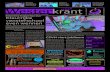
Autoradiogram and its interpretive drawing of a replicating E. coli chromosome. Page 1137 3 H-thymidine

Figure 30-2Autoradiogram and its interpretive drawing of a replicating E. coli chromosome. Page 1137 3 H-thymidine.
Dec 19, 2015
Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

Figure 30-2 Autoradiogram and its interpretive drawing of a replicating E. coli chromosome.
Pag
e 11
37
3H-thymidine

Figure 30-6 Electron micrograph of a replication eye in Drosophila melanogaster DNA.
Pag
e 11
39

Figure 30-3 Replication of DNA.
Pag
e 11
37

How would YOU go about determining the mechanism of
DNA replication?????
What would a geneticist do?
What would a biochemist do?

Figure 5-31 Action of DNA polymerases.
Pag
e 99
DNA polymerases assemble incoming deoxynucleoside triphosphates on single-stranded DNA templates such that the growing strand is elongated in its 5’ 3’ direction.

Table 30-1Properties of E. coli DNA Polymerases.
Pag
e 11
45

DNA PolymerasesEnzymes that replicate DNA using a DNA template (as opposed to enzymes that synthesize DNA using an RNA template --reverse transcriptases—and enzymes that make DNA without a template--terminal transferases). Most organisms have more than one type of DNA polymerase (E. coli has five DNA polymerases). For all:
1. Polymerization occurs only 5' to 3'2. Polymerization requires a template to copy: the complementary strand.3. Polymerization requires 4 dNTPs: dATP, dGTP, dCTP, dTTP (TTP is sometimes not designated with a 'd' since there is no ribose containing equivalent)4. Polymerization requires a pre-existing primer from which to extend. The primer is RNA in most organisms, but it can be DNA in some organisms; very rarely the primer is a protein in the case of certain viruses only.


Figure 30-10 Schematic diagram for
the nucleotidyl transferase
mechanism of DNA
polymerases.
A and B are coordinated Me+2.
A activates primer’s 3’-OH for inline nucleophilic attack on incoming dNTP’s α-phosphate.
B orients and stabilizes the triphosphate.

Figure 30-7 Priming of DNA synthesis by short RNA
segments.
Pag
e 11
39

Figure 5-32a Replication of duplex DNA in E. coli.
Pag
e 10
0

Figure 5-32b Replication of duplex DNA in E. coli.
Pag
e 10
0
Animation


Figure 30-28The replication of E. coli DNA.

DNA Polymerase I from E. coli was the first DNA polymerase characterized. approximately 400 molecules of the enzyme per cell. E. coli DNA polymerase I is abbreviated pol I. a single large protein with a molecular weight of approximately 103 kDa (103,000 grams per mole). a divalent cation (Mg++) for activity
Three enzymatic activities:1. 5'-to-3' DNA Polymerase activity2. 3'-to-5' exonuclease (Proofreading activity)3. 5'-to-3' exonuclease (Nick translation activity)
It is possible to remove the 5'-to-3' exonuclease activity using a protease to cut DNA pol I into two protein fragments
Both the polymerization and 3'-to-5' exonuclease activities are on the large Klenow fragment of DNA pol I, and the 5'-to-3' exonuclease activity is on the small fragment.

Like all known DNA polymerases, DNA polymerase I requires a primer from which to initiate replication and polymerizes deoxyribonucleotides into DNA in the 5' to 3' direction using the complementary strand as a template. Newly synthesized DNA is covalently attached to the primer, but only hydrogen-bonded to the template. The template provides the specificity according to Watson-Crick base pairing. Only the alpha phosphate of the dNTP is incorporated into newly synthesized DNA

Figure 30-8b X-Ray structure of E. coli DNA polymerase I Klenow fragment (KF) in complex with a dsDNA (a tube-and-arrow representation of the complex in the same orientation as Part a).
Pag
e 11
41


Figure 30-12 Nick translation as catalyzed by Pol I.
Pag
e 11
44

Figure 30-8aX-Ray
structure of E. coli DNA
polymerase I Klenow
fragment (KF) in complex with
a dsDNA.
Pag
e 11
41

Here’s a computer modelhttp://www.youtube.com/watch?v=4jtmOZaIvS0
Overview of DNA and replication
http://207.207.4.198/pub/flash/24/menu.swf
Another one with review questions
http://www.wiley.com/college/pratt/0471393878/student/animations/dna_replication/index.html
This is a pretty good outline:http://www.youtube.com/watch?v=teV62zrm2P0&NR=1


DNA Pol III holoenzyme.

Figure 30-13b The subunit of E. coli Pol III holoenzyme. Space-filling model of sliding clamp in hypothetical complex with B-
DNA.
Pag
e 11
46

http://www.callutheran.edu/Academic_Programs/Departments/BioDev/omm/poliiib_2/poliiib.htm
Sliding clamp

Figure 30-20The reactions
catalyzed by E. coli DNA ligase.
Pag
e 11
50

Figure 30-21
X-Ray structure of DNA ligase from
Thermus filiformis.
Pag
e 11
51
Related Documents