C ARGE FLI 10. OIE Seminar, Berlin 07 th June 2013 Metagenomics using Next Genera2on Sequencing technology Mar2n Beer, Bernd Hoffmann, Ma=hias Scheuch, Dirk Höper

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
CARGE FLI
10. OIE Seminar, Berlin 07th June 2013
Metagenomics using Next Genera2on Sequencing technology
Mar2n Beer, Bernd Hoffmann, Ma=hias Scheuch, Dirk Höper
Ins%tute of Diagnos%c Virology
Preview
! Introduc2on – pathogen detec2on
! The metagenomic approach
! Challenges of metagenomics
! From sample prepara2on to NGS to data analysis
! Summary and conclusions
BTV, SBV and Usutu outbreak as „indicator“
African Horse sickness
West Nile Virus
Rift Valley Fever
EHDV ASFV
CCHFV ??????
Ins%tute of Diagnos%c Virology
How to detect the unexpected or unknown?
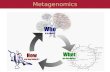
The basic definition of metagenomics is the analysis of genomic DNA from a whole community.
Gilbert JA, Dupont CL (2011). Ann Rev Mar Sci 3: 347-71. 10.1146/annurev-marine-120709-142811
What is Metagenomics (1)?
Ins%tute of Diagnos%c Virology
What is Metagenomics (2)?
Metagenomics is the applica0on of modern genomics techniques to the study of communi0es of microbial organisms directly in their natural environments, bypassing the need for isola2on and lab cul2va2on of individual species.
Chen K, Pachter L (2005). PLoS Comput Biol 1(2): e24. doi:10.1371/journal.pcbi.0010024
Ins%tute of Diagnos%c Virology
Why Do Metagenomic Pathogen Detec2on (1)?
Because every causative agent of infectious disease relies on nucleic acids (NAs) both for its genome and gene expression (only known exception so far are transmissible spongiform encephalopathies) AND Modern shotgun sequencing methods detect all NAs in a given sample with ± equal probability
Why Do Metagenomic Pathogen Detec2on (2)?
Suppose the following • We have animals suffering from an unknown disease • Standard targeted diagnos0cs do not reveal the causa0ve agent
• We expect as the causa2ve agent a pathogen containing nucleic acids
→ it is straighQorward to comprehensively sequence total NAs from these animals to detect the pathogen
For example: Roche Genome Sequencer Gs flex (up to 500 Mio. bp per run)
Next generation sequencing (NGS)
- Read: short fragments (80 to 500 bp) - Contig: larger sequence pieces assembled from reads
Key Issues for Diagnos2c Metagenomics
• Pathogen detec0on in a metagenomic dataset is finding the needle in a haystack
Ins%tute of Diagnos%c Virology
Zoom 1000 X
Key Issues for Diagnos2c Metagenomics
Example • CaZle genome: 2.97 Gbp • pCV2 genome: 1768 b → mass ra0o pCV2:CaZle = 1:1,679,864
Key Issues for Diagnos2c Metagenomics
Sensi0vity is determined by • the ra0o of the genome sizes (or in the RNA case
genome/total RNA) and • the copy numbers of the genomes (RNA molecules)
1. Sensi2vity can easily be scaled up by simply producing more sequences to yield at least one pathogen read 2. Choosing the appropriate sample material, i.e. material with high loads of pathogen (and minimum host genome) is crucial
Ins%tute of Diagnos%c Virology
Key Issues for Diagnos2c Metagenomics
• Huge datasets are generated, resul2ng in complex and 2me consuming data analysis
• Classifica0on of the sequences relies on finding similari0es to known pathogens
• Limited read length (with some instruments) impedes data analysis
• How to deal with unclassifiable sequences: are they random ar0ficial or natural but unknown sequences? These sequences require intense manual evalua2on
Ins%tute of Diagnos%c Virology
Sample Choice
-‐ Body fluid/0ssue material? -‐ Preferably the matrix with the lowest quan2ty of host nucleic acids (NA)
-‐ The highest possible quan0ty of pathogen NA -‐ DNA or RNA?
-‐ RNA is to be preferred over DNA because all replica0ng viruses generate RNA but only a few generate DNA
Ins%tute of Diagnos%c Virology
Sample Choice
Example 1 -‐ Fish are dying for unknown reason -‐ Sample from metabolically highly ac0ve gill 0ssue -‐ DNA virus expected
-‐ Sequencing library from RNA -‐ 229,000 sequencing reads -‐ 2 reads RNA virus (0.00087%)
Example 2 -‐ sequencing RNA isolated from serum from BVDV persistently infected caZle
-‐ 5% viral reads
Sample Choice
Sample Choice
Example 3 -‐ Schmallenberg virus samples RNA from serum -‐ 7/27,000 (0.026%) reads orthobunyavirus -‐ Library only sufficient to yield a total of approx. 85,000 reads
Sample Prepara2on
-‐ Usually high background of host nucleic acids -‐ Different nuclease diges0on dependent techniques for the deple0on of these: -‐ DNase SISPA -‐ Vidisca -‐ Kits for sample normaliza2on
-‐ BUT: nuclease digest means risk of irreversible informa0on loss
Sequencing
-‐ Various technologies available -‐ Read length is a cri2cal determinant -‐ Sanger with too low throughput -‐ Some plaQorms with high throughput require long sequencing 0me
For diagnos0cs 2me necessary 0ll comple0on may be an issue
Sequencing: Impact of Read Length
Success of read classifica0on depending on read length (GCReoV-‐Daten Histogramm)
Sequencing: Impact of Read Length
What if … the Schmallenberg-‐virus sequencing reads had been shorter?
Long reads Short Reads Mean length 315 96 Total No. reads 27420 27420 No. classified reads 26128 25310 No. Orthobunyavirus hits 7 1 Similarity with subject sequence 68.37 % -‐ 95.60 % 98.68 % No. unclassified reads 1292 2110
Data Analysis
A known virus/bacterium with a sufficiently similar genome or amino acid sequence is necessary The more sequences are available for diverse viruses/ organisms, the higher the chance gets to iden0fy a novel/unexpected pathogen by sequence comparison
Data Analysis
-‐ A bigger database causes longer computa0on to find the most similar sequence
-‐ Too small databases produce significant hits that are not meaningful!
-‐ The stringency of the search algorithm is crucial
Beyond detec2on: fulfillment of Koch’s postulates
-‐ Discussion is necessary about the possibility and necessity to fulfill Koch’s postulates
-‐ Failure to fulfill these might result in false diagnosis
-‐ In animal diagnos0cs fulfillment can be possible and desirable (see SBV)
Beyond detec2on: fulfillment of Koch’s postulates
Mokili -‐ Koch’s postulates • The diseased host metagenome in a sample is
significantly different from a control sample • Inocula0on of the clinical sample in a healthy subject
should result in the same disease state • Differen0al metagenomics traits iden0fied in step 1 have
to be recognized, only arer infec0on, in the metagenome of the inoculated subject
• Inocula0on of a sample of the diseased subject from step 2 must induce disease in a newly infected subject
J. L. Mokili, F. Rohwer, B. E. Du0lh, Curr. Opin. Virol. 2, 63 (2012)
Conclusions
• Metagenomics is a universal powerful tool for discovery of unexpected/unknown pathogens
• The key to success is a high rela0ve abundance of the pathogen compared to the host nucleic acid in a given sample
• In order to reduce the cost and to circumvent the boZleneck of data analysis, protocols for op0mized sample prepara0on are necessary
• Op0mized data analysis and databases are crucial
Related Documents