Synthesis of stable isotope labelled standards (SILs) hm 5 C + 3 Da m 5 C + 4 Da C + 3 Da UHPLC-MS/MS nucleoside quantification method Purified RNA is spiked with a known amount of stable isotope-labelled (SIL) standards and enzymatically digested to ribonucleosides. These are separated on an ultra-high performance liquid chromatography (UHPLC) column and quantified using tandem mass spectrometry (MS/MS). cells tissue RNA isolation enzymatic RNA digestion SIL spiking UHPLC- MS/MS N NH 2 O N O OH OH HO D 3 C D 3.8 4.0 4.2 4.4 4.6 0 250 500 750 1000 Time [min] 3.8 4.0 4.2 4.4 4.6 0 250 500 750 1000 Time [min] 3.8 4.0 4.2 4.4 4.6 0 250 500 750 1000 Time [min] N NH 2 O N H 3 C H R N NH 2 O N D 3 C D R N NH 2 O N HO R m 5 C: m/z = 258 → 126.06619 ± 5 ppm m 5 C + 4 Da: m/z = 262 → 130.09130 ± 5 ppm hm 5 C: m/z = 274 → 142.06110 ± 5 ppm NH O O N O OH OH HO N NH 2 O N O OH OH HO H 3 C NH N N O NH 2 N O OH OH HO N NH 2 O N O OH OH HO D 3 C D 4.33 min 4.04 min 4.04 min Synthesis of [2- 13 C,1,3- 15 N 2 ]-cytidine (C + 3 Da), 5-methyl-D 3 -[6-D]- cytidine (m 5 C + 4 Da), and 5-hydroxymethyl-[2- 13 C,1,3- 15 N 2 ]-cytidine (hm 5 C + 3 Da). Asterisks (*) represent a 15 N or 13 C isotopologue. Introduction More than 150 modified nucleosides exist in RNA, but the precise function and significance of nearly all of them remain unknown. [1] 5- methylcytosine (m 5 C) is one of the most abundant modifications and can influence RNA stability, processing, and protein binding. [2-4] We probe the dynamics of m 5 C and its metabolic derivatives in RNA to better understand the biological role of modified cytosine residues. Metabolism of 5-methylcytosine in RNA Sabrina M. Huber , Pieter van Delft, Eric A. Miska, Shankar Balasubramanian Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge, CB2 1EW, UK Contact: [email protected] [1] M. A. Machnicka et al., Nucleic Acids Res. 2013, 41, D262 – 267; [2] M. Schaefer et al., Genes Dev. 2010, 24, 1590 – 1595; [3] S. Hussain et al., Cell Reports 2013, 4, 255 – 261; [4] T. Amort et al., RNA Biology 2013, 10, 1 – 7. Bibliography hm 5 C is pervasive across all three domains of life hm 5 C was detected in total RNA from various model organisms. Its levels are species dependent and up to a 1000-fold lower than those of m 5 C. RNA N NH 2 O N O OH O O 13 C D D D N NH 2 O N O OH O O 13 C D HO D N NH 2 O N O OH O O 13 C O D CO 2 H NH 2 2 13 CD 3 S [Ox] [Ox] In vivo oxidation of m 5 C to hm 5 C and f 5 C Upon feeding mice with L-methionine-(methyl- 13 C,D 3 ), 13 C and D isotope incorporation was observed into m 5 C, 5-hydroxymethylcytosine (hm 5 C) and 5-formylcytidine (f 5 C), establishing them as metabolic derivatives of m 5 C. Transcription-independent formation of hm 5 C Freshly transcribed cellular RNA was labelled with 15 N 2 -cytidine and methylation and hydroxymethylation thereof was marked with 13 C and D isotopes. While m 5 C is established co-transcriptionally, its oxidation to hm 5 C boosts at a specific stage during the cell cycle. ! purification of specific RNA types and quantification of their modifications " polyA RNA, tRNA & rRNA, small RNAs ! identification of proteins binding specifically to m 5 C- or hm 5 C-modified RNA Future perspectives 0 5 10 15 0 17 34 51 68 15 N 2 13 CD 3 -m 5 C 15 N 2 -C 15 N 2 13 CD 2 -hm 5 C time [h] labelling efficiency [%] Poster: S. M. Huber et al., ChemBioChem 2015, 16, 752 – 755. N NH 2 O N H 3 C N NH 2 O N HO m 5 C hm 5 C [Ox] ? structure stability protein binding processing ??? M. jannaschii E. coli A. thaliana C. elegans Mouse brain 0.0001 0.001 0.01 0.1 1 10 m 5 C hm 5 C % C mod / ( C + m 5 C + hm 5 C) H 2 N NH 2 O H H O N H NH O O O OBz BzO BzO N NH 2 O N O OH OH HO polyphosphoric acid * * * * * * N NH O O * * * * * * N NH 2 O N O O O HO * * * HO N NH 2 O N O OH OH HO * * * HO N H NH O O D 3 C D O OH OH HO N NH NH 2 O D 3 C D O OBz BzO BzO N N NH 2 O * * * O OBz BzO BzO OAc 1. TMSCl, HMDS , TMSOTf 2. N N N P O 3 2. NH 4 OH 1. NaOMe 1. H 2 SO 4 , acetone 2. paraformaldehyde, KOH 90 % TFA NaOMe

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

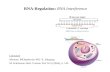
Synthesis of stable isotope labelled standards (SILs)
hm5C + 3 Da
m5C + 4 Da
C + 3 Da
UHPLC-MS/MS nucleoside quantification method
Purified RNA is spiked with a known amount of stable isotope-labelled (SIL) standards and enzymatically digested to ribonucleosides. These are separated on an ultra-high performance liquid chromatography (UHPLC) column and quantified using tandem mass spectrometry (MS/MS).
cells
tissue
RNA isolation
enzymatic RNA digestion
SIL spiking
UHPLC-MS/MS
NNH2
ON
O
OHOH
HO
D3C
D
3.8 4.0 4.2 4.4 4.60
250
500
750
1000
Time [min]3.8 4.0 4.2 4.4 4.60
250
500
750
1000
Time [min]3.8 4.0 4.2 4.4 4.60
250
500
750
1000
Time [min]
N
NH2
ON
H3C
HR
N
NH2
ON
D3C
DR
N
NH2
ON
HO
R
m5C:m/z = 258 → 126.06619 ± 5 ppm
m5C + 4 Da:m/z = 262 → 130.09130 ± 5 ppm
hm5C:m/z = 274 → 142.06110 ± 5 ppm
NHO
ONO
OHOH
HO
NNH2
ONO
OHOH
HO
H3C
NH
N
NO
NH2NO
OHOH
HO
NNH2
ON
O
OHOH
HO
D3C
D
4.33 min4.04 min4.04 min
Synthesis of [2-13C,1,3-15N2]-cytidine (C + 3 Da), 5-methyl-D3-[6-D]-cytidine (m5C + 4 Da), and 5-hydroxymethyl-[2-13C,1,3-15N2]-cytidine (hm5C + 3 Da). Asterisks (*) represent a 15N or 13C isotopologue.
Introduction
More than 150 modified nucleosides exist in RNA, but the precise function and significance of nearly all of them remain unknown.[1] 5-methylcytosine (m5C) is one of the most abundant modifications and can influence RNA stability, processing, and protein binding.[2-4] We probe the dynamics of m5C and its metabolic derivatives in RNA to better understand the biological role of modified cytosine residues.
Metabolism of 5-methylcytosine in RNA Sabrina M. Huber, Pieter van Delft, Eric A. Miska, Shankar Balasubramanian Department of Chemistry, University of Cambridge, Lensfield Road, Cambridge, CB2 1EW, UK
Contact: [email protected]
[1] M. A. Machnicka et al., Nucleic Acids Res. 2013, 41, D262 – 267; [2] M. Schaefer et al., Genes Dev. 2010, 24, 1590 – 1595; [3] S. Hussain et al., Cell Reports 2013, 4, 255 – 261; [4] T. Amort et al., RNA Biology 2013, 10, 1 – 7.
Bibliography
hm5C is pervasive across all three domains of life
hm5C was detected in total RNA from various model organisms. Its levels are species dependent and up to a 1000-fold lower than those of m5C.
hm5C in RNA may be evolutionary conserved RNA N
NH2
ON
O
OHO
O
13CD
D
D
NNH2
ON
O
OHO
O
13CD
HO
D
NNH2
ON
O
OHO
O
13C
O
DCO2H
NH2213CD3S
[Ox] [Ox]
In vivo oxidation of m5C to hm5C and f5C
Upon feeding mice with L-methionine-(methyl-13C,D3), 13C and D isotope incorporation was observed into m5C, 5-hydroxymethylcytosine (hm5C) and 5-formylcytidine (f5C), establishing them as metabolic derivatives of m5C.
Transcription-independent formation of hm5C
Freshly transcribed cellular RNA was labelled with 15N2-cytidine and methylation and hydroxymethylation thereof was marked with 13C and D isotopes. While m5C is established co-transcriptionally, its oxidation to hm5C boosts at a specific stage during the cell cycle.
! purification of specific RNA types and quantification of their modifications " polyA RNA, tRNA & rRNA, small RNAs
! identification of proteins binding specifically to m5C- or hm5C-modified RNA
Future perspectives
0 5 10 150
17
34
51
68
15N213CD3-m5C
15N2-C
15N213CD2-hm5C
time [h]
labe
lling
effi
cien
cy [%
]
Poster: S. M. Huber et al., ChemBioChem 2015, 16, 752 – 755.
N
NH2
ON
H3C N
NH2
ON
HO
m5C hm5C
[Ox]
?structure
stability
protein binding
processing???
M. jannas
chii
E. coli
A. thali
ana
C. eleg
ans
Mouse b
rain
0.0001
0.001
0.01
0.1
1
10m5C hm5C
% C
mod
/ ( C
+ m
5 C +
hm
5 C)
H2N NH2
O
HH
ONH
NH
O
O O
OBzBzO
BzO
N
NH2
ONO
OHOH
HO
polyphosphoricacid*
***
** N
NH
O
O*
**
* **
N
NH2
ONO
OO
HO * **HO N
NH2
ONO
OHOH
HO * **HO
NH
NH
O
O
D3C
D
O
OHOH
HO N
NH
NH2
O
D3C
D
O
OBzBzO
BzO N
N
NH2
O***
O
OBzBzO
BzO OAc
1. TMSCl, HMDS
, TMSOTf2.
NN
NPO
3
2. NH4OH
1.
NaOMe
1. H2SO4, acetone2. paraformaldehyde, KOH
90 % TFA
NaOMe
Related Documents