Yeast & Cloning Sergio Peisajovich Lim Lab June 2007

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

Yeast & Cloning
Sergio Peisajovich
Lim Lab June 2007

Experimental LabWhy Yeast?
The yeast Saccharomyces cerevisiae (also called “baker’s yeast”) is probably the ideal eukaryotic microorganism for biological studies.
Yeast genome: fully sequenced and easy to manipulate.
Basic mechanisms of yeast cell biology (such as DNA replication, recombination, cell division and metabolism) are highly similar to that of higher organisms (including humans).

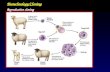
Experimental LabYeast Life Cycle

Experimental LabYeast: Ideal Platform for Synthetic Biology
Add parts, devices or even modules (in an “extra-genomic” format -plasmid-based- or “inte grating” them within the yeast genome. Delete specific yeast genes, to r emove “background” or interference. Add “reporter genes” to monitor in real time the function of the synthetic parts/devices/modules under study.
Life cycle fa st enough so that we could do all these genetic manipulations in a reasonable amount of time.

Parts/Devices/Modules are built in bacteria
Empty initial plasmid
Plasmid coding the desired
device
Transform into Yeast
Experimental LabYeast: Adding parts… in plasmids

Experimental LabYeast: Adding parts… in plasmids
growth in selective medium

Experimental LabYeast: Adding parts… in plasmids
growth in selective medium

Experimental LabYeast: Adding parts… into the genome
Homologous recombination allows genomic integration, but we still need to select:

Experimental LabYeast: Adding parts… into the genome
Part/Device/Module
URA3
plasmid
Digest with specific restriction enzyme
Part/Device/Module
plasmid
Linear DNA, ready for yeast transformation and integration
Part/Device/Module
URA3*
HomologousRecombination
YeastChromosome
Incoming Linear DNA
URA3* URA3Part/Device/Module
Integration(Note that 2 copies, one defective and one functional, of the marker
are generated)
YeastChromosome

Experimental LabYeast: Adding parts… into the genome
URA3
plasmid
URA3
PCR product
Linear DNA, ready for yeast transformation
and integration
yfg
HomologousRecombination
YeastChromosome
URA3
Integration(yfg is now disrupted)
YeastChromosome
URA3

Part 1
plasmidplasmid
Part 1
plasmidplasmid
Part 2
Part 1
plasmidplasmid
Part 2 Part 3
Experimental LabCombinatorial Cloning

A B B C C D
A D
A D
Experimental LabCombinatorial Cloning
Based on Type IIsrestriction enzymes

A B B C C D
A D
A D A D
Combinatorial Cloning
Experimental Lab
CombinatorialLibraries


Experimental LabSynthetic Biology as Engineering Engineering Negative
Feedback Loops
Negative Effectors to be used:OspF (MAPK Phosphothreonine Lyase) YopJ (MAPKK Ser/Thr acetylase)YopH (MAPK Tyr phosphatase)
Promoters to be used:Constitutive expression (Adhp, CycIp, Ste5p)Inducible by pathway activation (STLp, Fig1p)
Protein-interaction domains:Leucine Zippers (high and medium affinities,some with degradation motif)
Prom Tag Effector Zipper Term

Experimental LabSynthetic Biology as Engineering Engineering Negative
Feedback Loops
1- Combinatorial Cloning in Bacteria
2- Transfer Constructs into Yeast
3- Analyze Pathway Behavior

Experimental LabSynthetic Biology as Engineering Engineering Negative
Feedback Loops
1- Combinatorial Cloning in Bacteria
DONORS ACCEPTORS

Prom Tag Effector Zipper Term
Experimental LabSynthetic Biology as Engineering Engineering Negative
Feedback Loops
1- Combinatorial Cloning in Bacteria

Experimental LabSynthetic Biology as Engineering Engineering Negative
Feedback Loops
2- Transfer Constructs into Yeast
3- Analyze Pathway Behavior
FACS
Microscopy
Related Documents