1 Supporting Information Salt-dependent Conditional Protein Splicing of an Intein from Halobacterium salinarum Julie N. Reitter, Christopher E. Cousin, Michael C. Nicastri, Mario V. Jaramillo and Kenneth V. Mills* College of the Holy Cross, Department of Chemistry, Worcester, MA 01610 Figure S1. Diagram of the chemical steps of protein splicing. Figure S2. Sequence alignment of Hsa and Mma PolII inteins. Figure S3. Sequence of MIHHsaWT fusion protein. Figure S4. Temperature dependence of splicing. Figure S5. Comparison of protein splicing of MIHHsa fusion proteins. Figure S6. Time dependence of protein splicing and N-terminal cleavage. Figure S7. Influence of ion concentration and identity on splicing and cleavage reactions. Figure S8. Reductant-dependent mobility shift of precursor protein and Cys mutants. Figure S9. pH dependence of protein splicing. Table S1. MALDI-TOF/MS analysis of protein splicing and cleavage of MIHHsaWT, MIHHsaQN and MIHHsaQACA. Table S2. N-terminal sequencing analysis of protein splicing of MIHHsaWT. Experimental Methods References for Supplemental Information

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

1
Supporting Information
Salt-dependent Conditional Protein Splicing of an Intein from
Halobacterium salinarum
Julie N. Reitter, Christopher E. Cousin, Michael C. Nicastri, Mario V. Jaramillo and
Kenneth V. Mills*
College of the Holy Cross, Department of Chemistry, Worcester, MA 01610
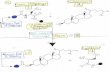
Figure S1. Diagram of the chemical steps of protein splicing.
Figure S2. Sequence alignment of Hsa and Mma PolII inteins.
Figure S3. Sequence of MIHHsaWT fusion protein.
Figure S4. Temperature dependence of splicing.
Figure S5. Comparison of protein splicing of MIHHsa fusion proteins.
Figure S6. Time dependence of protein splicing and N-terminal cleavage.
Figure S7. Influence of ion concentration and identity on splicing and cleavage reactions.
Figure S8. Reductant-dependent mobility shift of precursor protein and Cys mutants.
Figure S9. pH dependence of protein splicing.
Table S1. MALDI-TOF/MS analysis of protein splicing and cleavage of MIHHsaWT,
MIHHsaQN and MIHHsaQACA.
Table S2. N-terminal sequencing analysis of protein splicing of MIHHsaWT.
Experimental Methods
References for Supplemental Information

2
Figure S1. Diagram of the chemical steps of protein splicing. Protein splicing is a four-step
process: (1) an amide-thioester rearrangement of the peptide bond linking the N-extein and
intein, (2) transesterification resulting in transfer of the N-extein from the side chain of the first
residue of the intein to the first residue of the C-extein, (3) cyclization of the intein’s C-terminal
Asn coupled to peptide bond cleavage, and (4) conversion of the thioester linking the exteins to
an amide and potential hydrolysis of the C-terminal aminosuccinimide of the intein to Asn or
iso-Asn. Inteins with C-terminal Gln likely proceed via a similar mechanism involving Gln
cyclization.

3
Figure S2. Sequence alignment of Hsa and Mma PolII inteins. Alignment of intein sequences
by the EMBOSS Water sequence alignment tool.1 Conserved intein sequence motifs are
highlighted in green (Both inteins lack a central homing endonuclease domain, which would
comprise blocks C, D, and E.) Sequence identity shown as a bar between the two sequences,
sequence similarity shown with a dot. The gap in the Mma PolII intein sequence is the location
of the putative loop region in the Hsa PolII intein noted in the article.

4
Figure S3. Sequence of MIHHsaWT fusion protein. The sequence of the N-extein is given in
red (maltose binding protein and linker) and green (native N-extein residues), followed by the
intein in blue, native C-extein residues in green, and the linker and poly-His tag in red.

5
Figure S4. Temperature dependence of splicing. SDS-PAGE analysis of splicing of
MIHHsaWT. Splicing reactions consisted of 2.8 µM purified fusion protein in Buffer A
supplemented with 2 mM TCEP and 5 mM EDTA, with a final salt concentration of 2.0 M NaCl.
Splicing was initiated by incubation for 16 h at the temperatures indicated.

6
Figure S5. Comparison of protein splicing of MIHHsa fusion proteins. Analysis of protein
splicing and N-terminal cleavage assayed by SDS-PAGE, stained by InVision His-tag in-gel
stain. (Compare to Coomassie Blue stained gel in Figure 2.) Splicing or cleavage was initiated by
incubation for 16 h at 28°C of 2.8 µM purified fusion protein in Buffer A with the final salt
concentration given and supplemented with 5 mM EDTA. Lanes labeled “P” are untreated
proteins. MIHHsaWT and MIHHsaQN were incubated with 2 mM TCEP, and MIHHsaQACA
with 150 mM DTT. Gel was stained with InVision His-tag in-gel stain; band identities in italics
(M and I) note where bands in Figure 2 are not visualized in Figure S5, as the proteins lack a
His-tag. Predicted molecular masses for proteins: MIH precursor, 67.9 kDa; MH spliced product,
46.6 kDa; M cleavage product, 44.3 kDa; IH cleavage product, 23.6 kDa; I excised intein, 21.3
kDa.

7
Figure S6. Time dependence of protein splicing and N-terminal cleavage. SDS-PAGE
analysis of protein splicing of MIHHsaWT (A) or MIHHsaQN (B). Splicing was initiated by
28°C incubation of a reaction mixture, for the times indicated, of 2.8 µM purified fusion protein
in Buffer A supplemented with 2 mM TCEP and 5 mM EDTA, with a final salt concentration of
2.5 M NaCl.

8
Figure S7. Influence of ion concentration and identity on splicing and cleavage reactions.
SDS-PAGE analysis of splicing by MIHHsaWT. Splicing initiated by incubation for 16 h at
28°C of 2.8 µM fusion protein in buffer A supplemented with 2 mM TCEP and 5 mM EDTA.
Sample mixtures also were supplemented with the appropriate salt to the total concentrations
given. In panel A, the salt is supplied as NaCl or KCl. In panel B, the pH of the NaH2PO4 sample
was adjusted to 7.5 with NaOH, and the concentration above the lanes refers to the phosphate
concentration, not the sodium ion concentration. The first lane in each panel is a control that was
not incubated.

9
Figure S8. Reductant-dependent mobility shift of precursor protein and Cys mutants. SDS-
PAGE analysis of mobility shift on reduction of MIHHsaWT, MIHHsaQACA and MIHHsaC1A.
Precursor protein (3 µg) in buffer A analyzed by SDS-PAGE with or without 100 mM DTT in
the sample running buffer. Higher molecular weight bands likely due to disulfide linked dimers
in absence of DTT. We observe similar migration shifts of MIHHsaWT in unpurified cellular
extracts, suggesting the disulfide bonds are present in the E. coli.

10
Figure S9. pH dependence of protein splicing. SDS-PAGE analysis of protein splicing
facilitated by MIHHsaWT. Precursor fusion protein MIHHsaWT was exchanged into 20 mM
HEPES buffer at pH 7.0, 7.5, or 8.0 or 20 mM PIPES at pH 6.0 or 6.5, each with 500 mM NaCl.
Splicing was initiated by incubation for 16 h at 28°C of a reaction mixture of 2.8 µM purified
fusion protein, supplemented with 5 mM EDTA, 2.0 M NaCl, 2.0 mM TCEP and 5.0 mM
EDTA.

11
Table S1. MALDI-TOF/MS analysis of protein splicing and cleavage of MIHHsaWT,
MIHHsaQN and MIHHsaQACA. Splicing or cleavage was initiated by 28°C incubation for 16
h of a reaction mixture of 2.8 µM purified fusion protein in Buffer A supplemented with 5 mM
EDTA, with a final salt concentration of 2.5 M NaCl. MIHHsaWT and MIHHsaQN were
incubated with 2 mM TCEP, and MIHHsaQACA with 150 mM DTT. Samples were submitted to
the Tufts University Core Facility (Boston, MA). Not detected = n.d.
Purported
Protein ID
MIHHsa
WT
expected,
Mr
MIHHsa
WT
observed,
m/z
MIHHsa
QN
expected,
Mr
MIHHsa
QN
observed,
m/z
MIHHsa
QACA
expected,
Mr
MIHHsa
QACA
observed,
m/z
MIH 67,875 68,097 67,861 68,105 67,786 67,884
MIH
+2 peak
33,938 34,023 33,931 34,122 n.d. n.d.
MH 46,582 46,749 46,568 46,713 n.d. n.d.
M 44,265 44,434 44,265 44,438 44,265 44,383
IH 23,628 23,700 23,614 23,645 23,539 23,609
I 21,311 21,373 21,297 21,347 n.d. n.d.

12
Purported
Protein ID
Predicted N-terminal
Sequence
Observed N-terminal sequence
MH M-K-I-E-E-G M(S,G)-K-I-E-E-G
M M-K-I-E-E-G M(G,S)-K-I-E-E-G
IH C-F-H-P-E-T (W,G,S)-F-H-P(H,E)-E-T(E)
I C-F-H-P-E-T (W,G,S)-F-H-P-E-T(E)
Table S2. N-terminal sequencing analysis of protein splicing of MIHHsaWT. Splicing was
initiated by 28°C incubation for 16 h of a reaction mixture of 2.8 µM purified fusion protein in
Buffer A supplemented with 2 mM TCEP and 5 mM EDTA, with a final salt concentration of
2.5 M NaCl. Samples were submitted to the Tufts University Core Facility for analysis via
Edman degradation/N-terminal sequencing. Observed N-terminal sequence as called by core
facility, with lower confidence calls in parentheses; Cys residues are not detectable by this
method.

13
Experimental Procedures
Plasmid construction and protein expression. We created a plasmid to encode a fusion protein
containing the Hsa PolII intein. The fusion protein has an N-terminal Maltose binding protein
followed in frame by a short linker, the 10 C-terminal residues of the native N-extein, the 195
intein residues, the 6 N-terminal residues of the native C-extein, a short linker and a poly His tag.
(See Figure S3 for the sequence of the fusion protein.)
Genomic DNA was obtained from the American Type Culture Collection from strain NRC
34001. The intein containing gene was amplified by PCR using primers HsaPCRU, 5’-
CGTCCGTCGGCTACGCCAGGCCTTACTTCCA and HsaPCRL, 5’-
CCATCAGCAGCGAATTCCAGTCCTCGTC. The PCR product was digested with StuI and
EcoRI (New England Biolabs) and ligated into the same sites of the plasmid pMIHMma as
previously described.2 The sequence of the intein was verified by DNA sequencing and is
consistent with the translated protein sequence in the NCBI database (accession number
WP_010903745). The parent plasmid is a derivative of pMal-c2X from New England Biolabs.
Mutations at Cys1 of the intein to Ala (MIHHsaC1A), Gln195 and Cys+1 to Ala
(MIHHsaQACA) and Gln195 to Asn (MIHHsaQN) were facilitated by site-directed mutagenesis
using appropriate oligonucleotide primers.
We over-expressed the protein by transformation into E. coli BL21DE3 (Novagen-Merck
Biosciences), incubation at 37°C in rich media, induction at mid-log phase with 1 mM isopropyl-
β-D-1-thiogalactoside, and incubation of the cells with shaking at 20°C for 16 h. We pelleted the
cells by centrifugation at 3000 x g. Following fractionation of the lysate/extract and SDS-PAGE,

14
the over-expressed, unspliced intein precursor protein was found almost exclusively in the
soluble fraction, either with expression for 16 h at 20°C or for 3 h at 37°C, and after either
expression condition only unspliced precursor was produced, which was active on salt incubation
to the same extent.
Protein Purification and buffer exchange. Frozen cell pellets from 50 mL cultures were thawed
and suspended in 2.5 mL of buffer A (20 mM HEPES (4-(2-hydroxymethyl)-1-
piperazineethanesulfonic acid), pH 7.5, 500 mM NaCl) supplemented with BugBuster Extraction
buffer (Novagen-Merck Biosciences), 12 units/ml benzonase nuclease, 100 µM
phenylmethylsulfonyl fluoride and 10 µL Protease Inhibitor Cocktail P8849 (Sigma-Aldrich).
The soluble cell extract was purified using Talon metal affinity resin (Clontech), which was
prewashed with buffer A with a settled column bed volume of 300 µL. The loaded resin was
washed three times with 10 mL of buffer A supplemented with 10 mM imidazole and 0.1%
Tween 20 and one time with 10 mL of buffer A supplemented with 10 mM imidazole, and then
eluted three times with 500 µL of buffer A supplemented with 200 mM imidazole. Protein
concentration was determined via the Bradford method.3 Proteins were exchanged against buffer
A with a Millipore Ultracel-0.5 centrifugal filter with a nominal 3000 MWCO. In some cases, we
modified buffer A by substituting KCl for NaCl, or HEPES for PIPES (piperazine-N,N’-bis(2-
ethanesulfonic acid)) buffers at appropriate pH values.
Protein analysis by SDS-PAGE, MALDI-TOF/MS and Edman degradation. To study splicing,
we created 16 µL reaction mixtures as described in the figure legends. In each reaction, the
fusion protein was added at 3 µg, such that the initial fusion protein concentration was about 2.8

15
µM. Buffer A contains 500 mM NaCl; this is accounted for in the sodium ion concentrations
noted in the figures. Stock solutions of salts, EDTA and TCEP also were prepared in buffer A,
and, for the TCEP and NaH2PO4, the pH of the stock solution was adjusted to pH 7.5. TCEP was
used in place of DTT due to the nucleophilic nature of DTT; DTT was used for sample
preparation in reducing SDS-PAGE experiments. Sodium ions from the EDTA and any NaOH
used to adjust pH values are not accounted for in the sodium ion concentrations reported in the
figures, as they are relatively insignificant. For Figure S7, the pH of the NaH2PO4 samples was
adjusted with NaOH, resulting in a total sodium ion concentration of 3.7 M and 4.6 M for the 2.0
M and 2.5 M NaH2PO4 samples, respectively. A similar experiment in which the sodium ion
concentration is held at 2.0 M or 2.5 M, with corresponding phosphate concentrations of 1.1 M
or 1.4 M, respectively, gave similar results.
For SDS-PAGE analysis, we used precast 4-20% gradient Tris-glycine gels (Lonza) via the
Laemmli method.4 Sample buffer was supplemented with DTT to 100 mM unless otherwise
noted, and we used BenchMark protein ladder (Thermo-Fisher). Gels were stained with
Coomassie Blue, except for the gel in Figure S5, which was stained with InVision His-tag in-gel
stain (Life Technologies) according to manufacturer’s instructions and visualized with a BioRad
Gel Doc EZ Imager with a UV tray. To calculate percentage of splicing or N-terminal cleavage,
gels were scanned using an Epson Perfection V700 photo scanner and analyzed by Image J.5
Percentage splicing was calculated using densitometry data for appropriate bands as follows: 100
x [(MH)/46.6] / [((MH)/46.6) + ((MIH)/68) + ((M)/44.3)]. For N-terminal cleavage, percentage
was calculated by 100 x [(M)/44.3] / [((MH)/46.6) + ((MIH)/68) + ((M)/44.3)].

16
For MALDI-TOF analysis, we submitted samples to the Tufts University Core Facility. They
desalted the reaction mixtures with C18 ZipTips and spotted with 3,5-dimethoxy-4-
hydroxycinnamic acid, and analyzed the samples with an AB-Sciex Voyager DE Pro mass
spectrometer in linear mode. For N-terminal sequencing, we blotted onto polyvinylidene fluoride
membrane, stained with 0.1% Ponceau S in 1% aqueous acetic acid, destained in 50%
methanol/10% acetic acid, and washed with deionized water. The bands of interest were excised
and analyzed by the Tufts University Core Facility.
References
(1) Rice, P.; Longden, I.; Bleasby, A. Trends Genet 2000, 16, 276.
(2) Nicastri, M. C.; Xega, K.; Li, L.; Xie, J.; Wang, C.; Linhardt, R. J.; Reitter, J. N.; Mills, K. V.
Biochemistry 2013, 52, 5920.
(3) Bradford, M. M. Anal Biochem 1976, 72, 248.
(4) Laemmli, U. K. Nature 1970, 227, 680.
(5) Abramoff, M. D.; Magalhaes, P. J.; Ram, S. J. Biophotonics International 2004, 11, 36.
Related Documents