NGS LIBRARY Illumina Hi‐Seq 4000 Submission Form PO #: Please indicate special instructions (pooling schemes, combinationof projects in NOTE: one lane, etc) in the Comments section below. Email chains with Facility personnel will NOT qualify as evidence of instruction in case of disputes. UofC Clients mark “N/A” Contact Information Date (mm/dd/yyyy) Principal Investigat or Principal Investigator Email / Phone Department Cancer Center Member? Yes No Experiment Contact Experiment Contact Email / Phone Billing Administrator Billing Administrator Email / Phone Sample Preparation and Delivery AB-1 Use 1 5 mL Eppendorf tubes Simple Unique Labe Email Sample Names & Index Se Mail/Drop‐Off Prepare 13ul at 10nM “Initials‐number” [email protected]. edu 9:00am‐4:00pm M‐F Project Information Sample Species Human Mouse Rat Other: Number of Tubes Submitted: Number of Samples per Tube: Library Type: *Please note: Success of the experiment will be predicated upon properly completing the below fields: High‐Complexity Low‐Complexity (please specify RNA ChIP ATAC PCR product DNA‐Whole‐Genome DNA‐Other in comments) Please Submit Excel Sheet Listing Sample Labels and Index Sequences Number of Lanes Needed: Run Type: SR50 PE100 Index Length 6 bases 8 bases Dual (8/8) Other Index Manufacturer/Library Kit used: Comments: (please specify if libraries are non‐standard, i.e. low complexity/low nucleotide diversity, repeating elements, etc) University of Chicago | Genomics Facility ‐ Knapp Center for Biomedical Discovery (KCBD) 900 E. 57th Street, Room # 1230 Chicago, IL 60637 ACTTAC

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

NGS
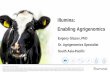
LIBRARY Illumina Hi‐Seq 4000 Submission Form PO #:
Please indicate special instructions (pooling schemes, combination of projects in NOTE: one lane, etc) in the Comments section below. Email chains with Facility
personnel will NOT qualify as evidence of instruction in case o f disputes.
UofC Clients mark “N/A”
Contact Inform
ation
Date (mm/dd/yyyy)
Principal Investigator Principal Investigator Email / Phone
Department Cancer CenterMember?Yes No
Experiment Contact Experiment Contact Email / Phone
Billing Administrator Billing Administrator Email / Phone
Sample Preparation and Delivery
AB-1
Use 1Φ5 mL Eppendorf tubes Simple Unique Labeƭǎ Email Sample Names & Index Selj Mail/Drop‐Off
Prepare 13ul at 10nM “Initials‐number” [email protected] 9:00am‐4:00pm M‐F
Project Inform
ation
Sample Species Human Mouse Rat Other:
Number of Tubes Submitted: Number of Samples per Tube:
Library Type: *Please note: Success of the experiment will be predicated upon properly completing the below fields:
High‐Complexity Low‐Complexity
(please specify RNA ChIP ATAC PCR product DNA‐Whole‐Genome DNA‐Other in comments)
Please Submit Excel Sheet Listing Sample Labels and Index Sequences
Number of Lanes Needed:
Run Type:
SR50 PE100
Index Length
6 bases 8 bases Dual (8/8) Other
Index Manufacturer/Library Kit used:
Comments: (please specify if libraries are non‐standard, i.e. low complexity/low nucleotide diversity, repeating elements, etc)
University of Chicago | Genomics Facility ‐ Knapp Center for Biomedical Discovery (KCBD) 900 E. 57th Street, Room # 1230
Chicago, IL 60637
ACTTAC
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
Gel Cut Required?
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
yes
lscarpitta
Typewritten Text
no
lscarpitta
Typewritten Text
If yes, specify size range needed:
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
NO PLATES, STRIP TUBES, OR 0.5ml TUBES!!
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
lscarpitta
Typewritten Text
*
lscarpitta
Typewritten Text
* full flowcell submissions only!
Related Documents