SUPPLEMENTAL DATA Supplemental Protocol S1. Laser capture microdissection, RNA isolation and amplification. Butyl methyl methacrylate (BMM) was used as an embedding medium for structural preservation of ovule tissues as specified in Materials and Methods. This medium has been used routinely in cytological analyses, particularly for in situ hybridization in sexual and apomictic Hieracium. The BMM matrix is readily removed with acetone prior to laser microdissection (Baskin et al., 1992; Tucker et al., 2003; Okada et al., 2007; Tucker et al., 2012a). Histological detail was well preserved after acetone treatment and individual cells such as AI cells and other tissue layers remained relatively easy to identify prior to LCM (Figure S1A). However, when the Leica laser was used under the conditions described in Materials and Methods to dissect cell types it left a broad trace (Figure S1B-C) where adjoining cells were considered to be destroyed as per other published experiments using LCM. This trace is comparable to that previously reported in other studies dissecting larger tissue masses (Day et al., 2005) but not as fine as the uniquely modified laser used by Wuest et al. (2010) to dissect individual Arabidopsis mature female gametophyte cells. We conducted experiments to examine if we were able to isolate RNA from distinct cell types in sufficient quantity for 454 sequencing analyses that would enable detection of low copy genes. In preliminary LCM experiments, we examined the recovery of RNA when ovules were excised from 5μm thick ovary sections by laser capture (Figure S1A; white dashed line). We estimated that each captured ovule section contained 250 “cells”. When 20 ovule sections containing approximately 5,000 “cells” were captured and the RNA extracted using a PicoPure kit (Arcturus Bioscience Inc, Mountain View, CA, USA), the RNA recovery was low and difficult to quantify (NanoDrop spectrophotometer, Thermo Scientific, Wilmington, DE, USA). Therefore, the quantity of RNA was estimated by indirect measurement using RT- PCR on known quantities of whole ovary RNA samples to detect the presence of a low level ovary expressed gene MAP3K (ID 7.01 Table S2). The intensity of a band generated from using RNA extracted from 20, and 2, captured ovule sections was compared to that generated from a PCR reaction with 25ng, 5 ng and 1 ng of whole ovary input RNA (Figure S1D). From this we estimated that the RNA isolated from 20 ovule sections was approximately 1–2 ng. As this equates to approximately 5,000 “cells”, we considered it was unrealistic to harvest

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

SUPPLEMENTAL DATA
Supplemental Protocol S1. Laser capture microdissection, RNA isolation and amplification.
Butyl methyl methacrylate (BMM) was used as an embedding medium for structural
preservation of ovule tissues as specified in Materials and Methods. This medium has been used
routinely in cytological analyses, particularly for in situ hybridization in sexual and apomictic
Hieracium. The BMM matrix is readily removed with acetone prior to laser microdissection
(Baskin et al., 1992; Tucker et al., 2003; Okada et al., 2007; Tucker et al., 2012a).
Histological detail was well preserved after acetone treatment and individual cells such as AI
cells and other tissue layers remained relatively easy to identify prior to LCM (Figure S1A).
However, when the Leica laser was used under the conditions described in Materials and
Methods to dissect cell types it left a broad trace (Figure S1B-C) where adjoining cells were
considered to be destroyed as per other published experiments using LCM. This trace is
comparable to that previously reported in other studies dissecting larger tissue masses (Day et
al., 2005) but not as fine as the uniquely modified laser used by Wuest et al. (2010) to dissect
individual Arabidopsis mature female gametophyte cells. We conducted experiments to
examine if we were able to isolate RNA from distinct cell types in sufficient quantity for 454
sequencing analyses that would enable detection of low copy genes.
In preliminary LCM experiments, we examined the recovery of RNA when ovules were
excised from 5µm thick ovary sections by laser capture (Figure S1A; white dashed line). We
estimated that each captured ovule section contained 250 “cells”. When 20 ovule sections
containing approximately 5,000 “cells” were captured and the RNA extracted using a PicoPure
kit (Arcturus Bioscience Inc, Mountain View, CA, USA), the RNA recovery was low and
difficult to quantify (NanoDrop spectrophotometer, Thermo Scientific, Wilmington, DE,
USA). Therefore, the quantity of RNA was estimated by indirect measurement using RT-
PCR on known quantities of whole ovary RNA samples to detect the presence of a low level
ovary expressed gene MAP3K (ID 7.01 Table S2). The intensity of a band generated from
using RNA extracted from 20, and 2, captured ovule sections was compared to that generated
from a PCR reaction with 25ng, 5 ng and 1 ng of whole ovary input RNA (Figure S1D).
From this we estimated that the RNA isolated from 20 ovule sections was approximately 1–2
ng. As this equates to approximately 5,000 “cells”, we considered it was unrealistic to harvest

AI “cells” under these experimental conditions and obtain sufficient RNA for transcriptome
generation without amplification
In the next series of experiments, RNA extracted from 2 captured ovule sections ~500
“cells”) predicted to contain 0.1-0.2 ng RNA was amplified once using the MessageAmpTM
II
aRNA Kit (Ambion Inc, Austin, Texas, USA). Indirect measurement by RT-PCR using 2 μl
out of the total 100 μl of aRNA was equivalent to the 5 ng RNA input amplification control
indicating a total yield of 250 ng aRNA. We concluded that harvesting “100–250” individual AI
“cells” from 5 µm sections by laser capture would be sufficient material for RNA isolation,
amplification and use in downstream transcriptome experiments, particularly if a second
amplification step was included. Figure S1F shows that the amplified RNA was free of
genomic DNA contamination and Figure S1G shows that the Hieracium FIE gene was detected
in all samples as expected, while the HDMC1 gene expressed during meiosis in megaspore
mother cells was absent in the three samples. Remaining experiments focused on accumulating
captured cell samples and isolating RNA, amplification as described in Materials and Methods
and quantifying yields (Table S2).

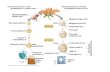
Supplemental Figure S1. Validation of quality and quantity of amplified RNA from laser
capture microdissected cell types from H. praealtum ovule sections.
A, An ovary longitudinal section showing an ovule outlined in white and an AI cell
outlined in yellow before laser microdissection. B, The same section in A after
tracing around the AI cell with the laser. The pink shading indicates the damage path
of the laser. The intact AI cell is outlined in yellow. C, View of the section in B after

the AI cell was harvested into the cap of the capture tube. Bars = 50 µm. D,
Estimation of the quantity of RNA. RNA isolated from 20 (Lane1) and 2 (Lane 2)
captured whole ovule sections (5µm thick) was used for RT-PCR; Lane 3-5, Stage
4 ovary RNAs (25, 5 and 1 ng, respectively) were used as controls for quantity
estimation. E, Quantification of amplified RNA. RNA isolated from 2 whole ovule
sections was subjected to one round of RNA amplification followed by RT-PCR
using 2 µl of amplified product (Lane 1). Lane 2-4, Stage 4 ovary RNAs (5, 1 and
0.2 ng, respectively) were used as controls for quantity estimation. F, Assessment
of genomic DNA contamination in the amplified RNA samples. RT-PCR was
carried out using equal quantities of aRNA from sporophytic ovule (SO) cells,
aposporous initial (AI) and early aposporous embryo sac (EAEs) samples, and
genomic (g) DNA. G, Validation of aRNA quality derived from LCM-harvested cells
using Hieracium genes with known ovule cell-type expression profiles. Expression of
HDMC1 and HFIE in Hieracium ovules has been characterized by RT-PCR and in
situ hybridization (Okada et al., 2007; Rodrigues et al., 2008).

Supplemental Figure S2. Validation of RT-PCR data obtained for low ovary-expressed genes
in LCM samples by quantitative real-time PCR. The three AI cell detected genes (class II,
Figure 1C; ID 9.45 RD22, ID 24.04 NLR and ID 27.18 LOX, Table S2) and two genes from
class III (Figure 1C; ID 9.20 BAM1 and ID 9.10 CKX) and one from class IV (Figure 1C, ID
25.03 S-protein; Table S2) were chosen for quantitative PCR analysis. Gene expression levels
relative to the reference HUBQ gene (Rodrigues et al., 2008) are plotted on the y-axis for each
examined cell type. Abbreviations; sporophytic ovule (SO) cell, aposporous initial (AI) cell and
early aposporous embryo sacs (EAEs).

Supplemental Figure S3. Examination of two AI cell-detected genes, an abscisic acid-responsive
RD22-like gene and a CC-NBS-LRR-like disease resistance gene, by in situ hybridization in
ovules of sexual and apomictic plants. A,B,E and F, antisense RNA (AS) probes were hybridized
with apomictic (Apo) H. praealtum R35 ovary sections and sexual (Sex) H.pilosella P36 ovary
sections in D and H. In C and G, sense RNA (S) probe was hybridised with apomictic H.
praealtum R35 ovary sections as a negative control. The 9.45 RD22 RNA probe was applied in A-
D, and the 24.04 NLR RNA probe in E-H.
Abbreviations: AI, aposporous initial cell; DM, degenerating megaspores; EAE sac, early
aposporous embryo sac; FM, functional megaspore; NE, nucellar epidermal cells. Bars = 20 µm.

Supplemental Figure S4. Flow chart of bioinformatic analyses of RNA sequencing data from
each laser-captured ovule cell type (AI, aposporous initial; EAE sacs, early aposporous
embryo sac; SO cells, sporophytic ovule cells) with thresholds and filtering as shown. Three
in silico analysis approaches were used. A, sequence based analysis of assembled contigs. B,
Pfam annotation analysis of un-assembled sequence reads, median read length was 260 base
pairs. C, Analysis of gene ontology annotations of assembled contigs assigned through
sequence homology, median contig length was 453 base pairs.

Supplemental Figure S5. Comparison of the number of sequence reads in 15 genes
assembled de novo and their expression in the three LCM samples by quantitative real-time
PCR. For each gene (see Table S2 for gene IDs) the normalized sequence read number in
reads per kilobase per million (RPKM) is presented in the left panel (y-axis) and Q-PCR
expression measures relative to the control UBC9 gene are presented in the right panel (y-
axis). The SO cell 64 gene was not verified and the AI 1771 and SO 1226 genes only
partially verified following the suite of comparative control genes (see Methods).

Supplemental Figure S6. Radar plots showing the similarity between Arabidopsis and
apomictic H.praealtum ovule LCM data sets based on detection of TAIR10 transcripts on
Agilent 4x44K arrays. The Hieracium datasets consist of TAIR10 homologues of Hieracium
contigs. Gene lists were filtered to remove genes expressed in every dataset, and the values
represent the similarity of each list to the base dataset indicated in each panel. NUC,
Arabidopsis nucellus at meiosis, OV (early), Arabidopsis whole ovule at meiosis, OV (late),
Arabidopsis whole ovule at FG2-4 stage, FG2-4, Arabidopsis female gametophyte stage FG3-4,
SO cells, Hieracium sporophytic ovule cells, AI, Hieracium aposporous initial cells, EAE sac,
Hieracium early aposporous embryo sac at FG2.

Supplemental Figure S7. Differentially expressed Arabidopsis homologs of H.praealtum
contigs from AI cell, SO cell and EAE sac data sets. Red dots indicate relative expression in
the FG2-4 vs ovule (late) and blue dots indicate the relative expression in the nucellus vs whole
ovule (early).

Cell type a Ovule b SO cells AI cells c EAE sacs c
Number of “cells” 500 2,500 270 100 Original RNA d 0.1-0.2 ng 0.5-1 ng 0.1-0.2 ng 0.1-0.2 ng 1st amplification e 600 ng 1.3 µg 646 ng 513 ng 2nd amplification f
N.A 189 µg 154 µg 126 µg
f, 10% (10 µl) of the first round amplified material was used for the second amplification. N.A, not available.
Supplemental Table S1 . Isolation of RNA from laser captured H. praealtum R35 apomictic ovule cell types and RNA yields after amplification.
a, Whole ovules, clusters of somatic ovule (SO) cells, aposporous initial (AI) cells and 2-4 nucleate early aposporous embryo (EAE) sacs were collected by LCM from 5 µm sections. b, Two LCM ovule sections estimated to contain 500 "cells" in cross section were used for RNA isolation and amplification.
c, This represents the number of individual AI “cells” or EAE sacs cut from 5µm thick sections. d, Quantity of original RNA was estimated using an indirect PCR method to amplify a control gene from a known amount of ovary RNA. e, Quantity of RNA after the first round of amplification was determined using a Nanodrop spectrophotometer.

Identifier Protein E-value SO AI EAE Forward Reverse
2.07 AT1G74770.1 unknown protein 2.00E-47 GO673075 VIATTACATGACCAC
ACGCAAG
AGGTAACGAGGAA
AGGATTC
2.11 AT3G56740.1
ubiquitin-associated
(UBA)/TS-N domain-
containing protein
2.00E-37 GO673077 + + + IACAACCCCTCCG
ATTAATTG
GTAAGCACACTCA
AAGTTGG
2.13 AT4G37190.1 unknown protein 4.00E-33 GO673078 VIICCAAAACCCCAAT
TTCGAAG
TTTATCAGGTTTC
GTGTCCG
2.18 No hits GO673081 IIIAACAACCAAGACG
TTCCAAC
TTTGAAGCCTCTA
TGCTGTC
2.20 AT5G59380.1MBD6 (methyl-CpG-binding
domain 6)2.00E-04 GO673083 I
GGGCAATTCTGTC
ATTTGTC
CTGGACAGAAATT
CAGATCC
2.21 AT3G04380.1SUVR4; histone-lysine N-
methyltransferase3.00E-09 GO673084 I
TAGGAAAGCTCTA
ATGCTGC
GATGGAACATGCT
TATCCTG
4.06 AT3G62770.1
AtATG18a (Arabidopsis
thaliana homolog of yeast
autophagy 18 (ATG18) a)
2.00E-08 GO673086 IATAGATTGCGATC
AATGGGG
GATGACTCAACTG
GAATATC
4.07 AT2G27880.1 argonaute protein (AGO5) 3.00E-05 GO673087 VITTCCATTGACCAA
TGGATGG
AAGAGCGTAGAAG
GTGAAAG
4.09 AT4G21710.1
EMB1989, RPB2, NRPB2 |
NRPB2 (EMBRYO
DEFECTIVE 1989); DNA
binding
1.00E-88 GO673089 IGTACGATACCGAA
TACGTTC
AAGAACGTCTCTT
TGACCAG
4.12 AT1G68720.1cytidine/deoxycytidylate
deaminase family protein2.00E-25 GO673090 I
TGTAATGGGAAGA
CACGAAG
AGAAGCTAAAAAG
GGTGCTG
4.18 No hits GO673092 ITCGCGTATTTAAG
TCGTCTG
TTTTGATCCTTCG
ATGTCGG
4.19 No hits GO673093 ICATAGCTTTTAGA
AGACGCG
GACAGAAATTGGG
ACAGAAG
4.20 AT5G19300.1 expressed protein 2.00E-17 GO673094 IIITGGCTAAAAAGAA
GAGGGAC
GACCCACAAATCT
TAAGCTG
5.04 AT5G65700.1
BAM1 (big apical meristem
1); ATP binding / kinase/
protein serine/threonine
kinase
2.00E-52 GO673095 IIITACGAGTACATGA
GAAACGG
GAACATTAGCCAG
TCTAGAG
5.05 AT5G47870.1 unknown protein 5.00E-40 GO673096 ND Paste photo ND VTGCAAACCGCAAA
AAATCCC
ATCACGTATTTCT
CCACACC
Supplemental Table S2. Expression of low level Hieracium piloselloides (D18) ovary genes in amplified AI cell, SO cell and EAE
sac H. praealtum aRNA samples by RT-PCR.
Primers for RT-PCRe
Classd
BlastX Against Arabidopsis protein (TAIR database)a
ID No.Accession
No.b
LCM RT-PCRc

Table S2 (continued)
5.11 AT3G49500.1
SGS2, SDE1, RDR6 | RDR6
(RNA-DEPENDENT RNA
POLYMERASE 6); nucleic
acid binding
5.00E-23 GO673100 IAATCACGACTTCA
AAAGCGG
CGAGTTTGAGAAG
TTTTCGG
5.13 AT4G38170.1FRS9 (FAR1-related
sequence 9); zinc ion binding9.00E-52 GO673101 IV
TTACATAAACGCC
ACCACTC
ATGAGGAAGGAAG
TGTAAGG
5.14 AT3G60860.1guanine nucleotide exchange
family protein4.00E-20 GO673102 I
AAACGAACCCTGT
CTAACAC
CCAAATAACCCAA
AGCAGAC
5.16 AT3G49600.1
UBP26 (ubiquitin-specific
protease 26); ubiquitin-
specific protease
4.00E-44 GO673104 IAATGGGCATTCAC
CAAGATC
AAATCGGCGGTTA
TCCAAAC
7.01 AT3G13530.1
MAP3KE1, MAPKKK7 |
MAPKKK7 (MAP3K EPSILON
PROTEIN KINASE); kinase
1.00E-44 GO673105 + + + IAACGCGATTTCAA
GAAGCAG
AAGACGCGGTTCA
AAAACTC
7.02 AT2G28360.1SIT4 phosphatase-
associated family protein9.00E-57 GO673106 III
AATTTCTTCCGGA
ACCATCC
CTGTCTCTACTTC
TCTCTTG
9.01 AT5G49910.1
HSC70-7, cpHSC70-2 |
cpHSC70-2 (HEAT SHOCK
PROTEIN 70-7); ATP binding
/ unfolded protein binding
2.00E-40 GO673107 IIIATTCTAATCCAAG
AACGGCC
TGGGAATCGTTGA
AGTATGC
9.02 AT5G65700.1
BAM1 (big apical meristem
1); ATP binding / kinase/
protein serine/threonine
kinase
5.00E-26 GO673108 ITCACTCTCCTCTT
CATATGC
AGCTGTAACAGGG
AGAAAAC
9.03 AT3G15010.1RNA recognition motif (RRM)-
containing protein5.00E-28 GO673109 III
AGAGAAAGACGGA
AGAGAAC
TTAACGCCAGAAT
TGTACCG
9.04 AT3G61250.1
AtMYB17 (myb domain
protein 17); DNA binding /
transcription factor
4.00E-31 GO673110 ITCCTAACACTGAG
TTCTAGG
CAAAATCCCAAAC
CTCCAAG
9.06 No hits GO673111 VCTCCATAATCAGT
CCATTCC
TCGGGTTTGTTTT
ACGAGAG
9.07 AT5G66680.1
DGL1 (defective glycosylation
1); dolichyl-
diphosphooligosaccharide-
protein glycotransferase
4.00E-59 GO673112 ITGGAGACAAGTTC
CATTTCC
ACAGGATCAATGA
TGACCTG
9.08 AT3G52260.1pseudouridine synthase
family protein 2.00E-38 GO673113 VI
CAACTCCATCTAA
TCCAACC
CAGCCATTGAGCA
AAAGTTC
9.09 AT4G11820.1
HMGS, MVA1, BAP1 | BAP1
(hydroxymethylglutaryl-CoA
synthase)
4.00E-50 GO673114 ITACAACATCACCC
CTTAACC
CAATTGGGGAACA
TGTACAC
9.10 AT1G75450.1
ATCKX5, ATCKX6, CKX5 |
CKX5 (CYTOKININ OXIDASE
5); cytokinin dehydrogenase
6.00E-16 GO673115 VTCAAATCGTGCCT
AACACTC
CATTGGATAATGG
GGAAGAG

Table S2 (continued)
9.11 AT5G42620.1metallopeptidase/ zinc ion
binding1.00E-19 GO673116 I
CATCAACACTTTG
TGCCATG
GCTGCTAAAAGAT
TAGCGTG
9.12 AT5G48930.1HCT | transferase family
protein9.00E-34 GO673117 IV
AAACCAACACCAA
ACCCATC
CACTTGAAACGTG
TAACCTG
9.14 AT5G50260.1 cysteine proteinase, putative 2.00E-55 GO673118 IIITGCATGGAGCTTT
GATTTCC
GTCGACTAGTTCT
TGTTCAG
9.15 AT2G39190.1ATATH8 | ATATH8 (ABC2
homolog 8)4.00E-31 GO673119 I
TAAACACTAATCC
TCACCGC
GATATTTCAAGAT
GGCGGAC
9.20 AT5G65700.1
BAM1 (big apical meristem
1); ATP binding / kinase/
protein serine/threonine
kinase
2.00E-85 GO673121 IIICTCTGAAAAACCA
GTGTGTG
TGTTGTTGTACAG
GTCAAGG
9.21 AT1G60200.1
splicing factor PWI domain-
containing protein / RNA
recognition motif (RRM)-
containing protein
3.00E-13 GO673122 ITGTTAGGGAAAGA
GATGTGC
TCGTATCCGGATT
AATTGGG
9.22 AT5G03690.1fructose-bisphosphate
aldolase, putative5.00E-24 GO673123 III
TGATGAGCTTATT
GCCAACG
AGAGAGTTTCTTC
GAAGAGG
9.23 AT4G00740.1dehydration-responsive
protein-related2.00E-19 GO673124 III
AATGGGCGAGAGT
AAAAGAG
TCATTCATGGGGT
TGCTATG
9.25 AT1G60730.1aldo/keto reductase family
protein3.00E-29 GO673125 I
TCCTGAAACCATC
AGAAGAG
TCCGAGTTCGTAT
AACTGTG
9.27 AT3G11980.1 MS2 (MALE STERILITY 2) 5.0E-09 GO673127 IIICAACTTCACAACC
AACATCC
TTCTTTCCAACAT
CAGGCAC
9.29 AT2G44490.1
PEN2 (PENETRATION 2);
hydrolase, hydrolyzing O-
glycosyl compounds
7.00E-21 GO673128 VIIAAATGCTGGTGAT
CGTTTGC
AGCAACAATTCGT
AGGAGTC
9.30 AT1G61620.1protein binding / zinc ion
binding1.00E-30 GO673129 I
TACACACTGCGAA
TGTTTCG
AACGAGGAAAAGA
CGGATTC
9.34 AT5G66420.1 unknown protein 1.00E-48 GO673131 ITGAACTCATGATC
ACACGTG
TGGAGGTGGAGAT
GATAAAC
9.38 No hits GO673133 IGAAGTGCATTTCC
AGATCTG
ATGGCAACAGTTT
TGGCAAG
9.39 AT1G64550.1
ATGCN3 (Arabidopsis
thaliana general control non-
repressible 3)
7.00E-15 GO673134 IIIATTTCGCCCTATA
GTGAGTC
GGCCAATTTGGTA
AGGAAAG

Table S2 (continued)
9.42 AT2G23910.1cinnamoyl-CoA reductase-
related 8.00E-17 GO673135 V
CAACCAATGTTGT
TACACGG
GAACATGGTGTCC
ATAAACG
9.44 AT2G44950.1
HUB1 (HISTONE MONO-
UBIQUITINATION 1); protein
binding / zinc ion binding
5.00E-23 GO673136 IACCCGACAACAAA
CTTTCAC
CACTCAAAGGAGA
CTACAAG
9.45 AT5G25610.1RD22 (RESPONSIVE TO
DESSICATION 22)2.00E-34 GO673137 II
TTGTGGGATGTCA
TGCAATG
CAGCGTAGAGGTA
ATTGAAC
9.47 AT3G21280.1
UBP7 (UBIQUITIN-SPECIFIC
PROTEASE 7); ubiquitin-
specific protease
1.00E-52 GO673138 VTACATATCAATGG
TGCGAGC
AGTGGATATCCAT
TGGAGTG
24.02 AT4G23500.1
glycoside hydrolase family 28
protein / polygalacturonase
(pectinase) family protein
2.00E-55 GO673139 ICATCTTGGCTATC
GATTCTG
TCAACATCACTCG
AAATCCC
24.03 AT4G28300.1hydroxyproline-rich
glycoprotein family protein7.00E-34 GO673140 III
ACTAAAAATCTCT
CTCCCCG
TGATGTACGATAG
TGAACCG
24.04 AT3G14470.1disease resistance protein
(NBS-LRR class)2.00E-18 GO673141 II
ATTGTGTCTGCCT
TGTACTC
GCACGATGCTAAA
GTTTCTG
24.12 No hits GO673142 IGAAGAACGTAGCA
AAATGCG
CAAGACACACGAC
ACATGAC
25.03 AT4G16195.1self-incompatibility protein-
related 2.0E-05 GO673145 IV
CTCAAGTCTTGTC
CTGTATC
TTGCGCGGATTTG
GATTATC
25.04 AT1G08760.1
similar to unknown protein
[Arabidopsis thaliana]
(TAIR:AT4G13370.1)
8.0E-24 GO673146 IVATTGTGACCCACA
ACTATGC
AAAGGGGGCGAT
CTTTTATG
25.10 AT1G55320.2
similar to acyl-activating
enzyme 17 (AAE17)
[Arabidopsis thaliana]
(TAIR:AT5G23050.1); similar
to acyl-activating enzyme 18,
putative
2.0E-46 GO673147 ICGTAACTCCTCCT
TCAATTG
TATTTGGGAGCAA
CGGATAG
26.01 AT5G67360.1 ARA12; subtilase 2.0E-16 GO673148 VACACACACACTCA
CTAAAGC
TATTTCAACTCCG
GCAACAC
26.04 AT4G12710.1armadillo/beta-catenin repeat
family protein6.0E-18 GO673149 VII
CATTCTCGTCTTT
TGCACAG
TTATGGAAGGGAA
GATGCAC
27.01 AT5G55310.1TOP1alpha
(TOPOISOMERASE I)8.0E-35 GO673152 IV
AAGCCACCTGTCA
ATCATTC
TCATCTGACTCAG
ATTCTGG
27.06 No hits GO673154 ICACTCTGCCACTT
ACAATAC
GCAGAATTCACCA
AGTGTTG

Table S2 (continued)
27.09 No hits GO673155 IAAGTTAGTCTTCA
GCCACAC
AAACGGTAAGAAA
ACACGGC
27.13 AT5G52510.1scarecrow-like transcription
factor 8 (SCL8)7.0E-47 GO673156 IV
CCATCCACCCAAA
ACTAATC
TTATCCGATCTCA
ATACGGC
27.14 AT4G23160.1 protein kinase family protein 6.0E-28 GO673157 VITTCAATGCAAGCA
TTGTGGG
TTTGGGTGATCGA
CCTTTAG
27.17 AT4G16280.1 FCA (FCA); RNA binding 1.0E-04 GO673158 ICTAATGGAGATCA
ATCACGG
ACAACAGACACAG
AGTCTTG
27.18 AT3G45140.1 ATLOX2, LOX2 | LOX2
(LIPOXYGENASE 2)4.0E-56 GO673159 II
AACAACAGTACAC
ACTCGTG
AGGCACTCTTGAA
ATGCTTC
27.27 AT1G19100.1
ATP-binding region, ATPase-
like domain-containing
protein-related
8.0E-04 GO673160 IIITTTATAACACTCC
CCCTTGG
TTGGTCCAGAAGT
ACCATAC
27.29 AT3G07160.1
ATGSL10 (GLUCAN
SYNTHASE-LIKE 10); 1,3-
beta-glucan synthase
7.0E-37 GO673162 IACAACAAGTTGCA
AAGCCAG
ACGAACAAAATCG
GGAACAG
27.30 AT3G03940.1 protein kinase family protein 2.0E-19 GO673163 VGTACACTTTTCCC
TTACACC
GAAACACAGGAGA
CATTACG
27.34 AT3G23640.1
HGL1 (HETEROGLYCAN
GLUCOSIDASE 1);
hydrolase, hydrolyzing
O-glycosyl compounds
2.0E-48 GO673164 IIIGGTTTTTGAACCG
ATTCTGG
TTGTTGATAGCCC
AATGACC
27.35 AT1G68530.1
CER6, G2, POP1, CUT1 |
CUT1 (CUTICULAR 1);
acyltransferase
6.0E-38 GO673165 IACGAAGCAACAAC
GTGAAAG
GTTGAAGCATCCA
GAATGAC
27.37 AT1G03280.1
transcription initiation factor
IIE (TFIIE) alpha subunit
family protein / general
transcription factor TFIIE
family protein
2.0E-53 GO673166 IACATCTCTAGGAA
GAACACC
GTTCGGGAAATGT
GGATTTG
HUBQ AT4G05320.1 UBQ10NM_001084
884 I
ACTCCACTTGGTC
TTGCGTCT
AGTACGGCCGTC
TTCAAGC
HFIE (D36) AT3G20740.1
FIS3, FIE1, FIE | FIE
(FERTILIZATION-
INDEPENDENT
ENDOSPERM 1); nucleotide
binding / transcription factor
e-172 EU439051 ICCAGGAGAGGGC
ACAGTTGATA
GGGCTAGTTTGCA
ATTCCCATA

Table S2 (continued)
HRBR AT3G12280.1
RBR, RB, RBL1, RBR1 |
RBR1 (RETINOBLASTOMA-
RELATED 1)
0 EU439049 VICATGTGTTGGAGA
GAGCACACA
ACTTGATGAAGCG
GGACCTTTC
HDMC1 AT3G22880.1
DMC1, ATDMC1 | ATDMC1
(RECA-LIKE GENE); ATP
binding / DNA-dependent
ATPase/ damaged DNA
binding
e-163 EF530197 - - - -CAGCTGGCTCAC
ACTCTCTG
TCAAGTACAGCTC
CAGCATCC
a, The most similar Arabidopsis genes searched by blasts with TAIR database (http://www.arabidopsis.org/index.jsp) are listed with AGI identifier number, similarity and E-value.
b, Accession number of the Hieracium gene sequence deposited on GenBank database
c, RT-PCR anaylsis using aRNA from LCM samples. SO, Somatic ovule cells; AI, aposporous initial cells; EAE, embryo sac; -, no expression
d, Class defined by expression pattern (see text and Figure 1). -, no expression in any cell type
e, Primer sequences used for RT-PCR analysis

Pfam Domain ID Pfam Domain GO ID GO Description Ontologya Number in
input listb
Number in
BG/Refb p-value
c
SO vs AI
Enriched in SO
PF03095 PTPA Phosphotyrosyl phosphate activator (PTPA) protein GO:0019211 phosphatase activator activity F 5 0 3.2E-03
PF03953 Tubulin_C Tubulin C-terminal domain GO:0003924 GTPase activity F 22 8 1.0E-03
PF00230 MIP Major intrinsic proteins GO:0005215 transporter activity F 23 9 1.3E-03
PF02984 Cyclin_C Cyclin GO:0005634 nucleus C 11 3 3.4E-03
PF01370 Epimerase NAD dependent epimerase/dehydratase family GO:0003824 catalytic activity F 8 2 3.3E-03
PF00657 Lipase_GDSL GDSL-like Lipase/Acylhydrolase GO:0016788 hydrolase activity, acting on ester bonds F 6 1 3.8E-03
PF01915 Glyco_hydro_3_C Glycoside hydrolase family 3 GO:0004553 hydrolase activity, hydrolyzing O-glycosyl compounds F 5 1 4.2E-03
PF08241 Methyltransf_11 Methyltransferase domain GO:0008168 methyltransferase activity F 5 1 4.2E-03
Enriched in AI
PF03931 Skp1_POZ Skp1 family, tetramerisation domain GO:0006511 ubiquitin-dependent protein catabolic process P 11 3 7.4E-04
PF00240 ubiquitin Ubiquitin GO:0005515 protein binding F 57 38 1.4E-03
PF08148 DSHCT DSHCT (NUC185) domain GO:0005524 ATP binding F 7 1 2.0E-03
PF00298 Ribosomal_L11 Ribosomal protein L11, RNA binding domain GO:0003735 structural constituent of ribosome F 12 4 2.7E-03
PF01466 Skp1 Skp1 family, dimerisation domain GO:0006511 ubiquitin-dependent protein catabolic process P 15 6 2.8E-03
PF01655 Ribosomal_L32e Ribosomal protein L32 GO:0003735 structural constituent of ribosome F 16 7 4.3E-03
AI vs EAE
Enriched in EAE
PF01095 Pectinesterase Pectinesterase (EC 3.1.1.11) GO:0030599 pectinesterase activity F 6 2 2.3E-03
PF00112 Peptidase_C1 Cysteine protease GO:0008234 cysteine-type peptidase activity F 11 7 2.3E-03
PF00235 Profilin Profilin GO:0003779 actin binding F 5 2 4.1E-03
Enriched in AI
PF00400 WD40 WD40 repeat GO:0005515 protein binding F 84 27 2.7E-03
Supplemental Table S3. Pfam enrichment analysis by reciprocal and pairwise comparisons between H. praealtum ovule cell types.

Table S3 (continued)
SO vs EAE
Enriched in SO
PF00400 WD40 WD40 repeat GO:0005515 protein binding F 82 27 3.3E-03
PF00230 MIP Major intrinsic proteins GO:0005215 transporter activity F 23 5 7.3E-03
PF03953 Tubulin_C Tubulin C-terminal domain GO:0003924 GTPase activity F 22 5 1.0E-02
PF01501 Glyco_transf_8 Glycosyl transferase family 8 GO:0016757 transferase activity, transferringGlycosylGroups F 11 1 6.8E-03
PF02535 Zip ZIP Zinc transporter GO:0046873 metal ion transmembrane transporter activity F 6 0 7.3E-03
PF03095 PTPA Phosphotyrosyl phosphate activator (PTPA) protein GO:0019211 phosphatase activator activity F 5 0 7.3E-03
PF01915 Glyco_hydro_3_C Glycoside hydrolase family 3 GO:0004553 hydrolase activity, hydrolyzing O-glycosyl compounds F 5 0 7.3E-03
PF08241 Methyltransf_11 Methyltransferase domain GO:0008168 methyltransferase activity F 5 0 7.3E-03
Enriched in EAE
PF01599 Ribosomal_S27 Ribosomal protein S27a GO:0003735 structural constituent of ribosome F 12 11 3.3E-03
PF03719 Ribosomal_S5_C Ribosomal protein S5, C-terminal domain GO:0003735 structural constituent of ribosome F 11 10 5.7E-03
PF01780 Ribosomal_L37ae Ribosomal L37ae protein GO:0003735 structural constituent of ribosome F 11 9 1.3E-03
PF01095 Pectinesterase Pectinesterase (EC 3.1.1.11) GO:0030599 pectinesterase activity F 6 4 4.5E-03
PF00366 Ribosomal_S17 Ribosomal protein S17 GO:0003735 structural constituent of ribosome F 6 4 4.5E-03
PF00164 Ribosomal_S12 Ribosomal protein S12 GO:0003735 structural constituent of ribosome F 9 7 6.1E-03
PF05162 Ribosomal_L41 Ribosomal L41 GO:0003735 structural constituent of ribosome F 9 7 6.1E-03
PF01655 Ribosomal_L32e Ribosomal protein L32 GO:0003735 structural constituent of ribosome F 9 7 6.1E-03
PF00338 Ribosomal_S10 Ribosomal protein S10p/S20e GO:0003735 structural constituent of ribosome F 7 4 6.8E-04
PF02298 Cu_bind_like Plastocyanin-like domain GO:0005507 copper ion binding F 8 5 2.5E-03
PF00112 Peptidase_C1 Cysteine protease GO:0008234 cysteine-type peptidase activity F 11 7 2.5E-03
PF00067 p450 Cytochrome P450 GO:0005506 iron ion binding F 8 4 3.0E-03
PF04573 SPC22 Signal peptidase subunit GO:0008233 peptidase activity F 6 1 8.5E-04
PF00120 Gln-synt_C Glutamine synthetase (EC 6.3.1.2) GO:0004356 glutamate-ammonia ligase activity F 6 1 8.5E-04
PF01929 Ribosomal_L14e Ribosomal protein L14 GO:0003735 structural constituent of ribosome F 10 5 1.3E-03
a, GO functional categories. P, Biological process; F, Molecular function; C, Cellular Component
b, Total numbers of Pfam domains used in input or background reference for each cell type are 38,667 (SO), 42,962 (AI) and 17,924 (EAE).
c, Enrichment of Pfam domains was analysed by Fisher test (p<0.05).

GO term Ontologya Description
Number in
input listb
Number in
BG/Refb p-value
c
SO vs AI
Enriched in SO
GO:0006468 P protein amino acid phosphorylation 133 72 5.6E-03
GO:0007166 P cell surface receptor linked
signaling pathway
61 27 3.9E-02
GO:0007167 P enzyme linked receptor protein
signaling pathway
57 25 3.9E-02
GO:0007169 P transmembrane receptor protein
tyrosine kinase signaling pathway
57 25 3.9E-02
GO:0023052 P signaling 486 390 3.9E-02
GO:0023034 P intracellular signaling pathway 78 41 5.0E-02
GO:0004672 F protein kinase activity 168 94 4.0E-04
GO:0016773 F phosphotransferase activity, alcohol
group as acceptor
203 130 2.6E-03
GO:0016798 F hydrolase activity, acting on glycosyl
bonds
78 39 1.5E-02
GO:0004674 F protein serine/threonine kinase
activity
122 73 1.5E-02
GO:0016772 F transferase activity, transferring
phosphorus-containing groups
444 359 2.4E-02
GO:0004553 F hydrolase activity, hydrolyzing O-
glycosyl compounds
71 36 2.4E-02
GO:0004871 F signal transducer activity 184 130 2.9E-02
GO:0060089 F molecular transducer activity 184 130 2.9E-02
GO:0016301 F kinase activity 380 303 2.9E-02
GO:0005975d
P carbohydrate metabolic process 14 36.9E-03
GO:0008168d
F methyltransferase activity 5 0 3.3E-02
Enriched in AI
GO:0006412 P translation 995 746 1.7E-06
GO:0042254 P ribosome biogenesis 259 177 2.6E-02
GO:0022613 P ribonucleoprotein complex
biogenesis
259 177 2.6E-02
GO:0010467 P gene expression 1466 1260 2.6E-02
GO:0044267 P cellular protein metabolic process 1714 1501 3.2E-02
GO:0003735 F structural constituent of ribosome 724 533 3.3E-05
GO:0005198 F structural molecule activity 860 671 3.2E-04
GO:0009908d
P flower development 11 3 2.5E-02
GO:0048229d
P gametophyte development 5 0 2.9E-02
GO:0051704d
P multi-organism process 19 9 3.6E-02
GO:0010876d
P lipid localization 8 2 5.0E-02
AI vs EAE
There are no significant GO terms found to be enriched in this comparison
Supplemental Table S4. GO enrichment analysis by reciprocal and pairwise comparisons between
H. praealtum ovule cell types.

Table S4 (continued)
SO vs EAE
Enriched in SO
GO:0050789 P regulation of biological process 776 329 7.3E-03
GO:0019219 P regulation of nucleobase,
nucleoside, nucleotide and nucleic
acid metabolic process
204 64 7.3E-03
GO:0019222 P regulation of metabolic process 290 101 7.3E-03
GO:0006350 P transcription 420 162 7.3E-03
GO:0051171 P regulation of nitrogen compound
metabolic process
210 66 7.3E-03
GO:0060255 P regulation of macromolecule
metabolic process
265 91 7.3E-03
GO:0045449 P regulation of transcription 195 62 7.8E-03
GO:0065007 P biological regulation 908 401 7.8E-03
GO:0006468 P protein amino acid phosphorylation 133 37 8.3E-03
GO:0010468 P regulation of gene expression 256 89 8.3E-03
GO:0080090 P regulation of primary metabolic
process
227 77 9.0E-03
GO:0031323 P regulation of cellular metabolic
process
239 83 1.1E-02
GO:0050794 P regulation of cellular process 693 299 1.1E-02
GO:0023033 P signaling pathway 96 24 1.2E-02
GO:0009889 P regulation of biosynthetic process 218 75 1.2E-02
GO:0031326 P regulation of cellular biosynthetic
process
218 75 1.2E-02
GO:0010556 P regulation of macromolecule
biosynthetic process
209 72 1.5E-02
GO:0023052 P signaling 486 201 1.5E-02
GO:0023034 P intracellular signaling pathway 78 19 2.9E-02
GO:0043412 P macromolecule modification 579 253 4.2E-02
GO:0007167 P enzyme linked receptor protein
signaling pathway
57 12 4.2E-02
GO:0007169 P transmembrane receptor protein
tyrosine kinase signaling pathway
57 12 4.2E-02
GO:0032774 P RNA biosynthetic process 125 39 4.2E-02
GO:0016740 F transferase activity 949 397 2.0E-04
GO:0016773 F phosphotransferase activity, alcohol
group as acceptor
203 59 7.6E-04
GO:0016772 F transferase activity, transferring
phosphorus-containing groups
444 166 7.9E-04
GO:0004672 F protein kinase activity 168 48 1.8E-03
GO:0016301 F kinase activity 380 142 2.3E-03
GO:0003700 F transcription factor activity 215 70 3.3E-03
GO:0030528 F transcription regulator activity 299 111 9.6E-03
GO:0004674 F protein serine/threonine kinase
activity
122 36 2.2E-02
GO:0004871 F signal transducer activity 184 65 4.3E-02
GO:0060089 F molecular transducer activity 184 65 4.3E-02
GO:0009057d
P macromolecule catabolic process 23 4 2.6E-02
GO:0003824d
F catalytic activity 144 53 1.7E-02
GO:0016787d
F hydrolase activity 42 11 2.9E-02

Table S4 (continued)
Enriched in EAE
GO:0006412 P translation 630 746 2.9E-15
GO:0019538 P protein metabolic process 1092 1698 7.4E-05
GO:0044267 P cellular protein metabolic process 981 1501 7.4E-05
GO:0010467 P gene expression 840 1260 7.4E-05
GO:0042254 P ribosome biogenesis 161 177 2.0E-04
GO:0022613 P ribonucleoprotein complex
biogenesis
161 177 2.0E-04
GO:0034645 P cellular macromolecule biosynthetic
process
847 1316 1.7E-03
GO:0009059 P macromolecule biosynthetic
process
850 1324 1.8E-03
GO:0044249 P cellular biosynthetic process 1005 1619 1.0E-02
GO:0009058 P biosynthetic process 1117 1824 1.4E-02
GO:0008152 P metabolic process 1944 3342 4.0E-02
GO:0006091 P generation of precursor metabolites
and energy
159 207 4.2E-02
GO:0003735 F structural constituent of ribosome 477 533 2.0E-14
GO:0005198 F structural molecule activity 570 671 2.0E-14
GO:0010876d
P lipid localization 6 2 1.9E-02
GO:0051704d
P multi-organism process 11 9 3.5E-02
GO:0048229d
P gametophyte development 3 0 3.7E-02
GO:0008289d
F lipid binding 5 2 4.4E-02
GO:0005840d
C ribosome 65 89 6.4E-03
GO:0022626d
C cytosolic ribosome 63 86 7.0E-03
GO:0030529d
C ribonucleoprotein complex 68 95 7.3E-03
GO:0015934d
C large ribosomal subunit 37 44 9.8E-03
GO:0033279d
C ribosomal subunit 59 81 1.0E-02
GO:0043232d
C intracellular non-membrane-bounded organelle79 117 1.1E-02
GO:0022625d
C cytosolic large ribosomal subunit 36 44 1.4E-02
GO:0032991d
C macromolecular complex 94 152 3.0E-02
d, Identification by post-hoc GO enrichment analysis using nested GO (nEASE) analysis
a, GO functional categories. P, Biological process; F, Molecular function; C, Cellular Component
b, Numbers of contigs with GO term annotation used in input or background reference for each cell
type are 4032 (SO), 3896 (AI) 1820 (EAE), see Table III.
c, Enrichment of GO terms was analysed by Fisher test (p<0.05) with FDR adjustment using AgriGO.

Gene name TAIR10 ID
SPOROCYTELESS AT4G27330
DISRUPTION OF MEIOTIC CONTROL 1 AT3G22880
SPORULATION 11 AT1G63990
RAD50 AT2G31970
ASYNAPTIC 1 AT1G67370
MUTS HOMOLOG 4 AT4G17380
SOLO DANCERS AT1G14750
SWITCH1/DYAD AT5G51330
MULTIPOLAR SPINDLE 1 AT5G57880
MUTL-HOMOLOGUE 1 AT3G24320
REC8 AT5G05490
PARTING DANCERS AT1G12790
TARDY ASYNCHRONOUS MEIOSIS AT1G77390
CHROMATIN-REMODELING PROTEIN 11 AT3G06400
pFM AT4G12250
MNEME AT5G39840
MtN3 AT5G40260
MtN3 AT5G23660
ARABINOGALACTAN PROTEIN 18 AT3G11700
Supplemental Table S5 Arabidopsis meiosis and megaspore
associated genes examined for presence in H. praealtum AI cell contigs.

Gene Name
Comparison GO term (P<0.05) TAIR10 ID AI EAE SO Nuc Ov ES Ov
Enriched in AI and EAE v SO
flower development AT1G52740* ● ● ● ● ● HISTONE H2A PROTEIN 9 (HTA9)
AT1G79000* ● ● ● ● ● HISTONE ACETYLTRANSFERASE OF THE CBP FAMILY 1 (HAC1)
AT2G40080 ● ● ● ● ● EARLY FLOWERING 4 (ELF4)
AT4G24960* ● ● ● ● HVA22 HOMOLOGUE D (HVA22D)
AT5G56030 ● ● ● ● ● HEAT SHOCK PROTEIN 81-2 (HSP81-2)
AT4G09960 ● ● ● ● ● SEEDSTICK (STK)
AT5G17690 ● ● ● ● ● TERMINAL FLOWER 2 (TFL2)
AT5G16780 ● ● ● ● DEFECTIVELY ORGANIZED TRIBUTARIES 2 (DOT2)
AT4G18960 ● ● ● ● ● ● AGAMOUS (AG)
AT2G38810 ● ● ● ● ● HISTONE H2A 8 (HTA8)
gametophyte development AT1G60490 ● ● ● ● VACUOLAR PROTEIN SORTING 34 (VPS34)
AT2G20490 ● ● ● ● ● ● NOP10, EDA27
AT4G24960* ● ● ● ● HVA22 HOMOLOGUE D (HVA22D)
AT2G35940* ● ● BEL1-LIKE HOMEODOMAIN 1 (BLH1)
AT2G47470* ● ● ● ● ● UNFERTILIZED EMBRYO SAC 5 (UNE5)
AT3G48750 ● ● ● ● CELL DIVISION CONTROL 2 (CDC2)
multi-organism process AT1G52740* ● ● ● ● ● HISTONE H2A PROTEIN 9 (HTA9)
AT3G03300 ● ● ● ● ● DICER-LIKE 2 (DCL2)
AT1G09770 ● ● ● ● ● ● CELL DIVISION CYCLE 5 (CDC5)
AT3G44750 ● ● ● ● ● HISTONE DEACETYLASE 3 (HDA3)
AT1G79000* ● ● ● ● ● HISTONE ACETYLTRANSFERASE OF THE CBP FAMILY 1 (HAC1)
AT4G00860 ● ● ● ● ● ● ATOZI1
AT3G51300 ● ● ● ● ● RHO-RELATED PROTEIN FROM PLANTS 1 (ROP1)
AT5G50320 ● ● ● ● ● ELONGATA 3 (ELO3)
AT3G59760 ● ● ● ● ● O-ACETYLSERINE (THIOL) LYASE ISOFORM C (OASC)
AT5G01650 ● ● ● ● ● TAUTOMERASE MIF SUPERFAMILY PROTEIN
AT2G47470* ● ● ● ● ● UNFERTILIZED EMBRYO SAC 5 (UNE5)
AT2G35940* ● ● BEL1-LIKE HOMEODOMAIN 1 (BLH1)
AT5G55390 ● ● ● ● ● ENHANCED DOWNY MILDEW 2 (EDM2)
AT3G16640 ● ● ● ● ● ● TRANSLATIONALLY CONTROLLED TUMOR PROTEIN (TCTP)
AT5G33340 ● ● CONSTITUTIVE DISEASE RESISTANCE 1 (CDR1)
AT1G48410 ● ● ● ● ● ARGONAUTE 1 (AGO1)
AT3G17220 PECTIN METHYLESTERASE INHIBITOR 2 (PMEI2)
lipid localisation AT1G03103 ●
AT5G38170 ● ●
AT2G18370 ● ●
AT3G18280 ● ● ●
AT3G51590 ● LIPID TRANSFER PROTEIN 12 (LTP12)
AT5G64080 ● ● ● ● XYLOGEN PROTEIN 1 (XYP1)
AT1G43666 ●
AT1G48750 ● ● ●
Enriched in SO v AI
carbohydrate metabolic process AT1G66980 ● ● ● ● SUPPRESSOR OF NPR1-1 CONSTITUTIVE 4 (SNC4)
AT1G79550 ● ● ● ● ● PHOSPHOGLYCERATE KINASE (PGK)
AT3G52990 ●
AT2G01290 ● ● ● ● ● RIBOSE-5-PHOSPHATE ISOMERASE 2 (RPI2)
AT4G38270 ● ● ● ● ● GALACTURONOSYLTRANSFERASE 3 (GAUT3)
AT1G80160 ● GLYOXYLASE I 7 (GLYI7)
AT5G55700 ● ● ● ● ● BETA-AMYLASE 4 (BAM4)
AT5G64380 ● ● ● ●
AT5G24400 ● ● ● ● ● EMBRYO DEFECTIVE 2024 (emb2024)
AT5G64570 ● ● ● ● ● BETA-D-XYLOSIDASE 4 (XYL4)
AT3G61490 ● ● ● ● ●
AT4G36890 ● ● ● ● ● IRREGULAR XYLEM 14 (IRX14)
methyltransferase activity AT5G26880 ● ● ● ● ● AGAMOUS-LIKE 26 (AGL26)
AT5G15380 ● ● ● DOMAINS REARRANGED METHYLASE 1 (DRM1)
AT3G12270 ● ● ● ● ● PROTEIN ARGININE METHYLTRANSFERASE 3 (PRMT3)
AT5G13710 ● ● ● ● ● STEROL METHYLTRANSFERASE 1 (SMT1)
AT4G25730 ● ● ● ● ●
Supplemental Table S6. Discriminatory gene annotations associated with significant GO terms found through nested GO enrichment analysis between H. praealtum ovule cell types and comparison
with Arabidopsis ovule genes on Agilent 4x44k arrays. Blue dot (●) indicates detectable expression, gray identifies genes not present on the array, asterisk (*) highlights genes contributing to more than 1 GO
term enrichment in this list.
HieraciumArabidopsis
(Meiosis)
Arabidopsis
(FG 2-4)
Related Documents