (DYNEIN, CYTOPLASMIC 1, INTERMEDIATE CHAIN 1; 603772)
145. DYNC1LI2 (DYNEIN, CYTOPLASMIC 1, LIGHT INTERMEDIATE CHAIN 2; 611406)
146. DYNLL1 (DYNEIN, LIGHT CHAIN, LC8 TYPE, 1; 601562)
147. DYNLL2 (DYNEIN, LIGHT CHAIN, LC8 TYPE, 2; 608942)
148. DYNLRB1 (DYNEIN, LIGHT CHAIN, ROADBLOCK TYPE, 1; 607167)
149. DYNLRB2 (DYNEIN, LIGHT CHAIN, ROADBLOCK TYPE, 2; 607168)
150. DYNLT1 (DYNEIN, LIGHT CHAIN, TCTEX TYPE, 1; 601554)
151. DYNLT3 (DYNEIN, LIGHT CHAIN, TCTEX TYPE, 3; 300302)
152. EB1 (MAPRE1/MICROTUBULE-ASSOCIATED PROTEIN, RP/EB FAMILY, MEMBER 1; 603108)
153. EB2 (MAPRE2; 605789)
154. EB3 (MAPRE3; 605788)
155. ELP3 (ELONGATION PROTEIN 3, S. CEREVISIAE, HOMOLOG OF; 612722) - tubulin acetylase
156. ENAH (ENAH enabled homolog; 609061)
157. EVPL (ENVOPLAKIN; 601590)
158. EZRIN/VIL2 (VILLIN 2; 123900) - mouse model with intestinal epithelial defects
159. FSCN1 (FASCIN, SEA URCHIN, HOMOLOG OF, 1; 602689)
160. HDAC5 (HISTONE DEACETYLASE 5; 300269) - likely tubulin deacetylase
161. HDAC8 (HISTONE DEACETYLASE 8; 300269) - likely tubulin deacetylase
162. PHF21A (PHD FINGER PROTEIN 21A/BRAF35/KIAA1696; 605315) - HDAC COMPLEX, 80-KD SUBUNIT, likely tubulin deacetylase
163. IQGAP1 (IQ MOTIF-CONTAINING GTPase-ACTIVATING PROTEIN 1; 603379)
164. IQGAP2 (IQ MOTIF-CONTAINING GTPase-ACTIVATING PROTEIN 2; 605401)
165. KATNA1 (KATANIN, p60 SUBUNIT, A1; 606696)
166. KATNAL1 (KATANIN p60 SUBUNIT A-LIKE 1; 614764)
167. KATNAL2 (KATANIN, p60 SUBUNIT, A-LIKE PROTEIN 2; 614697)
168. KATNB1 (KATANIN, p80 SUBUNIT, B1; 602703)
169. KIF2C or MCAK or KNSL6 (KINESIN FAMILY MEMBER 2C, MITOTIC CENTROMERE-ASSOCIATED KINESIN, KINESIN-LIKE 6; 604538) - +TIP and inhibitor of MT polymerisation
170. KIF5B (KINESIN FAMILY MEMBER 5B; 602809) - kif5B-/- mice were embryonic lethal with a severe growth retardation at 9.5 to 11.5 days postcoitum
171. KIF5C or NKHC2 (KINESIN FAMILY MEMBER 5C, NEURON-SPECIFIC KINESIN HEAVY CHAIN 2; 604593)
172. KIF20A or RABKINESIN 6 (KINESIN FAMILY MEMBER 20A; 605664) - frequently deleted in myeloid leukemias
173. KIF23 or KNSL5 or MKLP1 (KINESIN FAMILY MEMBER 23, KINESIN-LIKE 5; MITOTIC KINESIN-LIKE 1; 605064)
174. KLC1 or KLC or KNS2 (KINESIN LIGHT CHAIN 1, KINESIN 2; 600025) - mouse Klc1 predominantly in central and peripheral neuronal tissues
175. MACF1 (MICROTUBULE-ACTIN CROSS-LINKING FACTOR 1; 608271)
176. MAP1A (MICROTUBULE-ASSOCIATED PROTEIN 1A; 600178)
177. MAP1B (MICROTUBULE-ASSOCIATED PROTEIN 1B; 157129)
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 8
178. MAP2 (MICROTUBULE-ASSOCIATED PROTEIN 2; 157130) - some indirect effect in Jnk1
(MAPK8; 601158)-null mouse
179. MSN (MOESIN; 309845) - actin-binding
180. MYH10 (MYOSIN, HEAVY CHAIN 10, NONMUSCLE; 160776) - homozygous null mice have 70% fewer, but larger, myocytes
181. MYL4 (MYOSIN, LIGHT CHAIN 4, ALKALI, ATRIAL, EMBRYONIC; 160770)
182. NINL (NINEIN-LIKE PROTEIN; 609580)
183. PARD3 (PARTITIONING-DEFECTIVE PROTEIN 3; 606745) - MT binding in neuronal polarisation
184. PFN1 (PROFILIN 1; 176610) - mouse model with immune response deficits
185. PFN2 (PROFILIN 2; 176590)
186. PFN2 (PROFILIN 2; 612812)
187. PLS1 (PLASTIN; FIMBRIN, 602734)
188. PKP3 (PLAKOPHILIN 3; 605561)
189. PKP4 (PLAKOPHILIN 4; 604276)
190. PPL (PERIPLAKIN; 602871)
191. PPP1R9A (PROTEIN PHOSPHATASE 1, REGULATORY SUBUNIT 9A; SPINOPHILIN; NEURABIN I; 602468) - actin binding
192. SPTB (SPECTRIN, BETA, ERYTHROCYTIC; 182870) - Anemia, neonatal hemolytic, fatal and near-fatal; Elliptocytosis-3; Spherocytosis, type 2
193. SPTBN1 (SPECTRIN, BETA, NONERYTHROCYTIC, 1; 182790)
194. SPTBN5 (SPECTRIN, BETA, NONERYTHROCYTIC, 5; 605916)
195. STMN2 or SCGN10 / SUPERIOR CERVICAL GANGLIA, NEURAL SPECIFIC 10 (STATHMIN-LIKE 2; 600621) - upregulated in familial dysautonomia (Cheishvili et al., 2011, Hum Mol Genet 20, 1585ff.)
196. STMN3 (STATHMIN-LIKE 3; 608362) - tubulin sequestering
197. TLN1 (TALIN 1; 186745)
198. TLN2 (TALIN 2; 607349)
199. TMOD1 (TROPOMODULIN; 190930)
200. TMOD2 (TROPOMODULIN 2; 602928)
201. TMOD3 (TROPOMODULIN 3; 605112)
202. TMOD4 (TROPOMODULIN 4; 605834)
203. TMSB4X (THYMOSIN, BETA-4, X CHROMOSOME; 300159) - actin monomer sequestering protein
204. TNNI1 (TROPONIN I, SLOW-TWITCH SKELETAL MUSCLE ISOFORM; 191042) - binds to actin in thin myofilaments
205. TPM4 (TROPOMYOSIN 4; 600317)
206. TUBE1 (TUBULIN, EPSILON-1; 607345)
207. TUBG1 (TUBULIN, GAMMA-1; 191135)
208. TUBG2 (TUBULIN, GAMMA-2; 605785)
209. TUBGCP4 (TUBULIN-GAMMA COMPLEX-ASSOCIATED PROTEIN 4; 609610)
210. TUBGCP5 (TUBULIN-GAMMA COMPLEX-ASSOCIATED PROTEIN 5; 608147)
211. TWF1 (TWINFILIN, DROSOPHILA, HOMOLOG OF, 1; 610932) - G-actin binding
212. associated with sensory neuropathy (SPTLC1, SPTLC2, WNK1, IKBKAP, NTRK1, NGFB, DNMT1
213. VASP (VASODILATOR-STIMULATED PHOSPHOPROTEIN; 601703)
214. WASL (WISKOTT-ALDRICH SYNDROME GENE-LIKE; 605056)
A. Prokop et al. - Understanding cytoskeletal machinery of neurons 9
215. WASF1 (WISKOTT-ALDRICH SYNDROME PROTEIN FAMILY, MEMBER 1, WAVE1,
SCAR1; 605035)
216. WASF2 (WISKOTT-ALDRICH SYNDROME PROTEIN FAMILY, MEMBER 2, WAVE2, SCAR2; 605875)
217. WASF3 (WISKOTT-ALDRICH SYNDROME PROTEIN FAMILY, MEMBER 3, WAVE3, SCAR3; 605068)
Related Documents