Systematic approaches to central nervous system myelin Patricia de Monasterio-Schrader 1 , Olaf Jahn 2,3 , Stefan Tenzer 4 , Sven P. Wichert 1 , Julia Patzig 1 , and Hauke B. Werner 1* This is the author’s version as accepted for publication. The copyrighted e-offprint is available from Springer Basel AG, as published in Cellular and Molecular Life Sciences (2012), Digital Object Identifier (DOI) 10.1007/s00018-012-0958-9. Authors addresses 1 Department of Neurogenetics Max Planck Institute of Experimental Medicine, Göttingen, Germany 2 Proteomics Group Max Planck Institute of Experimental Medicine, Göttingen, Germany 3 DFG Research Center for Molecular Physiology of the Brain Göttingen, Germany 4 Institute of Immunology University Medical Center of the Johannes Gutenberg University Mainz, Germany * Corresponding author: Dr. Hauke Werner Max Planck Institute of Experimental Medicine Department of Neurogenetics Hermann-Rein-Str. 3 D-37075 Göttingen, Germany Tel.: +49 (551) 389-9759 Fax.: +49 (551) 389-9758 E-mail: [email protected] Running title Systematic approaches to CNS myelin Manuscript organization 22 pages, 4 figures, 2 tables, 1 supplemental table Acknowledgements We thank W. Möbius for providing the electron micrograph in Figure 1, J. M. Edgar for critical reading of the manuscript, C. M. Kassmann for discussion and K.-A. Nave for continuous support. ST is supported by the Deutsche Forschungsgemeinschaft (SFB 490 Z3) and the Forschungszentrum Immunologie (FZI) at the University of Mainz, and HBW is supported by the BMBF (DLR-Leukonet) and the European Commission (FP7-LeukoTreat).

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
Systematic approaches to central nervous system myelin
Patricia de Monasterio-Schrader1, Olaf Jahn2,3, Stefan Tenzer4, Sven P. Wichert1, Julia Patzig1, and Hauke B. Werner1*
This is the author’s version as accepted for publication. The copyrighted e-offprint is available from Springer Basel AG, as published in Cellular and Molecular Life Sciences (2012), Digital Object Identifier (DOI) 10.1007/s00018-012-0958-9. Authors addresses 1 Department of Neurogenetics
Max Planck Institute of Experimental Medicine, Göttingen, Germany 2 Proteomics Group
Max Planck Institute of Experimental Medicine, Göttingen, Germany 3 DFG Research Center for Molecular Physiology of the Brain Göttingen, Germany 4 Institute of Immunology University Medical Center of the Johannes Gutenberg University Mainz, Germany * Corresponding author: Dr. Hauke Werner Max Planck Institute of Experimental Medicine Department of Neurogenetics Hermann-Rein-Str. 3 D-37075 Göttingen, Germany Tel.: +49 (551) 389-9759 Fax.: +49 (551) 389-9758 E-mail: [email protected] Running title Systematic approaches to CNS myelin Manuscript organization 22 pages, 4 figures, 2 tables, 1 supplemental table Acknowledgements We thank W. Möbius for providing the electron micrograph in Figure 1, J. M. Edgar for critical reading of the manuscript, C. M. Kassmann for discussion and K.-A. Nave for continuous support. ST is supported by the Deutsche Forschungsgemeinschaft (SFB 490 Z3) and the Forschungszentrum Immunologie (FZI) at the University of Mainz, and HBW is supported by the BMBF (DLR-Leukonet) and the European Commission (FP7-LeukoTreat).
Abstract
Rapid signal propagation along vertebrate axons is facilitated by their insulation with myelin, a
plasma membrane specialization of glial cells. The recent application of ‘omics’ approaches to
the myelinating cells of the central nervous system, oligodendrocytes, revealed their mRNA
signatures, enhanced our understanding of how myelination is regulated and established that
the protein composition of myelin is much more complex than previously thought. This
review provides a meta-analysis of the >1200 proteins thus far identified by mass
spectrometry in biochemically purified central nervous system myelin. Contaminating proteins
are surprisingly infrequent according to bioinformatic prediction of subcellular localization and
comparison with the transcriptional profile of oligodendrocytes. The integration of datasets
also allowed the subcategorization of the myelin proteome into functional groups comprising
genes that are coregulated during oligodendroglial differentiation. An unexpectedly large
number of myelin-related genes cause - when mutated in humans - hereditary diseases
affecting the physiology of the white matter. Systematic approaches to oligodendrocytes
and myelin thus provide valuable resources for the molecular dissection of developmental
myelination, glia-axonal interactions, leukodystrophies and demyelinating diseases.
Keywords
Oligodendrocyte; myelin; differentiation; proteome; transcriptome; leukodystrophy;
schizophrenia; multiple sclerosis
Abbreviations
CNS, central nervous system
Ig-CAM, immunoglobulin-like cell adhesion molecule
MBP, myelin basic protein
mOL, myelinating oligodendrocyte
MS, mass spectrometry / mass spectrometric
OPC, oligodendrocyte precursor cell
P0, myelin protein zero
PLP, proteolipid protein
PNS, peripheral nervous system
Introduction
Fast signal propagation along vertebrate axons requires the restriction of action potentials to
short axonal segments termed the nodes of Ranvier. This is achieved by the insulation of the
long segments between the nodes (‘internodes’) with a sleeve of membranous structure
termed myelin [1, 2]. Myelin is a specialization of the plasma membrane of glial cells, which is
organized as multiple, spiral and compacted layers (Fig. 1A). The non-compacted adaxonal
(innermost) myelin layer is connected to the glial cell body through cytoplasmic channels,
which contain cytoskeleton, vesicles and outposts of endoplasmic reticulum and Golgi
apparatus [3] and is considered relevant for the maintenance and turnover of myelin.
Schwann cell-derived myelin of the peripheral nervous system (PNS) differs from
oligodendrocyte-derived myelin of the central nervous system (CNS) regarding ontogenic
origin (subventricular zone versus neural crest), number of myelinated axonal segments per
glial cell (1:1 versus up to 60:1), and major proteins (the Ig-CAM myelin protein zero, (P0,
official gene name Mpz) versus the tetraspan proteolipid protein (PLP, Plp1)) (for review see
[4]).
Many aspects of myelin biogenesis, maintenance and metabolism, and of the function of
oligodendrocytes in preventing myelinated axons from degeneration [5, 6] are not well
understood at the molecular level. In this review, we focus on systematic analyses aimed at a
molecular understanding of CNS myelin. First we - briefly - consider systematic analyses at
the level of mRNA, miRNA and DNA, which have recently provided a wealth of information
regarding the mechanisms that regulate myelination and remyelination. Secondly, we review –
more extensively – current advancements in the analysis of the protein composition of myelin
itself. The addition of proteomic techniques to the methodological spectrum applied in myelin
research has accelerated our knowledge of which proteins are actually myelin constituents,
thereby providing a versatile basis for the systematic analysis of their functions in the
physiology and pathophysiology of myelinated fiber tracts. We have recently reviewed myelin
proteome analyses with a focus on technical [7] and disease-related [8] aspects. Here we
provide a meta-analysis of CNS myelin protein composition on the basis of unbiased protein
identification by mass spectrometry (MS) and the bioinformatic integration with other
systematic approaches such as mRNA microarray analysis.
Regulators of myelination identified by systematic mRNA, miRNA and DNA
analyses
Profiling of mRNA abundance during the differentiation of oligodendrocyte progenitor cells
(OPC) to post-mitotic, pre-myelinating oligodendrocytes (OL) and myelinating
oligodendrocytes (mOL), led to the identification of extensive changes affecting gene
clusters involved in the cell cycle, cell motility, adhesion, cytoskeletal remodeling, lipid
metabolism and ultimately myelination [9-12]. Filtering of such datasets allowed the
elucidation of functions as diverse as the regulation of OPC migration by semaphorins and
neuropilins [13], the transcriptional control of oligodendrocyte maturation and myelination
by ‘myelin gene regulatory factor‘ (Mrf, also termed gene model 98, Gm98) [14] and the
modulation of OPC differentiation and remyelination involving the nuclear ‘retinoid X receptor
gamma‘ (Rxrg) [15]. Additionally, novel proteins, such as an oligodendrocytic cell adhesion
molecule termed ‘proline-rich membrane protein‘ (PRMP/TMEM58, official gene name Shisa4)
[11] and ‘oligodendrocytic myelin paranodal and inner loop protein‘ (Opalin, also termed
transmembrane protein 10, TMEM10) [16, 17] have been identified.
Importantly, the expression of genes in oligodendrocytes is not only controlled by
transcriptional [18, 19], but also by post-transcriptional and post-translational modifications.
For example, oligodendroglial differentiation [20] coincides with alterations in the abundance
of regulatory RNAs [21-24] including miR-219, which represses proteins that both, promote
OPC proliferation and negatively regulate maturation. The regulation of oligodendrocyte
differentiation and myelination by epigenetic factors, including histone modifications, DNA
methylation and microRNAs, has been extensively reviewed [25-32], also in the context of
demyelinating disease [32-40]. Collectively, the studies discussed in these overview articles
founded the concept that the final step of oligodendroglial differentiation necessitates not
only transcriptional activation of myelin-related genes but also the concurrent de-repression
of various inhibitors of myelination [41, 42]. Taken together, systematic analyses at the level
of mRNA, miRNA and DNA have recently provided many important insights into regulatory
networks in oligodendrocytes, and more generally in their physiology and pathophysiology.
However, since not all mRNAs expressed in oligodendrocytes encode myelin proteins and not
all myelin proteins are signified by the same mRNA abundance profile, studies at the
nucleotide level are not well suited to systematically assess the protein content of myelin.
Towards the myelin proteome
In the early 1970s William Norton and Shirley Poduslo introduced a method to biochemically
enrich myelin from homogenized nervous tissue utilizing sucrose gradient centrifugation
[43], with myelin accumulating as low-density membranous material at the interface between
0.32 M and 0.85 M sucrose. As the method has proven to efficiently yield highly pure myelin,
only moderate modifications have been introduced since. These include the use of a Polytron
to substitute the Dounce homogenizer [44], a supplementary second density gradient
centrifugation that reduces axonal contaminations [45] and the addition of protease
inhibitors [46]. About the same time, the belief has formed that the number and variety of
myelin proteins is very low. This view emerged because upon one-dimensional gel-separation
of purified myelin and subsequent protein staining only very few bands were visible. These
are mainly constituted by PLP, myelin basic protein (Mbp) and 2’,3’-cyclic nucleotide
phosphodiesterase (Cnp1, previously termed ‘Wolfgram’s protein’) [47-51]. However, these
highly abundant proteins overshadowed low-abundance proteins that did not constitute
visible bands in early studies. Since then, the use of gradient-gels, two-dimensional gels, and
modified buffers and detergents has enhanced the migration of myelin into the gels and its
separation. Additionally, the sensitivity of modern staining procedures based on silver or
colloidal Coomassie is superior to the previously used Sudan Black and Fast Green. Together,
these improvements have allowed the visualization of many more bands (Fig. 1B), reflecting
that the protein composition of myelin is much more complex than previously anticipated.
To date, elaborated MS technology has turned out to be the most sensitive method to
systematically identify the protein constituents of myelin. While only one study of the PNS
myelin proteome is available so far [52], eight proteomic studies of CNS myelin have been
published, providing 11 datasets for myelin from mouse [7, 53-56], rat [57, 58] and human
[56, 59]. The approaches taken differ regarding details of sample preparation,
protein/peptide separation, MS instrumentation, and data analysis, including stringency
criteria for database search and acceptance of a protein hit. These differences
notwithstanding, we have compiled and compared these datasets ‘as published’ for the
present meta-analysis.
A compendium of mass spectrometrically identified myelin proteins
The integration of all myelin proteome datasets yielded a compendium of 1280 proteins,
among them 1261 for which it was possible to determine unique gene identifiers
(Supplemental Tab. S1). The number of proteins identified per approach markedly increased
in the more recent studies (Fig. 2A), reflecting advancements in proteomic methodology in
general and of MS instrumentation in particular. To measure the reproducibility of MS-based
myelin protein analyses we have determined which protein in any given dataset has
additionally been identified in at least one other approach. Reassuringly, more than half of all
proteins (52%) were present in at least two datasets (Fig. 2B), raising confidence in their
association with CNS myelin. The rate of reproduced identifications was almost 100% among
the earlier approaches (in which fewer than 150 proteins were identified) and well above
60% among the more recent approaches (with over 350 identified proteins) (Fig. 2C).
Importantly, the only exception is the single dataset in which - instead of conventionally
purified myelin - a remyelination-inhibiting subfraction was analyzed [58]. In that study, the
extensive prefractionation by ion exchange chromatography and multi-dimensional gel
electrophoresis led to a distinct sub-proteome, which is not directly comparable to that of
the complete myelin fraction. Together, this supports the view that the profile of myelin-
associated proteins as determined by sequential sucrose gradient centrifugation for myelin
purification and proteome analysis has been very robust across different laboratories,
irrespective of differences regarding the analyzed species, proteomic approach or MS
technology. We expect that further progress in MS performance will further increase the
coverage and the confidence level of the myelin proteome.
Interestingly, the only protein identified in all 11 approaches is not a ‘classical’ myelin protein
but a molecular chaperone termed heat shock 70kDa protein 8 (Hspa8; alternative name
constitutive heat shock protein, Hsc70). HSPA8 functions as a chaperone for MBP [60, 61]
and may thus be co-transported into myelin by molecular association. Five proteins were
each identified in 10 approaches, namely MBP, CNP, sirtuin 2 (SIRT2), septin 8 (SEPT8) and
dihydropyrimidinase-like 2 (DPYSL2, also termed collapsin response mediator, CRMP2) [53,
62-65]. Importantly, there was no simple correlation between the frequency of identification
and the abundance of a protein (Tab. 1). For example, the absence of transmembrane
proteins (such as PLP) among those most frequently identified was expected and is due to
the bias implemented by the use of conventional 2D gel electrophoresis in three approaches
[53-55]. This method comprises isoelectric focusing (IEF) as the first dimension of
separation, in which proteins migrate along an immobilized pH gradient until they reach their
isoelectric point (IP). However, hydrophobic membrane proteins tend to precipitate during
IEF, which has been attributed to the fact that proteins are least soluble at their IP while
potent ionic detergents such as SDS are not compatible with IEF. Together, this prevents the
transfer of membrane proteins to the second dimension SDS-PAGE [66], and thus their
identification.
Coinciding with the enhanced sensitivity of current MS instrumentation, the number of
identified contaminants from different cellular origins has increased. For example, proteomic
compendia of synaptic vesicles [67] or mammalian brain mitochondria [68] comprise
proteins derived from contaminating myelin, such as the myelin-specific proteins PLP, MBP,
myelin-associated glycoprotein (MAG) and myelin-oligodendrocyte basic protein (MOBP). Vice
versa, the most common contaminants of the myelin-enriched fraction are mitochondria and
synaptic vesicles, which can be attributed to similar floatation properties in sucrose or Percoll
gradients. Therefore it is not surprising that a considerable number of entries in the myelin
protein compendium may qualify as synaptic or mitochondrial according to prior knowledge.
However, information on subcellular localization, as assembled in large protein databases,
may be only partially true or even incorrect. For example, information for a particular protein
may have been assumed for an isoenzyme or another protein from the same biochemical
cascade without further validation. Moreover, many proteins supposed to be specific to
mitochondria actually do have a dual localization [68]. Thus, particular proteins may indeed
be present in both mitochondria and myelin, though antibody-based validation is lacking. We
also note that some proteins, such as neural cell adhesion molecule 1 (Ncam1), neurofascin
(Nfasc), and contactin 2 (Cntn2, also termed transient axonal glycoprotein, TAG-1) reside in
both, axonal membranes and (adaxonal) myelin [69-71]. In consequence, we have not
excluded proteins from the present compendium based on anticipated knowledge of
subcellular or cellular localization other than myelin.
The integration of the myelin proteome with other systematically gained information
circumvents some of the limitations of prior knowledge. For example, transcriptomic datasets
can clarify cellular origins and developmental mRNA abundance profiles (see below).
Bioinformatic approaches cannot only predict subcellular localizations, but also group
proteins according to structural criteria such as the presence of particular domains. For the
present compendium we have systematically predicted mitochondrial proteins utilizing Cello
(cello.life.nctu.edu.tw), TargetP (cbs.dtu.dk/services/TargetP) and Wolfpsort (wolfpsort.org)
and transmembrane proteins with the algorithms TMHMM2 v2.0
(cbs.dtu.dk/services/TMHMM), Phobius (phobius.sbc.su.se) and TMpred
(ch.embnet.org/software/TMPRED_form.html) (Supplemental Tab. S1). The numbers of
transmembrane and suspected mitochondrial proteins are indicated in Fig. 2A.
Cross-species comparison
The protein composition of myelin is considered highly similar among mammalian species [72,
73]. However, the cross-species comparison of MS-identified CNS myelin proteins of human
versus rodent origin (Fig. 3A) showed a surprisingly moderate overlap of only approximately
50%. Likely, the limited overlap reflects species-specific variations of amino acid sequences
affecting the proteolytic peptides rather than the existence of proteins exclusive to myelin of
one mammalian species. This view is supported by the finding that several proteins that upon
MS-analysis seemed specific to either human or rodent myelin were subsequently also
identified in the respective other species by immunoblot [56].
Comparison of CNS with PNS myelin
The proportion of proteins identified in both peripheral and central myelin is unexpectedly
high when the PNS myelin proteome [52] is compared with the present compendium (Fig.
3B). Proteins present in both types of myelin are likely relevant for functions or
morphological features that unify myelin in both the CNS and the PNS. For example, MBP,
which is considered relevant for the intracellular compaction of myelin membranes, is equally
abundant in both, CNS and PNS myelin. On the other hand, claudin 11 (Cldn11, also termed
oligodendrocyte specific protein, OSP) and periaxin (Prx) define molecular specializations of
the radial component particular to CNS myelin [74] and of the PNS-specific bands of Cajal
[75], respectively. Consequently, claudin 11 and periaxin were identified exclusively in CNS or
PNS myelin, respectively. This indicates that molecules specific to either type of myelin may
signify particular functions or morphological specializations. The relative abundance of
selected myelin proteins according to quantification by MS [7, 52] is compiled in Tab 1.
Comparison with transcriptomics data indicates that at least 90% of the
myelin-associated proteins are indeed robustly expressed in oligodendrocytes
The proteins identified in purified myelin by MS may include contaminants of a different
cellular origin that may have co-purified with myelin. As the individual validation of so many
proteins using antibody-based techniques is not feasible, we sought systematic information
regarding which proteins are actually expressed in oligodendrocytes. For this purpose we
have exploited a previously established transcriptional profile of oligodendrocytes, which
were isolated at distinct differentiation stages (PDGFRα+/MOG- OPC; GalC+ OL; MOG+/NG2-
mOL) by the team around John Cahoy and Ben Barres [9]. By filtering that dataset for the
proteins identified by MS in purified myelin, we have gained a bioinformatic comparison of the
myelin proteome and the oligodendrocyte transcriptome (Fig. 4, Supplemental Tab. S1). An
unambiguous correlation was possible for 1249 proteins and the corresponding mRNA.
For 79 proteins, no corresponding oligonucleotides were present on the arrays. For a further
106 proteins, corresponding mRNA was not detected in oligodendrocytes. This group may
include false-negatives for which the oligonucleotides on the arrays are not suitable, but also
proteins indeed not expressed in oligodendrocytes but co-purified with CNS myelin. The latter
category probably includes marker proteins of peripheral myelin such as P0 (Mpz), ‘peripheral
myelin protein 2‘ (Pmp2) [76] and ‘discs large homolog 1‘ (Dlg1) [77]. The identification of
these proteins in a very small subset of CNS myelin analyses may be attributed to myelin of
peripheral nerves inadvertently attached to the brains used as the starting material for
myelin purification.
Most importantly, robust mRNA abundance in oligodendrocytes at either differentiation stage
was found for at least 1064 (>90%) of the 1170 proteins identified in CNS myelin for which
such information was available. This suggests that contamination of the myelin-enriched brain
fraction by material from different cellular origins was fairly moderate.
Abundance profi les of mRNAs encoding CNS myelin proteins
To determine groups of myelin-associated genes signified by co-regulated expression, we
performed k-means cluster analysis for those 1064 mRNAs that were robustly detected in
oligodendrocytes. 631 mRNAs displayed significant abundance changes during
oligodendroglial differentiation (Fig. 4).
The majority of ‘classical’ myelin proteins clustered together with a significant mRNA-
abundance increase coinciding with the differentiation from OPCs to post-mitotic
oligodendrocytes (cluster ‘Early-UP’ in Fig. 4). These include Plp1 [4], Mbp [62], Cnp1 [64],
Sirt2 [53], Mag [78], Mobp [79], Cldn11 [74], Nfasc [70], myelin and lymphocyte protein
(Mal) [80], prominin 1 (Prom1) [81], the myelin-associated inhibitor reticulon 4 (Rtn4, Nogo)
[82], tubulin beta 4 (Tubb4) [83] and band 4.1 protein G (Epb4.1l2) [84]. A small cluster
(Up-DOWN in Fig. 4) comprised mRNAs with a temporarily increased abundance at the onset
of myelination such as the non-receptor tyrosine kinase fyn, which is involved in the
maturation of oligodendrocytes [85]. A third cluster of mRNAs that display a continuous
abundance increase during the differentiation from OPC to OL to mOL (cluster ‘Ascending’ in
Fig. 4) comprised the myelin marker carbonic anhydrase 2 (Car2) [86], Opalin [16, 17],
Cd82 [87], Rab3a [88], myelin oligodendrocyte glycoprotein (Mog) [89], integrin-associated
signal transducer (Cd47) [90], ermin (Ermn) [91], fatty acid synthase (Fasn) [92],
tetraspanin 2 (Tspan2) [93] and vesicle-associated membrane protein 3 (Vamp3) [94],
among many other proteins. It will be a task of future analyses to determine their role in
myelin biology.
Enhanced expression at the onset of myelination has often been considered circumstantial
evidence of a protein being a ‘myelin protein’. On the other hand, developmental repression
appears equally relevant. This is illustrated by the effect of polysialic acid (PSA), a
posttranslational modification of neural cell adhesion molecule (NCAM). Diminishment of
polysialylation coincides with the differentiation from OPC to mOL [69] and is a prerequisite
for efficient myelin biogenesis [95]. Thus, the early expression of both glial and axonal [96]
PSA-NCAM is inhibitory for premature myelination. Interestingly, the abundance of Ncam1
mRNA in oligodendrocytes is also repressed during the differentiation from OPC to mOL
(cluster ‘Descending’ in Fig. 4).
The 18kDa membrane-tetraspan peripheral myelin protein (PMP22) has long been assumed to
be absent from CNS myelin but was recently detected in purified CNS myelin by immunoblot
[97], though at a low level. Of note, identification of PMP22 by MS as in one approach [59] is
difficult by conventional proteomic approaches based on digestion with trypsin because its
cleavage sites are atypically distributed (leading to an unusually large 10 kDa fragment not
suitable for MS sequencing), and 65% of its amino acid residues constitute transmembrane
domains (leading to membrane-spanning peptides usually not detected by MS). Importantly,
by microarray analysis and in situ-hybridization, Pmp22 mRNA was detected in
oligodendrocytes [9, 21], in which its abundance decreases coinciding with the
differentiation from OPC to OL (‘Early-DOWN’ in Fig. 4), involving repression by miR-9 [21].
It is of note that developmental repression occurs frequently in oligodendrocytes: Among all
myelin-associated proteins, clusters with generally repressed abundance of the corresponding
mRNA (‘Early-DOWN’, ‘Descending’, ‘Late-DOWN’) comprise about as many genes as those
with increased mRNA abundance (‘Early-UP’, ‘Ascending’, ‘Late-UP’).
The abundance of 433 mRNAs encoding myelin-associated proteins, such as integrin beta 1
(Itgb1) [98], cell division cycle 42 (Cdc42), ras-related C3 botulinum toxin substrate 1
(Rac1) [99] as well as glyceraldehyde-3-phosphate dehydrogenase (Gapdh), was unchanged
during oligodendrocyte differentiation. Taken together, all clusters comprised previously
known myelin-related genes, supporting the notion that myelin proteins are not necessarily
signified by a particular developmental mRNA abundance profile.
Protein groups defined by homology or structural similarity
The systematic identification of protein groups with homology may be beneficial for
functional analysis. For example, by bioinformatic analysis of the compendium, those myelin-
associated proteins can be systematically identified that contain a FERM-domain - i.e. a
sequence motif with homology to four-point-one (4.1)-proteins, ezrin, radixin and moesin -
for molecular interactions (Epb41l1/4.1N, Epb41l2/4.1G, Epb41l3/4.1B, Ezr, Rdx, Msn, Tln1,
Tln2), or immunoglobulin-like domains that signify cell adhesion molecules (Ig-CAMs) (Alcam,
Bsg, Cadm2/NECL3, Cadm3/NECL1, Cadm4/NECL4, Cntn1, Cntn2/TAG1, Cntnap1/CASPR1,
Cntnap2/CASPR2, Dscaml1, Hepacam/GlialCAM, Igh, Igk, Igl, Igsf8, Itgb1, Itgb8, L1cam, Mag,
Mog, Mpzl1, Ncam1, Ncam2, Nrcam).
To further illustrate such systematic identification we have filtered tetraspan-transmembrane
proteins upon predicting the transmembrane domains for all proteins in the compendium
(Supplemental Tab. S1). Tetraspans are particularly abundant in myelin and have a proven
relevance for its biogenesis and pathology [100]. Tetraspan-encoding mRNAs with increased
abundance during maturation include Cd82, which restricts OPC migration [87] (Cluster
‘Ascending’ in Fig. 4), and those that determine the structure of myelin at the intraperiod
lines (Plp1) [4], radial component (Cldn11) [74] and paranodes (Mal) [80] (Cluster ‘Early-UP‘
in Fig. 4). The functions of other myelin tetraspans with ‘Ascending‘ mRNA profiles (in Fig.
4) may be related but remain to be established, including those of Tspan2, CKLF-like
MARVEL transmembrane domain containing 5 (Cmtm5), synaptogyrin 2 (Syngr2) and
transmembrane protein 205 (Tmem205). On the other hand, the repression of Pmp22 in
differentiating oligodendrocytes (see above) may be attributed to its capacity to repress
cellular growth and to form cellular junctions [101-104]. Similarly, the developmental
repression of synaptogyrin 3 (Syngr3, cluster ‘Early-DOWN‘), Cd151, glycoprotein M6A
(Gpm6a) (‘Descending‘) and Cd9 (‘Late-DOWN‘) may reflect functions of these tetraspans
not required during, or even inhibiting oligodendrocyte differentiation. Conversely, the
abundance of several tetraspan-encoding mRNAs (Pllp, Cd81, Connexin 43 (Gja1),
glycoprotein M6B (Gpm6b), secretory carrier membrane proteins 1 and 5 (Scamp1,
Scamp5)) is developmentally unchanged, suggesting continued requirement of the gene
product.
Notably, the membrane of vesicular exosomes derived from multivesicular bodies [105, 106]
comprises a particular abundance of tetraspans, in which they may facilitate fission and
fusion [107]. Oligodendrocyte-derived exosomes may be required to locally dispose off
superfluous membrane and to transfer material to axons [108] and microglia [109].
However, they also counteract the extension of oligodendroglial plasma membrane, at least in
vitro [110], possibly reflecting a function in the spatial segregation of myelin sheaths in vivo
related to that of Nogo (Rtn4) [82, 111]. Strikingly, there is a high overlap between the
tetraspans of exosomes and of myelin, including PLP, CD9, CD81, CD82 and CD63
(tetraspanin 30). It is attractive to speculate on commonalities of the two compartments
regarding biosynthesis and function. Together, the integration of proteomic and
transcriptomic datasets provides a background to study protein groups with structural and
functional similarity and co-regulated expression.
Heritable myelin-related diseases
Hereditary diseases affecting myelination or the physiology of the white matter
(leukodystrophies or leukoencephalopathies) vary considerably regarding the affected genes
and the pathophysiology. Importantly, only a subset of the causative genes encodes myelin
proteins [112-117]. For example mutations affecting the PLP1-gene, which encodes the
most abundant protein of CNS myelin, PLP, cause the hypomyelinating leukodystrophy
‘Pelizaus-Merzbacher disease‘ or the allelic ‘spastic paraplegia type 2‘ [118-120].
As oligodendrocytes, astrocytes, microglia and neurons intimately interact, the cellular
pathologies of leukodystrophies are very complex. Indeed, the identification of novel
leukodystrophy disease genes, which has accelerated in recent years [121-125], has
facilitated - often unexpected - insights into the biology and the interactions of glial cells
[126]. Importantly, many leukodystrophies are caused by mutations that do not affect
‘classical‘ myelin genes, or even genes considered to be expressed specifically in cells other
than oligodendrocytes. This is exemplified by ‘hypomyelination and congenital cataract‘
(HCC), which is caused by mutations affecting the FAM126A gene [127] encoding the
primarily neuronal protein hyccin [128], or by ‘hereditary diffuse leukoencephalopathy with
spheroids‘ (HDLS), which is caused by mutations affecting the gene encoding ‘colony
stimulating factor 1 receptor‘ (CSF1R) [125], considered exclusively expressed in microglia
[129]. However, such prior knowledge may infer an unjustified bias in the search for the
pathomechanism. For example, astrocytes are commonly considered the primary site of
pathology in ‘Alexander’s disease‘. Here, the white matter degeneration coincides with the
emergence of aggregates (termed Rosenthal fibers) in astrocytes. Rosenthal fibers comprise
the product of the causative gene, which encodes the intermediate filament glial fibrillary
acidic protein (Gfap) [130, 131]. However, whether astrocytic Rosenthal fibers indeed
contribute to the emergence of myelin abnormalities has not been satisfactorily shown.
Strikingly, GFAP is commonly thought to be exclusive to astrocytes, while our meta-analysis
emphasizes that GFAP was identified by MS in purified myelin and that the corresponding
mRNA was detected in oligodendrocytes (cluster ‘ASCENDING‘; Fig. 4). This suggests that
the expression of GFAP is less restricted than anticipated, and that it ought not be excluded
that oligodendrocytes and myelin are primary sites of pathology in Alexander’s disease.
In Table 2 we have compiled a list of genes that fulfill three criteria: (1) the protein was
identified by MS in myelin, (2) the mRNA was robustly detected in oligodendrocytes, and (3)
mutations affecting the corresponding human gene cause a disease that includes pathology
of myelin or the white matter, at least in a subset of patients. For example, one of the
causative genes for megalencephalic leukoencephalopathy with subcortical cysts (MLC), glial
cell adhesion molecule (GlialCAM, official gene name Hepacam; Cluster Down-UP in Fig. 4),
encodes an abundant CNS myelin protein [132] but is also expressed in astrocytes [124,
133]. In the latter, GlialCAM is involved in the intracellular trafficking of MLC1, a
transmembrane protein with distant homology to sodium channels, to its normal localization
at the junctions between astrocytes and neighboring astrocytes or the endothelial cells of
the blood-brain-barrier. As MLC1 is also a causative gene in MLC [134], a failure of GlialCAM-
dependent trafficking of MLC1 to astrocytic junctions is very likely disease relevant.
However, the emergence of myelin vacuoles in a subset of MLC patients may potentially be
attributed to the presence of GlialCAM in normal CNS myelin.
Taken together, oligodendrocytes and myelin may well contribute to the pathogenesis in
white matter diseases in which the affected genes are erroneously thought to be expressed
mainly or exclusively in other cells. However, proof for the involvement of particular cell
types in the pathogenesis of any leukodystrophy must come from the analysis of cell-type
specific mutant mice, which is also prerequisite for the development of rational therapy
concepts. We propose that the present compendium of myelin proteins also provides a useful
resource to identify causative genes in association studies in which only chromosomal
segments (comprising many genes) are currently known.
Tools and perspectives
Until recently, the constituents of myelin were mainly approached by single gene analysis.
However, with the advent of ‘omics’ techniques it became evident that all myelin proteins are
embedded in a context of molecular networks involving co-regulated expression and physical
protein-protein interactions. The present meta-analysis integrates systematic information
gained by proteomic analysis of normal CNS myelin in 11 published datasets and by
transcriptional profiling of differentiating oligodendrocytes upon immunopanning, i.e. cell
purification using antibodies directed against stage-specific surface antigens [9].
In an alternative approach to obtain samples for transcriptional profiling of distinct cell types,
BAC-transgenic mice were generated in which cell-type specific promoters drive the
expression of the ribosomal protein L10a with an EGFP-tag suited to affinity-purify labeled
polysomes for the subsequent analysis of the associated RNA (‘translating ribosome affinity
purification‘, TRAP) [135]. For the oligodendrocyte lineage, the Olig2 promoter (active from
OPCs to mOL) and the Cmtm5 promoter (active in mOL) were used [136]. Interestingly,
CMTM5 is a proven constituent of peripheral myelin [52], while antibody-based validation as
an oligodendroglial protein is yet lacking. However, its occurrence in the present compendium
(Cluster ‘Ascending‘ in Fig. 4) suggests that CMTM5 is a myelin protein also in the CNS.
Thus, the TRAP study has identified over 1000 probes representing hundreds of mRNAs with
a high probability of being translated in oligodendrocytes, which is also supported by the
considerable number of known myelin-related genes in the dataset (supplemental table S5 in
[136]. Considering that transgene expression under control of the Cmtm5 promoter was
comparatively weak, the future variation of TRAP utilizing a stronger oligodendrocyte-specific
promoter may allow the complementation of immunopanning for future transcriptional
profiling approaches in comparative analyses of mouse models of myelin-related diseases.
However, the application of either technique may remain limited, e.g. when oligodendroglial
surface antigens or the activity of driver-promoters are altered as part of the pathology.
Large-scale in situ-hybridization as supplied in the ‘Allen Brain Atlas‘ (mouse.brain-map.org)
and subsequent sorting of labeling patterns (Supplemental Table 11 in [137] by the time of
publication allowed the identification of 79 mRNAs with a high probability of oligodendrocyte-
enriched expression. Reassuringly, 37 (47%) among them are also represented in the current
myelin proteome compendium. Many of the remaining oligodendrocyte-enriched mRNAs
encode enzymes of the lipid metabolism (Abca2, Edg8, Fa2h, Fabp5, Hmgcs1, Lass2, Npc1,
Ugt8) and known oligodendroglial transcription factors (Olig1, Olig2, Mrf/Gm98, Sox10).
A more direct strategy to identify transcription factors in the oligodendrocyte lineage
involved in silico-screening [138] of a previously established transcription factor expression
atlas based on in situ-hybridization in the developing mouse CNS (‘Mahoney atlas‘) [139].
Out of 1445 transcription factors in the pictorial, 87 displayed an embryonic mRNA labeling
pattern compatible with expression in glial progenitors, and 20 displayed sustained glia-like
labeling at birth. Among those, 8 were already known to be involved in glial development
while 9 were newly identified as enriched in OPCs. One of them, the HMG-box transcription
factor 7-like 2 (Tcf7l2), was shown to be functionally involved in OPC maturation [138, 140,
141]. Together, the exploitation of pictorials allows the identification of molecules relevant
for oligodendrocytes, and thus is promising also beyond the application to transcription
factors. However, the interpretation of expression atlas data, their validation and their
integration with other systematic datasets [142-147] remains a challenge.
In conclusion, a wealth of systematically gained molecular information has recently emerged
for the normal development of oligodendrocytes and their accompanying non-myelinating
cells. As of today, the exploitation of these resources is still in its infancy, and the field is
confronted with the luxury problem to choose the most interesting candidate proteins for
individual functional analysis. We believe that the integration of systematic datasets - as
illustrated here - can facilitate the selection of proteins for in-depth follow-up studies. The
current technical limitations of systematic approaches, including non-represented genes and
unsuited probes (affecting microarrays and pictorials), non-specific antibodies
(immunohistochemistry) and proteins unsuited for MS sequencing upon trypsin digest
(proteomics), may well be overcome, e.g. by whole-transcriptome sequencing, more specific
antibodies and alternative proteases, respectively. Rather, the systematic application beyond
normal tissue is laborious and expensive and may thus remain limited at last. However,
disease-relevant insights into the pathophysiology of the white matter eventually require
comprehensive knowledge of the spatio-temporal expression of all mRNAs, regulatory RNAs
and proteins, not only in the normal brain but also in disease models and ultimately in
patients. While there are obvious limitations to the availability of human brain samples,
techniques for differential analyses of models of myelin disease have been established at the
proteomic [53, 148] and transcriptomic [14, 149, 150] level. It is encouraging that also the
application to complex traits in humans, such as multiple sclerosis [151-157] and psychiatric
diseases [158-162], has been initiated.
References
1. Simons, M. and J. Trotter, Wrapping it up: the cell biology of myelination. Curr Opin Neurobiol, 2007. 17(5): p. 533-40.
2. Baron, W. and D. Hoekstra, On the biogenesis of myelin membranes: sorting, trafficking and cell polarity. FEBS Lett, 2010. 584(9): p. 1760-70.
3. Perkins, G.A., et al., Electron tomographic analysis of cytoskeletal cross-bridges in the paranodal region of the node of Ranvier in peripheral nerves. J Struct Biol, 2008. 161(3): p. 469-80.
4. Mobius, W., et al., Phylogeny of proteolipid proteins: divergence, constraints, and the evolution of novel functions in myelination and neuroprotection. Neuron Glia Biol, 2008. 4(2): p. 111-27.
5. Nave, K.A., Myelination and the trophic support of long axons. Nat Rev Neurosci, 2010. 11(4): p. 275-83.
6. Nave, K.A. and B.D. Trapp, Axon-glial signaling and the glial support of axon function. Annu Rev Neurosci, 2008. 31: p. 535-61.
7. Jahn, O., S. Tenzer, and H.B. Werner, Myelin proteomics: molecular anatomy of an insulating sheath. Mol Neurobiol, 2009. 40(1): p. 55-72.
8. Werner, H.B. and O. Jahn, Myelin matters: proteomic insights into white matter disorders. Expert Rev Proteomics, 2010. 7(2): p. 159-64.
9. Cahoy, J.D., et al., A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci, 2008. 28(1): p. 264-78.
10. Dugas, J.C., et al., Functional genomic analysis of oligodendrocyte differentiation. J Neurosci, 2006. 26(43): p. 10967-83.
11. Nielsen, J.A., et al., Identification of a novel oligodendrocyte cell adhesion protein using gene expression profiling. J Neurosci, 2006. 26(39): p. 9881-91.
12. Lin, G., et al., Neonatal and adult O4(+) oligodendrocyte lineage cells display different growth factor responses and different gene expression patterns. J Neurosci Res, 2009. 87(15): p. 3390-402.
13. Cohen, R.I., et al., A role for semaphorins and neuropilins in oligodendrocyte guidance. J Neurochem, 2003. 85(5): p. 1262-78.
14. Emery, B., et al., Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell, 2009. 138(1): p. 172-85.
15. Huang, J.K., et al., Retinoid X receptor gamma signaling accelerates CNS remyelination. Nat Neurosci, 2011. 14(1): p. 45-53.
16. Golan, N., et al., Identification of Tmem10/Opalin as an oligodendrocyte enriched gene using expression profiling combined with genetic cell ablation. Glia, 2008. 56(11): p. 1176-86.
17. Kippert, A., et al., Identification of Tmem10/Opalin as a novel marker for oligodendrocytes using gene expression profiling. BMC Neurosci, 2008. 9: p. 40.
18. Nicolay, D.J., J.R. Doucette, and A.J. Nazarali, Transcriptional control of oligodendrogenesis. Glia, 2007. 55(13): p. 1287-99.
19. Wegner, M., A matter of identity: transcriptional control in oligodendrocytes. J Mol Neurosci, 2008. 35(1): p. 3-12.
20. Rosenberg, S.S., B.L. Powell, and J.R. Chan, Receiving mixed signals: uncoupling oligodendrocyte differentiation and myelination. Cell Mol Life Sci, 2007. 64(23): p. 3059-68.
21. Lau, P., et al., Identification of dynamically regulated microRNA and mRNA networks in developing oligodendrocytes. J Neurosci, 2008. 28(45): p. 11720-30.
22. Budde, H., et al., Control of oligodendroglial cell number by the miR-17-92 cluster. Development, 2010. 137(13): p. 2127-32.
23. Zhao, X., et al., MicroRNA-mediated control of oligodendrocyte differentiation. Neuron, 2010. 65(5): p. 612-26.
24. Dugas, J.C., et al., Dicer1 and miR-219 Are required for normal oligodendrocyte differentiation and myelination. Neuron, 2010. 65(5): p. 597-611.
25. Shen, S. and P. Casaccia-Bonnefil, Post-translational modifications of nucleosomal histones in oligodendrocyte lineage cells in development and disease. J Mol Neurosci, 2008. 35(1): p. 13-22.
26. Emery, B., Regulation of oligodendrocyte differentiation and myelination. Science, 2010. 330(6005): p. 779-82.
27. Emery, B., Transcriptional and post-transcriptional control of CNS myelination. Curr Opin Neurobiol, 2010. 20(5): p. 601-7.
28. Nave, K.A., Oligodendrocytes and the "micro brake" of progenitor cell proliferation. Neuron, 2010. 65(5): p. 577-9.
29. Dugas, J.C. and L. Notterpek, MicroRNAs in oligodendrocyte and Schwann cell differentiation. Dev Neurosci, 2011. 33(1): p. 14-20.
30. He, X., et al., Unwrapping Myelination by MicroRNAs. Neuroscientist, 2012. 18(1): p. 45-55.
31. Jacob, C., F. Lebrun-Julien, and U. Suter, How histone deacetylases control myelination. Mol Neurobiol, 2011. 44(3): p. 303-12.
32. Li, J.S. and Z.X. Yao, MicroRNAs: Novel Regulators of Oligodendrocyte Differentiation and Potential Therapeutic Targets in Demyelination-Related Diseases. Mol Neurobiol, 2012. 45(1): p. 200-12.
33. Taveggia, C., M.L. Feltri, and L. Wrabetz, Signals to promote myelin formation and repair. Nat Rev Neurol, 2010. 6(5): p. 276-87.
34. Junker, A., R. Hohlfeld, and E. Meinl, The emerging role of microRNAs in multiple sclerosis. Nat Rev Neurol, 2011. 7(1): p. 56-9.
35. Kremer, D., et al., The complex world of oligodendroglial differentiation inhibitors. Ann Neurol, 2011. 69(4): p. 602-18.
36. Dutta, R. and B.D. Trapp, Mechanisms of neuronal dysfunction and degeneration in multiple sclerosis. Prog Neurobiol, 2011. 93(1): p. 1-12.
37. Guerau-de-Arellano, M., et al., miRNA profiling for biomarker discovery in multiple sclerosis: From microarray to deep sequencing. J Neuroimmunol, 2011.
38. Kemppinen, A., S. Sawcer, and A. Compston, Genome-wide association studies in multiple sclerosis: lessons and future prospects. Brief Funct Genomics, 2011. 10(2): p. 61-70.
39. Kotter, M.R., C. Stadelmann, and H.P. Hartung, Enhancing remyelination in disease--can we wrap it up? Brain, 2011. 134(Pt 7): p. 1882-900.
40. Yao, Z.X., MicroRNA Patents in Demyelinating Diseases: A New Diagnostic and Therapeutic Perspective. Recent Pat DNA Gene Seq, 2012.
41. Liu, J. and P. Casaccia, Epigenetic regulation of oligodendrocyte identity. Trends Neurosci, 2010. 33(4): p. 193-201.
42. Swiss, V.A., et al., Identification of a gene regulatory network necessary for the initiation of oligodendrocyte differentiation. PLoS One, 2011. 6(4): p. e18088.
43. Norton, W.T. and S.E. Poduslo, Myelination in rat brain: method of myelin isolation. J Neurochem, 1973. 21(4): p. 749-57.
44. Wiggins, R.C. and G.N. Fuller, Analysis of distribution of rat sciatic nerve protein among soluble, insoluble, and myelin subfraction. Neurochem Res, 1981. 6(6): p. 719-27.
45. Haley, J.E., F.G. Samuels, and R.W. Ledeen, Study of myelin purity in relation to axonal contaminants. Cell Mol Neurobiol, 1981. 1(2): p. 175-87.
46. Larocca, J.N. and W.T. Norton, Isolation of myelin. Curr Protoc Cell Biol, 2007. Chapter 3: p. Unit3 25.
47. Lees, M.B. and S.A. Paxman, Myelin proteins from different regions of the central nervous system. J Neurochem, 1974. 23(4): p. 825-31.
48. McIntyre, R.J., et al., Isolation and characterization of myelin-related membranes. J Neurochem, 1978. 30(5): p. 991-1002.
49. Banik, N.L. and M.E. Smith, Protein determinants of myelination in different regions of developing rat central nervous system. Biochem J, 1977. 162(2): p. 247-55.
50. Deber, C.M. and S.J. Reynolds, Central nervous system myelin: structure, function, and pathology. Clin Biochem, 1991. 24(2): p. 113-34.
51. Norton, W.T. and S.E. Poduslo, Myelination in rat brain: changes in myelin composition during brain maturation. J Neurochem, 1973. 21(4): p. 759-73.
52. Patzig, J., et al., Quantitative and integrative proteome analysis of peripheral nerve myelin identifies novel myelin proteins and candidate neuropathy loci. J Neurosci, 2011. 31(45): p. 16369-86.
53. Werner, H.B., et al., Proteolipid protein is required for transport of sirtuin 2 into CNS myelin. J Neurosci, 2007. 27(29): p. 7717-30.
54. Vanrobaeys, F., et al., Profiling of myelin proteins by 2D-gel electrophoresis and multidimensional liquid chromatography coupled to MALDI TOF-TOF mass spectrometry. J Proteome Res, 2005. 4(6): p. 2283-93.
55. Taylor, C.M., et al., Proteomic mapping provides powerful insights into functional myelin biology. Proc Natl Acad Sci U S A, 2004. 101(13): p. 4643-8.
56. Ishii, A., et al., Human myelin proteome and comparative analysis with mouse myelin. Proc Natl Acad Sci U S A, 2009. 106(34): p. 14605-10.
57. Roth, A.D., A. Ivanova, and D.R. Colman, New observations on the compact myelin proteome. Neuron Glia Biol, 2006. 2(1): p. 15-21.
58. Baer, A.S., et al., Myelin-mediated inhibition of oligodendrocyte precursor differentiation can be overcome by pharmacological modulation of Fyn-RhoA and protein kinase C signalling. Brain, 2009. 132(Pt 2): p. 465-81.
59. Dhaunchak, A.S., et al., A proteome map of axoglial specializations isolated and purified from human central nervous system. Glia, 2010. 58(16): p. 1949-60.
60. Aquino, D.A., et al., The constitutive heat shock protein-70 is required for optimal expression of myelin basic protein during differentiation of oligodendrocytes. Neurochem Res, 1998. 23(3): p. 413-20.
61. Aquino, D.A., C. Lopez, and M. Farooq, Antisense oligonucleotide to the 70-kDa heat shock cognate protein inhibits synthesis of myelin basic protein. Neurochem Res, 1996. 21(4): p. 417-22.
62. Boggs, J.M., Myelin basic protein: a multifunctional protein. Cell Mol Life Sci, 2006. 63(17): p. 1945-61.
63. Buser, A.M., et al., The septin cytoskeleton in myelinating glia. Mol Cell Neurosci, 2008.
64. Edgar, J.M., et al., Early ultrastructural defects of axons and axon-glia junctions in mice lacking expression of Cnp1. Glia, 2009. 57(16): p. 1815-24.
65. Hensley, K., et al., Collapsin response mediator protein-2: an emerging pathologic feature and therapeutic target for neurodisease indications. Mol Neurobiol, 2011. 43(3): p. 180-91.
66. Braun, R.J., et al., Two-dimensional electrophoresis of membrane proteins. Anal Bioanal Chem, 2007. 389(4): p. 1033-45.
67. Takamori, S., et al., Molecular anatomy of a trafficking organelle. Cell, 2006. 127(4): p. 831-46.
68. Pagliarini, D.J., et al., A mitochondrial protein compendium elucidates complex I disease biology. Cell, 2008. 134(1): p. 112-23.
69. Trotter, J., D. Bitter-Suermann, and M. Schachner, Differentiation-regulated loss of the polysialylated embryonic form and expression of the different polypeptides of the
neural cell adhesion molecule by cultured oligodendrocytes and myelin. J Neurosci Res, 1989. 22(4): p. 369-83.
70. Zonta, B., et al., Glial and neuronal isoforms of Neurofascin have distinct roles in the assembly of nodes of Ranvier in the central nervous system. J Cell Biol, 2008. 181(7): p. 1169-77.
71. Traka, M., et al., The neuronal adhesion protein TAG-1 is expressed by Schwann cells and oligodendrocytes and is localized to the juxtaparanodal region of myelinated fibers. J Neurosci, 2002. 22(8): p. 3016-24.
72. Waehneldt, T.V., Phylogeny of myelin proteins. Ann N Y Acad Sci, 1990. 605: p. 15-28.
73. Kirschner, D.A., et al., Myelin membrane structure and composition correlated: a phylogenetic study. J Neurochem, 1989. 53(5): p. 1599-609.
74. Gow, A., et al., CNS myelin and sertoli cell tight junction strands are absent in Osp/claudin-11 null mice. Cell, 1999. 99(6): p. 649-59.
75. Court, F.A., et al., Restricted growth of Schwann cells lacking Cajal bands slows conduction in myelinated nerves. Nature, 2004. 431(7005): p. 191-5.
76. Brostoff, S.W., H. Sacks, and C. DiPaola, The P2 protein of bovine root myelin: partial chemical characterization. J Neurochem, 1975. 24(2): p. 289-94.
77. Bolino, A., et al., Disruption of Mtmr2 produces CMT4B1-like neuropathy with myelin outfolding and impaired spermatogenesis. J Cell Biol, 2004. 167(4): p. 711-21.
78. Quarles, R.H., Myelin-associated glycoprotein (MAG): past, present and beyond. J Neurochem, 2007. 100(6): p. 1431-48.
79. Montague, P., et al., Myelin-associated oligodendrocytic basic protein: a family of abundant CNS myelin proteins in search of a function. Dev Neurosci, 2006. 28(6): p. 479-87.
80. Schaeren-Wiemers, N., et al., The raft-associated protein MAL is required for maintenance of proper axon--glia interactions in the central nervous system. J Cell Biol, 2004. 166(5): p. 731-42.
81. Corbeil, D., et al., Expression of distinct splice variants of the stem cell marker prominin-1 (CD133) in glial cells. Glia, 2009. 57(8): p. 860-74.
82. Chong, S.Y., et al., Neurite outgrowth inhibitor Nogo-A establishes spatial segregation and extent of oligodendrocyte myelination. Proc Natl Acad Sci U S A, 2012. 109(4): p. 1299-304.
83. Wu, C., et al., Beta4 tubulin identifies a primitive cell source for oligodendrocytes in the mammalian brain. J Neurosci, 2009. 29(24): p. 7649-57.
84. Ivanovic, A., et al., The cytoskeletal adapter protein 4.1G organizes the internodes in peripheral myelinated nerves. J Cell Biol, 2012. 196(3): p. 337-44.
85. Kramer-Albers, E.M. and R. White, From axon-glial signalling to myelination: the integrating role of oligodendroglial Fyn kinase. Cell Mol Life Sci, 2011. 68(12): p. 2003-12.
86. Cammer, W., et al., Brain carbonic anhydrase: activity in isolated myelin and the effect of hexachlorophene. J Neurochem, 1976. 27(1): p. 165-71.
87. Mela, A. and J.E. Goldman, The tetraspanin KAI1/CD82 is expressed by late-lineage oligodendrocyte precursors and may function to restrict precursor migration and promote oligodendrocyte differentiation and myelination. J Neurosci, 2009. 29(36): p. 11172-81.
88. Schardt, A., et al., The SNARE protein SNAP-29 interacts with the GTPase Rab3A: Implications for membrane trafficking in myelinating glia. J Neurosci Res, 2009. 87(15): p. 3465-79.
89. Johns, T.G. and C.C. Bernard, The structure and function of myelin oligodendrocyte glycoprotein. J Neurochem, 1999. 72(1): p. 1-9.
90. Gitik, M., et al., Myelin down-regulates myelin phagocytosis by microglia and macrophages through interactions between CD47 on myelin and SIRPalpha (signal regulatory protein-alpha) on phagocytes. J Neuroinflammation, 2011. 8: p. 24.
91. Brockschnieder, D., et al., Ermin, a myelinating oligodendrocyte-specific protein that regulates cell morphology. J Neurosci, 2006. 26(3): p. 757-62.
92. Chakraborty, G. and R. Ledeen, Fatty acid synthesizing enzymes intrinsic to myelin. Brain Res Mol Brain Res, 2003. 112(1-2): p. 46-52.
93. Birling, M.C., et al., A novel rat tetraspan protein in cells of the oligodendrocyte lineage. J Neurochem, 1999. 73(6): p. 2600-8.
94. Feldmann, A., et al., Transport of the major myelin proteolipid protein is directed by VAMP3 and VAMP7. J Neurosci, 2011. 31(15): p. 5659-72.
95. Fewou, S.N., et al., Down-regulation of polysialic acid is required for efficient myelin formation. J Biol Chem, 2007. 282(22): p. 16700-11.
96. Jakovcevski, I., Z. Mo, and N. Zecevic, Down-regulation of the axonal polysialic acid-neural cell adhesion molecule expression coincides with the onset of myelination in the human fetal forebrain. Neuroscience, 2007. 149(2): p. 328-37.
97. De Gasperi, R., et al., Peripheral Myelin Protein-22 Is Expressed in Cns Myelin. Transl Neurosci, 2010. 1(4): p. 282-285.
98. Camara, J., et al., Integrin-mediated axoglial interactions initiate myelination in the central nervous system. J Cell Biol, 2009. 185(4): p. 699-712.
99. Thurnherr, T., et al., Cdc42 and Rac1 signaling are both required for and act synergistically in the correct formation of myelin sheaths in the CNS. J Neurosci, 2006. 26(40): p. 10110-9.
100. Bronstein, J.M., Function of tetraspan proteins in the myelin sheath. Curr Opin Neurobiol, 2000. 10(5): p. 552-7.
101. Sancho, S., P. Young, and U. Suter, Regulation of Schwann cell proliferation and apoptosis in PMP22-deficient mice and mouse models of Charcot-Marie-Tooth disease type 1A. Brain, 2001. 124(Pt 11): p. 2177-87.
102. Notterpek, L., et al., Peripheral myelin protein 22 is a constituent of intercellular junctions in epithelia. Proc Natl Acad Sci U S A, 2001. 98(25): p. 14404-9.
103. Roux, K.J., et al., Modulation of epithelial morphology, monolayer permeability, and cell migration by growth arrest specific 3/peripheral myelin protein 22. Mol Biol Cell, 2005. 16(3): p. 1142-51.
104. Berger, P., A. Niemann, and U. Suter, Schwann cells and the pathogenesis of inherited motor and sensory neuropathies (Charcot-Marie-Tooth disease). Glia, 2006. 54(4): p. 243-57.
105. Simons, M. and G. Raposo, Exosomes--vesicular carriers for intercellular communication. Curr Opin Cell Biol, 2009. 21(4): p. 575-81.
106. Mathivanan, S., H. Ji, and R.J. Simpson, Exosomes: extracellular organelles important in intercellular communication. J Proteomics, 2010. 73(10): p. 1907-20.
107. Rana, S. and M. Zoller, Exosome target cell selection and the importance of exosomal tetraspanins: a hypothesis. Biochem Soc Trans, 2011. 39(2): p. 559-62.
108. Krämer-Albers, E.-M., et al., Oligodendrocytes secrete exosomes containing major myelin and stress-protective proteins: Trophic support for axons? Proteomics Clin. Appl., 2007. 1: p. 1446–1461.
109. Fitzner, D., et al., Selective transfer of exosomes from oligodendrocytes to microglia by macropinocytosis. J Cell Sci, 2011. 124(Pt 3): p. 447-58.
110. Bakhti, M., C. Winter, and M. Simons, Inhibition of myelin membrane sheath formation by oligodendrocyte-derived exosome-like vesicles. J Biol Chem, 2011. 286(1): p. 787-96.
111. Scholze, A.R. and B.A. Barres, A Nogo signal coordinates the perfect match between myelin and axons. Proc Natl Acad Sci U S A, 2012. 109(4): p. 1003-4.
112. Schiffmann, R. and O. Boespflug-Tanguy, An update on the leukodsytrophies. Curr Opin Neurol, 2001. 14(6): p. 789-94.
113. Di Rocco, M., et al., Genetic disorders affecting white matter in the pediatric age. Am J Med Genet B Neuropsychiatr Genet, 2004. 129B(1): p. 85-93.
114. Patay, Z., Diffusion-weighted MR imaging in leukodystrophies. Eur Radiol, 2005. 15(11): p. 2284-303.
115. Lyon, G., A. Fattal-Valevski, and E.H. Kolodny, Leukodystrophies: clinical and genetic aspects. Top Magn Reson Imaging, 2006. 17(4): p. 219-42.
116. Boespflug-Tanguy, O., et al., Genes involved in leukodystrophies: a glance at glial functions. Curr Neurol Neurosci Rep, 2008. 8(3): p. 217-29.
117. Kohler, W., Leukodystrophies with late disease onset: an update. Curr Opin Neurol, 2010. 23(3): p. 234-41.
118. Saugier-Veber, P., et al., X-linked spastic paraplegia and Pelizaeus-Merzbacher disease are allelic disorders at the proteolipid protein locus. Nat Genet, 1994. 6(3): p. 257-62.
119. Garbern, J.Y., et al., Proteolipid protein is necessary in peripheral as well as central myelin. Neuron, 1997. 19(1): p. 205-18.
120. Gruenenfelder, F.I., et al., Axon-glial interaction in the CNS: what we have learned from mouse models of Pelizaeus-Merzbacher disease. J Anat, 2011. 219(1): p. 33-43.
121. Henneke, M., et al., RNASET2-deficient cystic leukoencephalopathy resembles congenital cytomegalovirus brain infection. Nat Genet, 2009. 41(7): p. 773-5.
122. Vaurs-Barriere, C., et al., Pelizaeus-Merzbacher-Like disease presentation of MCT8 mutated male subjects. Ann Neurol, 2009. 65(1): p. 114-8.
123. Ghezzi, D., et al., SDHAF1, encoding a LYR complex-II specific assembly factor, is mutated in SDH-defective infantile leukoencephalopathy. Nat Genet, 2009. 41(6): p. 654-6.
124. Lopez-Hernandez, T., et al., Mutant GlialCAM causes megalencephalic leukoencephalopathy with subcortical cysts, benign familial macrocephaly, and macrocephaly with retardation and autism. Am J Hum Genet, 2011. 88(4): p. 422-32.
125. Rademakers, R., et al., Mutations in the colony stimulating factor 1 receptor (CSF1R) gene cause hereditary diffuse leukoencephalopathy with spheroids. Nat Genet, 2011. 44(2): p. 200-5.
126. Aubourg, P., The leukodystrophies: a window to myelin. Nat Genet, 1993. 5(2): p. 105-6.
127. Zara, F., et al., Deficiency of hyccin, a newly identified membrane protein, causes hypomyelination and congenital cataract. Nat Genet, 2006. 38(10): p. 1111-3.
128. Gazzerro, E., et al., Hyccin, the Molecule Mutated in the Leukodystrophy Hypomyelination and Congenital Cataract (HCC), is a Neuronal Protein. PLoS One, 2012.
129. Erblich, B., et al., Absence of colony stimulation factor-1 receptor results in loss of microglia, disrupted brain development and olfactory deficits. PLoS One, 2011. 6(10): p. e26317.
130. Mignot, C., et al., Alexander disease: putative mechanisms of an astrocytic encephalopathy. Cell Mol Life Sci, 2004. 61(3): p. 369-85.
131. Sawaishi, Y., Review of Alexander disease: beyond the classical concept of leukodystrophy. Brain Dev, 2009. 31(7): p. 493-8.
132. Favre-Kontula, L., et al., GlialCAM, an immunoglobulin-like cell adhesion molecule is expressed in glial cells of the central nervous system. Glia, 2008. 56(6): p. 633-45.
133. Lopez-Hernandez, T., et al., Molecular mechanisms of MLC1 and GLIALCAM mutations in megalencephalic leukoencephalopathy with subcortical cysts. Hum Mol Genet, 2011. 20(16): p. 3266-77.
134. Leegwater, P.A., et al., Mutations of MLC1 (KIAA0027), encoding a putative membrane protein, cause megalencephalic leukoencephalopathy with subcortical cysts. Am J Hum Genet, 2001. 68(4): p. 831-8.
135. Heiman, M., et al., A translational profiling approach for the molecular characterization of CNS cell types. Cell, 2008. 135(4): p. 738-48.
136. Doyle, J.P., et al., Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell, 2008. 135(4): p. 749-62.
137. Lein, E.S., et al., Genome-wide atlas of gene expression in the adult mouse brain. Nature, 2007. 445(7124): p. 168-76.
138. Fu, H., et al., A genome-wide screen for spatially restricted expression patterns identifies transcription factors that regulate glial development. J Neurosci, 2009. 29(36): p. 11399-408.
139. Gray, P.A., et al., Mouse brain organization revealed through direct genome-scale TF expression analysis. Science, 2004. 306(5705): p. 2255-7.
140. Fu, H., S. Kesari, and J. Cai, Tcf7l2 is Tightly Controlled During Myelin Formation. Cell Mol Neurobiol, 2011.
141. Pedre, X., et al., Changed histone acetylation patterns in normal-appearing white matter and early multiple sclerosis lesions. J Neurosci, 2011. 31(9): p. 3435-45.
142. Visel, A., C. Thaller, and G. Eichele, GenePaint.org: an atlas of gene expression patterns in the mouse embryo. Nucleic Acids Res, 2004. 32(Database issue): p. D552-6.
143. Magdaleno, S., et al., BGEM: an in situ hybridization database of gene expression in the embryonic and adult mouse nervous system. PLoS Biol, 2006. 4(4): p. e86.
144. Geschwind, D.H. and G. Konopka, Neuroscience in the era of functional genomics and systems biology. Nature, 2009. 461(7266): p. 908-15.
145. Richardson, L., et al., EMAGE mouse embryo spatial gene expression database: 2010 update. Nucleic Acids Res, 2010. 38(Database issue): p. D703-9.
146. Diez-Roux, G., et al., A high-resolution anatomical atlas of the transcriptome in the mouse embryo. PLoS Biol, 2011. 9(1): p. e1000582.
147. Finger, J.H., et al., The mouse Gene Expression Database (GXD): 2011 update. Nucleic Acids Res, 2011. 39(Database issue): p. D835-41.
148. Fewou, S.N., et al., Myelin protein composition is altered in mice lacking either sulfated or both sulfated and non-sulfated galactolipids. J Neurochem, 2010. 112(3): p. 599-610.
149. Howng, S.Y., et al., ZFP191 is required by oligodendrocytes for CNS myelination. Genes Dev, 2010. 24(3): p. 301-11.
150. Chew, L.J., et al., SRY-box containing gene 17 regulates the Wnt/beta-catenin signaling pathway in oligodendrocyte progenitor cells. J Neurosci, 2011. 31(39): p. 13921-35.
151. Graumann, U., et al., Molecular changes in normal appearing white matter in multiple sclerosis are characteristic of neuroprotective mechanisms against hypoxic insult. Brain Pathol, 2003. 13(4): p. 554-73.
152. Robinson, W.H., P.J. Utz, and L. Steinman, Genomic and proteomic analysis of multiple sclerosis. Opinion. Curr Opin Immunol, 2003. 15(6): p. 660-7.
153. Kinter, J., T. Zeis, and N. Schaeren-Wiemers, RNA profiling of MS brain tissues. Int MS J, 2008. 15(2): p. 51-8.
154. Han, M.H., et al., Proteomic analysis of active multiple sclerosis lesions reveals therapeutic targets. Nature, 2008. 451(7182): p. 1076-81.
155. Zeis, T., et al., Molecular changes in white matter adjacent to an active demyelinating lesion in early multiple sclerosis. Brain Pathol, 2009. 19(3): p. 459-66.
156. Zeis, T., et al., Gene expression analysis of normal appearing brain tissue in an animal model for multiple sclerosis revealed grey matter alterations, but only minor white matter changes. J Neuroimmunol, 2008. 205(1-2): p. 10-9.
157. Derfuss, T., et al., Contactin-2/TAG-1-directed autoimmunity is identified in multiple sclerosis patients and mediates gray matter pathology in animals. Proc Natl Acad Sci U S A, 2009. 106(20): p. 8302-7.
158. Hakak, Y., et al., Genome-wide expression analysis reveals dysregulation of myelination-related genes in chronic schizophrenia. Proc Natl Acad Sci U S A, 2001. 98(8): p. 4746-51.
159. Tkachev, D., et al., Oligodendrocyte dysfunction in schizophrenia and bipolar disorder. Lancet, 2003. 362(9386): p. 798-805.
160. Lin, C.Y., A. Sawa, and H. Jaaro-Peled, Better understanding of mechanisms of schizophrenia and bipolar disorder: from human gene expression profiles to mouse models. Neurobiol Dis, 2012. 45(1): p. 48-56.
161. Sequeira, P.A., M.V. Martin, and M.P. Vawter, The first decade and beyond of transcriptional profiling in schizophrenia. Neurobiol Dis, 2012. 45(1): p. 23-36.
162. English, J.A., et al., The neuroproteomics of schizophrenia. Biol Psychiatry, 2011. 69(2): p. 163-72.
Figure legends
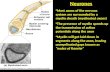
Figure 1. Central nervous system myelin.
(A) The optic nerve of an adult, wild-type mouse was visualized by transmission electron
microscopy upon fixation by high-pressure freezing and freeze substitution. Several
myelinated axons are shown in cross-section. Note the periodic arrangement of myelin
membranes. Electron micrograph kindly provided by W. Möbius. (B) One-dimensional gel-
separation of CNS myelin. Myelin purified from wild-type mouse brains was separated by SDS-
PAGE in different buffer systems providing improved resolution either in the low (MES) or
high (MOPS) molecular weight range. Proteins were visualized with colloidal Coomassie
(Coom., 5 µg protein load) or silver staining (0.5 µg protein load). Bands are denoted, which
are constituted by known myelin proteins according to mass spectrometric identification.
MAG, myelin associated glycoprotein; TUBA, α-tubulin; CNP, 2’,3’-cyclic nucleotide
phosphodiesterase; SIRT2, sirtuin 2; GAPDH, glyceraldehyde-3-phosphate dehydrogenase;
CLDN11, claudin 11/OSP; MOG, myelin oligodendrocyte glycoprotein; PLP, proteolipid protein;
MBP, myelin basic protein; CKB, brain creatine kinase; CA2, carbonic anhydrase 2; MOBP,
myelin-associated oligodendrocytic basic protein. In bands marked with arrowheads, only
proteins not previously associated with myelin were identified.
Figure 2. Assembling a compendium of CNS myelin proteins.
(A) The number of proteins identified in different approaches to the CNS myelin proteome is
plotted. The total number of myelin-associated proteins is unknown. Transmembrane proteins
(black) were systematically predicted by TMHMM2, Phobius and TMpred software. Proteins
derived from mitochondria (which are diminished but not entirely lost during myelin
purification) were predicted by Cello and Wolfpsort software and according to the literature.
T [55]; V [54]; R [57]; W [53]; B [58]; J [7]; D [59]. I [56] provide datasets for mouse (I-
m) and human (I-h) myelin. The integration of all datasets (‘All’) yields a catalogue of 1261
proteins for which a unique gene identifier was available. (B) Single and multiple
identifications. For all proteins identified in CNS myelin it was plotted in how many
approaches they were identified. Note that fewer than half of the proteins (48%) were
identified in only one approach. (C) Cross-study reproducibility. For all approaches to the
CNS myelin proteome it was plotted which percentage of identified proteins were additionally
identified in at least one other approach. Note the high overall reproducibility. The seemingly
low reproducibility of the dataset B [58] is due to the subfractionation of myelin by ion
exchange chromatography (IEX) (see text for details).
Figure 3. Venn diagram comparing the number of proteins identified in human versus rodent
CNS myelin (A), and in CNS versus PNS myelin (B) according to [52].
Figure 4. Profi les of myelin-associated mRNAs in oligodendrocytes.
(A) The mRNA abundance profiles in oligodendrocyte progenitor cells (OPC), pre-myelinating,
post-mitotic oligodendrocytes (OL) and myelinating oligodendrocytes (mOL), as determined
by [9], were filtered for the proteins identified by MS in purified myelin. Upon k-means
clustering, the normalized mRNA-abundance profiles were plotted with regard to the
differentiation stage. Genes with significant mRNA-abundance changes were categorized in
eight clusters. Known myelin-related genes are in bold, and genes encoding proteins probably
derived from mitochondrial, blood or nuclear contamination are in gray. mRNAs in the clusters
‘UP’ or ‘ASCENDING’ display significantly increased abundance during oligodendrocyte
differentiation while mRNAs in the clusters ‘DOWN’ or ‘DESCENDING’ are significantly
suppressed during development. (B) The numbers of mRNAs per cluster are given.
Supplemental Table S1. Compendium of proteins identified by mass
spectrometry in CNS myelin.
Out of the 1280 proteins listed, 1261 proteins could be assigned to a unique gene identifier,
of which 1249 could be correlated with the oligodendroglial transcriptome as established by
[9] (last column). The proteins are classified into three groups: known myelin proteins,
proteins identified by MS in myelin, and proteins presumably derived from mitochondria that
contaminate the myelin-enriched fraction. Identification of a protein in one of the 11 available
proteomic datasets (see main text for references) is indicated by "1". Potential mitochondrial
localization was classified by prior knowledge (MitoCarta) and prediction of subcellular
localization according to three algorithms (TargetP, Cello, Wolfpsort). The number of
transmembrane domains was predicted by three algorithms (TMpred, TMHMM, Phobius). If
available, correlation with a large-scale in situ-hybridization dataset (‘Allen Brain Atlas‘) is
provided (OL, oligodendrocyte; AS, astrocyte; NE, neuron) according to supplemental table
11 in [137].
Table 1. Relative abundance of major myelin proteins by mass spectrometric quantification Protein CNS myelin (%) PNS myelin (%) PLP 17 0.2 MBP 8 8 CNP 4 0.5 MOG 1 nd MAG 1 0.3 SIRT2 1 nd OSP 1 nd P0 nd 21 Periaxin nd 16 FASN nd 1 4.1G nd 1 Others 67 52 Proteins associated with purified CNS or PNS myelin were identified and quantified by LC-MSE [7, 52]. Selected myelin proteins are sorted by their relative abundance in CNS myelin. nd, not detected.
Table 2. Comparison of proteins identified in CNS myelin and disease genes associated with pathology of myelin or the white matter Gene Protein name mRNA profile OMIM Disease Remarks AHCY S-Adenosylhomocysteine Hydrolase Unchanged *180960
#613752 Hypermethioninemia Slow myelination, white
matter atrophy CTSD Cathepsin D Unchanged *116840
#610127 Ceroid Lipofuscinosis White matter
abnormalities GFAP Glial Fibrillary Acidic Protein Ascending *137780
#203450 Alexander Disease Leukodystrophy
GLUL Glutamate-Ammonia Ligase Late-UP *138290 #610015
Glutamine Deficiency Hypomyelination
HEPACAM Glial Cell Adhesion Molecule Down-UP *614133 #613926 #613925
Megalencephalic Leukoencephalopathy with Subcortical Cysts 2
Leukoencephalopathy
L1CAM L1 Cell Adhesion Molecule Descending *308840 #304100 #303350
Spastic Paraplegia type 1; Agenesis of the Corpus Callosum; MASA
White matter abnormalities
MBP Myelin Basic Protein Early-UP *159430 #601808
18q Deletion Syndrome Dysmyelination; Deleted chromosome segment includes MBP
MOG Myelin-Oligodendrocyte Glycoprotein Ascending *159465 #614250
Narcolepsy 7 Unknown pathogenesis
NPC1 Niemann-Pick C1 protein Unchanged *607623 #257220
Niemann-Pick Disease type C1
Impaired myelination
PHGDH Phosphoglycerate Dehydrogenase Late-UP *606879 #601815
PHGDH deficiency Dysmyelination
PLP1 Myelin Proteolipid Protein Early-UP *300401 #312080 #312920
Pelizaeus-Merzbacher Disease; Spastic Paraplegia type 2
Hypomyelinating leukodystrophy
PSAP Prosaposin Unchanged *176801 #249900 #611722
Metachromatic Leukodystrophy; Krabbe Disease
Leukodystrophy
PSAT1 Phosphoserine Aminotransferase Early-UP *610936 #610992
Phosphoserine amino-transferase deficiency
Poor white matter development
SLC12A6 Na/Cl-Cotransporter 3 Down-UP *604878 #218000
Agenesis of the Corpus Callosum
White matter abnormalities
SOD1 Superoxide Dismutase, Cytosolic Unchanged *147450 #105400
Amyotrophic lateral sclerosis Demyelination
Peroxisomal or dual localization ALDH3A2 Aldehyde Dehydrogenase 3a2 Ascending *609523
#270200 Sjogren-Larsson Syndrome Spasticity, paraplegia
HSD17B4
17-Beta-Hydroxysteroid Dehydrogenase IV
Unchanged *601860 #261515
Bifunctional protein deficiency
Dysmyelination
Mitochondrial or dual localization ETFA Electron Transfer Flavoprotein a Late-DOWN *608053
# 231680 Glutaric acidemia IIA Multisystemic, with
leukoencephalopathy HSPD1 Heat Shock 60kDa Protein 1 Late-DOWN *118190
#612233 Hypomyelinating Leukodystrophy 4
Leukodystrophy
NDUFV1 NADH-Ubiquinone Oxidoreductase Flavoprotein 1
Unchanged *161015 #203450 # 256000
Alexander Disease; Leigh syndrome
Demyelination
SLC25A12 Aspartate-Glutamate Carrier Aralar Unchanged *603667 #612949
Global cerebral hypomyelination
Hypomyelination
Probably mitochondrial localization AUH Au-specific RNA-binding Protein Ascending *600529
#250950 3-Methylglutaconic Aciduria White matter lesions
Proteins are listed that fulfill three criteria: (1) the protein was identified by MS in purified CNS myelin, (2) the corresponding mRNA was detected in oligodendrocytes and (3) mutations affecting the corresponding gene are associated with diseases that reportedly can include pathology of myelin or the white matter. Localization to mitochondria, which partly co-purify with myelin, was designated according to software-based prediction and a brain mitochondrial proteome study [68]. Note that mutations affecting mitochondrial proteins may infer myelin-related pathology. For proteins with additional expression in astrocytes, microglia, or neurons it is presently unknown whether loss/gain of function in oligodendrocytes is indeed causative of the disease. The mRNA abundance profile according to k-means cluster analysis (Fig. 4) is given.
Figure 1. Central nervous system myelin. (A) The optic nerve of an adult, wild-type mouse was visualized by transmission electron microscopy upon �xation by high-pressure freezing and freeze substitution. Several myelinated axons are shown in cross-section. Note the periodic arrangement of myelin membranes. Electron micrograph kindly provided by W. Möbius. (B) One-dimensional gel-separation of CNS myelin. Myelin puri�ed from wild-type mouse brains was separated by SDS-PAGE in di�erent bu�er systems providing improved resolution either in the low (MES) or high (MOPS) molecular weight range. Proteins were visualized with colloidal Coomassie (Coom., 5 µg protein load) or silver staining (0.5 µg protein load). Bands are denoted, which are constituted by known myelin proteins according to mass spectrometric identi�cation. MAG, myelin associated glycoprotein; TUBA, α-tubulin; CNP, 2’,3’-cyclic nucleotide phosphodiesterase; SIRT2, sirtuin 2; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; CLDN11, claudin 11/OSP; MOG, myelin oligodendrocyte glycoprotein; PLP, proteolipid protein; MBP, myelin basic protein; CKB, brain creatine kinase; CA2, carbonic anhydrase 2; MOBP, myelin-associated oligodendrocytic basic protein. In bands marked with arrowheads, only proteins not previously associated with myelin were identi�ed.
Figure 2. Assembling a compendium of CNS myelin proteins. (A) The number of proteins identi�ed in di�erent approaches to the CNS myelin proteome is plotted. The total number of myelin-associated proteins is unknown. Transmembrane proteins (black) were systematically predicted by TMHMM2, Phobius and TMpred software. Proteins derived from mitochondria (which are diminished but not entirely lost during myelin puri�cation) were predicted by Cello and Wolfpsort software and according to the literature. T [55]; V [54]; R [57]; W [53]; B [58]; J [7]; D [59]. I [56] provide datasets for mouse (I-m) and human (I-h) myelin. The integration of all datasets (‘All’) yields a catalogue of 1261 proteins for which a unique gene identi�er was available. (B) Single and multiple identi�cations. For all proteins identi�ed in CNS myelin it was plotted in how many approaches they were identi�ed. Note that fewer than half of the proteins (48%) were identi�ed in only one approach. (C) Cross-study reproducibility. For all approaches to the CNS myelin proteome it was plotted which percentage of identi�ed proteins were additionally identi�ed in at least one other approach. Note the high overall reproducibility. The seemingly low reproducibility of the dataset B [58] is due to the subfractionation of myelin by ion exchange chromatography (IEX) (see text for details).
Figure 3. Venn diagram comparing the number of proteins identi�ed in human versus rodent CNS myelin (A), and in CNS versus PNS myelin (B) according to [52].
Figure 4. Pro�les of myelin-associated mRNAs in oligodendrocytes.(A) The mRNA abundance pro�les in oligodendrocyte progenitor cells (OPC), pre-myelinating, post-mitotic oligodendrocytes (OL) and myelinating oligodendrocytes (mOL), as determined by [9], were �ltered for the proteins identi�ed by MS in puri�ed myelin. Upon k-means clustering, the normalized mRNA-abundance pro�les were plotted with regard to the di�erentiation stage. Genes with signi�cant mRNA-abundance changes were categorized in eight clusters. Known myelin-related genes are in bold, and genes encoding proteins probably derived from mitochondrial, blood or nuclear contamination are in gray. mRNAs in the clusters ‘UP’ or ‘ASCENDING’ display signi�cantly increased abundance during oligodendrocyte di�erentiation while mRNAs in the clusters ‘DOWN’ or ‘DESCENDING’ are signi�cantly suppressed during development. (B) The numbers of mRNAs per cluster are given.
Related Documents