Recent experience to pandemic ( ) A(H1N1)2009 in Japan and collaboration with Asia collaboration with Asia Haruo Watanabe National Institute of Infectious Disease National Institute of Infectious Disease, Japan

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
Recent experience to pandemic ( )A(H1N1)2009 in Japan and collaboration with Asiacollaboration with Asia
Haruo Watanabe National Institute of Infectious DiseaseNational Institute of Infectious Disease,
Japan
Surveillance system for influenza in Japan
Pre‐existing system• Sentinel surveillance system for ILI: ‐ January 1987‐y‐ year‐round, weekly report‐ clinically diagnosed / lab‐confirmed y g /‐ sentinel sites: 5,000 ( including 3,000 pediatric clinic/hospital)
• Sentinel influenza virus surveillance‐ 1973‐‐ year‐round, daily report‐monitor the virological feature for vaccine
Surveillance system for influenza in Japan, Pre‐existing system‐cont’d
• School absentee reporting system for ILI• School absentee reporting system for ILI
‐ 1973‐
‐ Late Oct – Mid July ( about 40 – 30 epi wk )
‐ day‐care center, Kindergarten, elementary and middle high school
‐ school should report the number of absentees pand closed classes
‐ local health care center should take samples from‐ local health care center should take samples from the cluster(s) of absentee in a flu‐season for laboratory testlaboratory test
Surveillance system for A(H1N1)2009 in Japan
Case‐based surveillance‐ 28 April ‐ 23 July‐ Lab‐confirmed cases‐ Daily report by faxDaily report by fax ‐ Case fefinition:
‐ suspected case of influenza A(H1N1)v virus infection is defined as a person with high fever (>38°C) OR at least two acute respiratory symptoms (nasal g p y y pobstruction/rhinorrhea, sore throat, cough, fever/feverishness) AND who meets at least one of the following criteria:
a) within the last seven days returned from a country or region with an ) y y gepidemic of influenza A(H1N1)v;b) was in close contact (within two meters) with a confirmed case within the
past seven days;c) handled samples suspected of containing influenza A(H1N1)v virus in ac) handled samples suspected of containing influenza A(H1N1)v virus in a
laboratory or other setting within the past seven days;
‐ confirmed case of influenza A(H1N1)v virus infection is defined as a person with high fever (>38°C) OR at least two acute respiratory symptoms (nasalwith high fever (>38 C) OR at least two acute respiratory symptoms (nasal obstruction/rhinorrhea, sore throat, cough, fever/feverishness) AND influenza A(H1N1)v virus infection that has been laboratory confirmed by real‐time PCR and/or viral isolation.
Timeline of situation of A(H1N1)2009 i Jin Japan
Notification about Swine flu
2009 WHO declared the emergence of the novel flu25 Apr
2009
28
WHO declared the emergence of the novel flu•Japanese gov’t: launched case‐based surveillance for novel flu and
strengthened quarantine to travelers from affected areas29 Primer for RT‐PCR became
il bl(5 May)
9 May1st cases :travelers •Ceased strengthened quarantine
available
Outbreaks among several schools
1st domestic case
returned from Canada 16 May • Shifted to cluster surveillance and hospitalized‐case surveillance13
June
g q
• Expanded School absentee reporting
27July
Extensive school closure: over 4,200 schools w/ 650,000
system16July
All 47 prefectures t dchildren/students
15Aug
1st fatal case
reported cases
Reported cases w/ A(H1N1)2009, 28 Apr‐23 July ( 4 496*)(n=4,496*)
(By case‐based surveillance;*Cases the data of onset available )
Cumulative number, RtReported cases, Lt
Effectiveness of school closures inEffectiveness of school closures in Osaka and Kobe
Extensive school closure
Number of ILI cases/sentinel/week Number of ILI cases/sentinel/week in in Japan(from 2000 to 2010)Japan(from 2000 to 2010)Number / sentinel60.00Number / sentinel
50.00
0
Lower peak, and highest numberof ILR patients in 2009
40.000
1
2
3
4
of ILR patients, in 2009
30.00 5
6
7
8
20.00 9
10
10.00
0.00
1 3 5 7 9 11 13 15 17 19 21 23 25 27 29 31 33 35 37 39 41 43 45 47 49 51 53
2000~2010, week 13(upto April 4, 2010)
W k 36
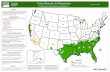
Geographical spread of A(H1N1)2009 during Aug. to Nov.Week 36
Week 37
Week 40
Week 39 Week 44 Week 47
Reported and estimated No. of patientsReported and estimated No. of patients
Reported ILI Total estimated ILIp2003‐04 0.77 million 9.23 m2004‐05 1.5 m 17.70 m
2009.29w~2010.13w ( by H1N1pdm) Estimated ILI cases 20.7m
(infection rate per population: 15.1%)at 30 March Confirmed Admitted patients No. 17,646
with underling diseases 6,599in ICU 1,022pregnant women 74pregnant women 74Influenza encephalopathy 543 Fatal case No. 171
* Japanese population 130 m
Age distribution of ILI (2009.28w~2010.10w)
600
インフルエンザ推計受診患者数年齢群別推移(2009年第28~2010年第10週)単位:万人
E ti t d t t l ILI 20 7インフルエンザ推計受診患者数男女別割合(2009年第28週~2010
年第10週)
500
Estimated total ILI=20.7m 年第10週)
malefemale
400 男, 1066, 51.6%
女, 1000, 48.4%
男
女
520476
300
229280
219100
200
155
10047 17 15
0
100
0~4歳 5~9歳 10~14歳 15~19歳 20~29歳 30~39歳 40~49歳 50~59歳 60~69歳 70歳~0~4歳 5~9歳 10~14歳 15~19歳 20~29歳 30~39歳 40~49歳 50~59歳 60~69歳 70歳~
Age distribution of admitted patients
新型インフルエンザ入院例年齢群別グラフ(2009年11月3日まで)
Age distribution of admitted patients
2500
1500
2000
1000
1500
500
0
0 4歳 5 9歳 10 14歳 15 19歳 20 39歳 40 59歳 60 79歳 80歳以上0~4歳 5~9歳 10~14歳 15~19歳 20~39歳 40~59歳 60~79歳 80歳以上
2009年11月3日現在
Age distribution for fatal cases of PI
32
34
新型インフルエンザ死亡例年齢群別基礎疾患別グラフ(2010年3月16日まで)新型インフルエンザ死亡例基礎疾患の有無割合グラフ(2010年3月16日まで)
24
26
28
30
32
138, 69.7%
60, 30.3%
基礎疾患有り
基礎疾患無し
16
18
20
22
24
With underlingdiseases
8
10
12
14
16
0
2
4
6
8
0
1歳未満 1~4歳 5~9歳 10~14歳 15~19歳 20~29歳 30~39歳 40~49歳 50~59歳 60~69歳 70~79歳 80歳以上
基礎疾患有り 基礎疾患無し
2010年3月16日現在
Mortality rate of PI among 100,000 population Mortality rate of PI among 100,000 population in several countiesin several counties
US Canada Mexico Australia UK France NZ Japan
Date collected
in several countiesin several counties
Date collected date 2/13 3/13 3/12 3/12 3/14 3/16 3/21 3/23
Death case No. Estimate12,000 429 1,111 191 457 309 20 198
Mortality rate (3.96) 1.32 1.05 0.93 0.76 0.50 0.48 0.15Infection rate ; 15~17% in all countries※尚、各国の死亡数に関してはそれぞれ定義が異なり、一義的に比較対象とならないことに留意が必要。
亡率
11.21.4
死亡率Canada
MexicoAustralia
0 20.40.60.81 Australia
UKFrance NZ
Japan0
0.2 Japan
出典:各国政府・WHOホームページから厚生労働省で作成 15
Influenza‐associated encephalopathy in Japan, ue a assoc a ed e cep a opa y apa ,week 1‐45, 2009
Nu
pathy
: AH1 pdm: Type A (week 28‐): Type A (‐week 28): Type B
umber of re
enceph
alop
: Type B: Undetermined: Number of reported cases of influenza per sentinel sites
eported cainflu
enza e
ses of influd cases of i
uenza* per of re
ported
Wk
sentinelNum
ber o
Wk
*including both lab‐confirmed and clinically diagnosed cases
A di ib i f i fl i d h l h i JAge distribution of influenza‐associated encephalopathy in Japan,week 28‐45, 2009
s orted case
mbe
r of rep
Num
y/o20s 30s 40s‐
Concluding Observations
Thi d i ' ll i t i ild f t t l• This pandemic's overall impact is mild, fortunately. • Patient number is estimated as 15% of population, however mortality rate
is very low (0.15/100,000 population), in Japan.
• The global and domestic context is becoming increasingly complex & unforgiving and handling future pandemics & global public health events will become more challengingwill become more challenging
– Where are the main gaps?How do we become better prepared & more flexible?– How do we become better prepared & more flexible?
– How do we respond more effectively?
• Influenza (not only pandemic) surveillance and control measure should be• Influenza (not only pandemic) surveillance and control measure should be strengthen more
Role and International Cooperation between NIID and Neighboring Asian Countries
Influenza Virus Research Center at NIID & WHO CC on
Influenza (Tokyo Center)
National Influenza Centers (NICs)
(Korea, China, Taiwan,Myanmar Laos MongolMyanmar, Laos, Mongol,
Singapore, Vietnam)Surveillance kit (Ref viruses, antisera)
Tech Supports (Manuals, Reagents) Outbreak information
Sending staff for training of lab staff
Host for Tech WS
Virus isolates and specimens
Sending staff to attend T‐WS
WHO CCsWHO HQ
Analyzed DataQ
NIID‐WG for Vaccine strain selection Vaccine strain selection for Northern & Southern Hems
L l G
Preparation of GuidelinesMOH, Japan
Local GovernmentsVaccine Manufacturers
Public
Vaccine ManufacturersPublic (Global)
Development of H1pdm detection systemDate Events24. Apr The sequence of H1N1 pdm virus was disclosed by CDC USA25 A Primers and probes for conv and real time RT PCR were designed by NIID25.Apr Primers and probes for conv. and real-time RT-PCR were designed by NIID
27. Apr Primers and probes were obtained28. Apr Primers and probes were obtained8 p e s a d p obes e e ob a ed29. Apr Usefulness of our detection system using swine Flu (H1N2) virus was verified
30. Apr Regents, positive control RNA(swine H1N2), primers and probes were provided to 76 Local Institutes of Health and 15 Quarantines in Japanprovided to 76 Local Institutes of Health and 15 Quarantines in Japan
2. May A/California/4/2009 (H1N1) pdm virus was sent from CDC 3. May Our detection system using H1N1 pdm virus was validated3. May Our detection system using H1N1 pdm virus was validated~ 5. May H1N1 pdm diagnosis system was established in all Local Institutes of Health
and Quarantines
6 May The information of our detection system was reported to the WHO6. May The information of our detection system was reported to the WHO (Published on 21. May)
8. May The first case traveled in Canada was detected in Narita Airport Quarantine
13. May The information of our detection system was reported to the ASEAN16. May The first case with no travel history was detected in Kobe city
Local Institutes of Health and Quarantine Laboratory in Japan
Local Institutes of Health76 places
Quarantine Laboratory15 places
NIID
fl i d d i l b l h b (Influenza virus detected in Local Pub Health Lab (2008 36w~2010 15w), in Japan
1600
1800
インフルエンザウイルス分離報告数週別グラフ(2008年第36~2010年15週)
1200
1400
600
800
1000
200
400
0
3637383940414243444546474849505152 1 2 3 4 5 6 7 8 9 1011121314151617181920212223242526272829303132333435363738394041424344454647484950515253 1 2 3 4 5 6 7 8 9 101112131415
2008 2009 2010AH1 AH3 B AH1pdm
2010年4月24日現在
OTV resistant H1N1pdm virusesOTV resistant- H1N1pdm viruses (as of April 2010)
67 Resistants / 5422 tested (1.2%)
All were sensitive to zanamivir
Training on laboratory diagnosis y g
P i i B i C b di I d i L PDR M l iParticipants; Burnei, Cambodia, Indonesia, Lao PDR, Malaysia, Philippines, Singapore, Thailand, Vietnam
Training and lecture; PT-PCR, Real time PCR, Sequencing(lecture)
Reagents provided;g p ;・RNase Inhibitor (Appliedbiosystems)・QuantiTect Probe RT-PCR Kit (QIAGEN) ・TaqMan MGB Probe and primers for real-time RT-PCRTaqMan MGB Probe and primers for real time RT PCR
Type A detection, H1 pdm detection・Primers for conventional RT-PCR
Type A detection H1 pdm detection H1(seasonal) detectionType A detection, H1 pdm detection, H1(seasonal) detectionH3(seasonal) detection
・Control RNA (A/Narita/1/2009(H1N1pdm)
Sakai/1
Hyogo/2
Sakai/2Osaka/180
Hyogo/1Kobe/1
Beijing/502
Shiga/1
Shiga/2Himeji/1
Okinawa/257
Osaka/1
Utsunomiya/1
Mexico/InDRE13551 MexicoCity/WR1301NOkinawa/254
Canada-PQ/RV1758
Osaka-C/2
Niigata/700
LaPaz/WR0096T
Amagasaki/1
Hunan/SWL3
Beijing/3
Korea/01
Canada-QC/RV1954
Osaka/2
Osaka-C/1
Utsunomiya/2
Okinawa/256
Amagasaki/2
Mar.13
Mar.24 1.2MC1
MC11
MC12Canada-SK/RV2486
Canada-MB/RV2023
Thailand/CU-B5
Singapore/ON1087
Shiga/1
Singapore/GP2686
New_York/3408
Mexico/InDRE13555
Utah/05Niigata/749
Shiga/3
Moscow/IIV04
Shizuoka/793
Italy/127
New_York/3551
Niigata/690Nonthaburi/102
New_York/3352
NewYork/4820
Shandong/1
Mexico/4604
New_York/4197
New_York/3260
I /1
Tokushima/1
New_York/3468
Akita/1
Shizuoka/759
Silver_Spring/SP509New_York/3389
Taiwan/T1773
New_York/3351
Kanagawa/137
1.3MC12
MC4New_York/3014
Zhejiang/1
New_York/18
Chiba-C/51
New_York/3651
California/14
New_York/3715
Denmark/524Denmark/523
Moscow/03
Chiba-C/48Yamaguchi/21
New_York/3413
New_York/3532New_York/3168
Denmark/528
Shanghai/71T
Yamaguchi/23
New_York/06
Iwate/1Yokohama/1
Moscow/IIV02
New_York/3166
Moscow/IIV03
Shizuoka/759
New York/11N Y k/3502
New_York/3170Guangdong/02
Taiwan/T0724
New_York/3898
New_York/3193New_York/3172
MC6
MC4
Hiroshima/201
Taiwan/T1338
Canada-ON/RV1589Canada-ON/RV1527
Shiga/43
New_York/3049
MexicoCity/WR1308TMie/41
Fukushima/1
New_York/3169
Taiwan/T0826
Okinawa/283
New_York/3214
Himeji/L1
New_York/20
Italy/49
Moscow/0253T
New_York/3463
New_York/23
Okinawa/259
Fuzhou/01
New_York/3234
New_York/4290
New_York/3176
New_York/11
NewYork/4870
New_York/3502
New_York/3012
MexicoCity/WR1312N
Fujian/1
New_York/3354
Sapporo/1
Kagoshima/56
Apr. 03
Cluster 2
MC7MC2
NewYork/4777
Hiroshima/230
Zhejiang-Yiwu/11
NewYork/4576
SanSalvador/0169T
Thailand/CU H9
Brandenburg/20
Okinawa/258
Mie/41
New_York/3007
New_York/10
Niigata/717
Mie/52
Bogota/0466N
Iwate/3
Hiroshima/220
Iwate/2
Zhejiang/DTID-ZJU02
Shanghai/1
Hiroshima/207
Taiwan/T1339
Zhejiang/DTID-ZJU03
New_York/3261
Wakayama/57
New_York/3264
Taiwan/T1821
y/
Kyrgyzstan/01
Tokushima/2
Aichi/202
Shanghai/37T
Aichi/198
Malaysia/820Mar. 13 MC3
MC5Hiroshima/216
Osaka/2031
Bethesda/SP508Thailand/CU-H9
Japan/PR1070
Italy/05
New_York/3313Kobe/91993
Taiwan/526
Shiga/44
Hiroshima/200New_York/3008
Italy/85
Osaka/2032
Michigan/02
New_York/3237
Kobe/91992
Texas/08
New_York/3992
Texas/09
Tochigi/154
NewYork/4790
Gifu-C/67
Mexico/InDRE13547
Singapore/ON1050
Saitama-C/88
Saitama/85
Bethesda/SP506
Texas/04
Nanjing/3
Singapore-Q/M168
Moscow/IIV05Jan.30
MC9
MC8
Canada-NS/RV1565
Beijing/02
Canada-NS/RV1535
New_York/1682
New_York/3455
Toronto/R8564
New_York/1669
Santo_Domingo/573N
Shizuoka-C/97
Kanagawa/140
E l d/195
Toronto/T5308
SantoDomingo/WR1068NSanto_Domingo/572N
Texas/09
Guangdong/1
Brawley/40082
Beijing/01Beijing/4
Narita/1
Washington/14
California/06
Brandenburg/19
New_York/3099
Canada-AB/RV1644
Mexico/4486
SantoDomingo/WR1059N
Indiana/09
Mexico/4108
Toronto/0462Mexico/InDRE13495
Canada-SK/RV1798
Santo_Domingo/568T
Cluster 1Feb.02
30.0
Paris/2590
Fukuoka-C/1
New_York/3307
Vladivostok/IIV17
Managua/0536NNanjing/2
Kagoshima/1
Mexico/InDRE4487
Fukuoka-C/2
England/195
Mexico/InDRE13494
SantoDomingo/WR1068NParis/2580
California/04
Paris/2592
California/07
Sichuan/1
Fukuoka-C/3
Vladivostok/IIV18
New_York/3178
Apr .1 May .1 Jun .1 Jul .1 Aug .1 Sep .1 Nov .1MC10
1. May 1. Jun. 1. Jul. 1. Aug. 1. Sep.
Time of virus introduced in Japan, which was analyzed from the genome evolution
SapporoIwateAkita
Niigata
TochigiUtsunomiya
5.15
6.511
3
2Fukushima
gSaitama
Chiba
KanagawaShizuoka
Ai hi
Narita
Yokohama
Nagano
45.17
5.163
6 99
*
**
AichiMieGifuShiga
OsakaSakaiAmagasaki
Wakayama
1
65.25
6.9
AmagasakiKobeHyogoHimejiTokushimaHiroshima
FukuokaYamaguchi
4.225.19
5
75.25
85.29
10FukuokaKagoshimaOkinawa 5.21
10
7.2412
Extensive school closure
ConclusionConclusion
1) Rapid development of new virus detection methodsis required especially in the WHO CC on influenza
2) Technical transfer and sharing of information amongcountries are very important to control thespreading of diseases
3) Characterization of virus genomes will tell us the3) a ac e a o o us ge o es e us eevolutional steps of virus and may explain theeffectiveness of countermeasures to the influenzaisolation
Laboratory-based net work among Asia and Pacific Rim
●Construction of PulseNet Asia-●Construction of PulseNet AsiaPanpacific
●Laboratory-based net work among Asia and Pacific rimNational National Institute of InfectiousInstitute of Infectious
Japan Asia
US CDCRapid detection of pathogens emerging in Asia and prevention
ASEAN Plus Three (APL)DiseaseDisease (NIID, Japan(NIID, Japan))
US.CDC
Philippine
Taiwan. CDCemerging in Asia and prevention of their spreading
Target pathogens;Bacteria (Enteric bacteria)
1)Collection of domestic i f ti
Australia
MalaysiaBacteria (Enteric bacteria)Virus ( Dengue fever virus, JE)Parasites (Malaria etc.)
1)Standardization of protocol
WHO,WPROinformation
2)Genetic and phenotypic characterization of the pathogens
3)Development of new 1)Standardization of protocolfor the detection of pathogensand genotyping
2)Quality control, validationof the protocol
Bangladesh,ICDDR B India NICED
New Zealandmethods
China, CDCKorea,NIH/CDC
of the protocol3)Construction of data-base4)Exchange of information and
researchers
ICDDR・B India,NICED
Local Health InstituteThailand, NIH/CDC
Vietnam,NIHE
Local Health Institute, Quarantine station
Related Documents