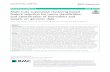
RAMclust/RAMsearch: efficient post-XCMS feature clustering and annotation of MS-based metabolomics datasets. Corey D. Broeckling and Jessica E. Prenni Colorado State University, Proteomics and Metabolomics Facility Acknowledgements: Asa Ben-Hur, and Fayyaz A Afsar helped to develop ramclustR algorithms. Steffen Neumann helped package ramclustR NIST (Steve Stein, David Sparkman, Dmitri Tchekhovskoi) Kevin Brown and Ben Sutton helped develop RAMsearch Introduction: • Non-targeted profiling by UPLC-MS is a powerful tool for metabolic profiling. • Electrospray ionization is soft, but many signals are generated for a given metabolite: • Isotopes • Adducts • Multimers • In-source fragments • Most metabolomics processing workflows fail to account for these phenomenon. • In-source phenomenon collectively result in a mass spectrum of signals representing a compound. • MASS SPECTRA are the most accurate representation of metabolites. • Prediction of spectra would require knowledge of structure and prediction of fragments. MS spectrum of Deoxycholic acid Approach : • Devise a ‘chemically blind’ clustering approach to deal with unpredictable phenomena . • Common MS signals derived from the same compound will both coelute and covary . Approach: • Covariation and coelution of two features represent strong evidence that they derive from same compound these features should be grouped together ! • ( • (OPTIONALLY ) Utilize indiscriminant MS/MS data to generate fragmentation data for every feature . • Feature similarity scores are based on product of correlational (covariance) and retention time ( coelution ) similarity . • Feature groups (including signal intensities) are representative of compound spectra that can be used for spectral matching to facilitate compound annotation . • Feature clustering enables confident recreation of mass spectra from XCMS output. • Spectra can be used for compound annotation. Results (RAMclustR): • Simultaneous reduction in complexity • Reduction in dataset wide variance Results (RAMsearch): • Feature groups export as MS spectra in *.msp format. • Enables efficient spectral searching. • Wrap NIST MSpepSearch into a GUI. Enables: Batch searching spectral libraries rapid manual validation of spectral matches Assignment of annotation confidence scores (MSI) Export of evidence, including match spectra, for import into RAMclustR Conclusions: More realistic - features are derived from compounds and feature clusters (spectra) fully represent compounds. More streamlined - ~ 5-10 fold reduction of features (RAMclustR). Batch searching is enabled (RAMsearch). More sensitive - Aggregation of features into spectra reduces analytical variance. More confidence - Several mass spectral signals offer more annotation confidence than a single accurate mass alone. Open source: https://github.com/cbroeckl/RAMClustR Anal. Chem., 2014, 86 (14), pp 6812–6817

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

RAMclust/RAMsearch: efficient post-XCMS feature clustering and annotation of MS-based metabolomics datasets.
Corey D. Broeckling and Jessica E. Prenni
Colorado State University, Proteomics and Metabolomics Facility
Acknowledgements: Asa Ben-Hur, and Fayyaz A Afsar helped to develop ramclustR algorithms. Steffen Neumann helped package ramclustRNIST (Steve Stein, David Sparkman, Dmitri Tchekhovskoi)Kevin Brown and Ben Sutton helped develop RAMsearch
Introduction: • Non-targeted profiling by UPLC-MS is a
powerful tool for metabolic profiling.
• Electrospray ionization is soft, but manysignals are generated for a givenmetabolite:
• Isotopes• Adducts• Multimers• In-source fragments
• Most metabolomics processing workflowsfail to account for these phenomenon.
• In-source phenomenon collectively result ina mass spectrum of signals representing acompound.
• MASS SPECTRA are the most accuraterepresentation of metabolites.
• Prediction of spectra would requireknowledge of structure and prediction offragments.
MS spectrum of Deoxycholic acid
Approach:• Devise a ‘chemically blind’ clustering
approach to deal with unpredictablephenomena.
• Common MS signals derived from the samecompound will both coelute and covary.
Approach: • Covariation and coelution of two features represent strong
evidence that they derive from same compound thesefeatures should be grouped together!
• (
• (OPTIONALLY) Utilize indiscriminant MS/MS data to generatefragmentation data for every feature.
• Feature similarity scores are based on product of correlational(covariance) and retention time (coelution) similarity.
• Feature groups (including signal intensities) arerepresentative of compound spectra that can be used forspectral matching to facilitate compound annotation.
• Feature clustering enables confident recreation of mass spectra from XCMS output.
• Spectra can be used for compound annotation.
Results (RAMclustR):
• Simultaneous reduction in complexity
• Reduction in dataset wide variance
Results (RAMsearch): • Feature groups export as MS spectra in
*.msp format.• Enables efficient spectral searching.• Wrap NIST MSpepSearch into a GUI.
Enables: Batch searching spectral libraries rapid manual validation of spectral
matches Assignment of annotation confidence
scores (MSI) Export of evidence, including match
spectra, for import into RAMclustR
Conclusions: More realistic - features are derived fromcompounds and feature clusters (spectra) fullyrepresent compounds.More streamlined - ~ 5-10 fold reduction offeatures (RAMclustR). Batch searching isenabled (RAMsearch).More sensitive - Aggregation of features intospectra reduces analytical variance.More confidence - Several mass spectralsignals offer more annotation confidence thana single accurate mass alone.
Open source:https://github.com/cbroeckl/RAMClustRAnal. Chem., 2014, 86 (14), pp 6812–6817
Related Documents










![Using Feature Clustering for GP-Based Feature Construction ...xuebing/Papers/... · feature selection methods [6,10,13,24]. However, applying feature clustering to FC is still limited.](https://static.cupdf.com/doc/110x72/5f41d0d40cf80c01af18e155/using-feature-clustering-for-gp-based-feature-construction-xuebingpapers.jpg)