GENETIC IMBALANCES REVEALED BY COMPARATIVE GENOMIC HYBRIDIZATION IN OSTEOSARCOMAS Toshifumi OZAKI 1 , Karl-Ludwig SCHAEFER 2,6 , Daniel WAI 2 , Horst BUERGER 2 , Silke FLEGE 3 , Norbert LINDNER 1 , Matthias KEVRIC 3 , Raihanatou DIALLO 2,6 , Agnes BANKFALVI 2 , Christian BRINKSCHMIDT 4 , Heribert JUERGENS 3 , Winfried WINKELMANN 1 , Barbara DOCKHORN-DWORNICZAK 5 , Stefan S. BIELACK 3 and Christopher POREMBA 2,6 * 1 Department of Orthopaedics, Westfa ¨lische Wilhelms-University, Mu ¨nster, Germany 2 Gerhard-Domagk-Institute of Pathology, Westfa ¨lische Wilhelms-University, Mu ¨nster, Germany 3 Pediatric Hematology and Oncology, Westfa ¨lische Wilhelms-University, Mu ¨nster, Germany 4 Institute of Pathology, Starnberg, Germany 5 Institute of Pathology, Kempten, Germany 6 Institute of Pathology, Heinrich-Heine-University, Du ¨sseldorf, Germany Osteosarcomas are the most frequent bone sarcomas. The molecular chromosomal aberrations in osteosarcomas were analyzed by comparative genomic hybridization (CGH). We studied 47 frozen tumors (41 primary samples, 6 relapses) in osteosarcoma patients registered in the Cooperative Osteo- sarcoma Study (COSS) protocol. Genomic imbalances were detected in 40 of 41 primary tumors and 6 of 6 relapsed tumors. Gains were more frequent than losses (ratio of 1.3: 1). The median number of changes was 16 and 12 in primary and relapsed osteosarcomas, respectively. The median num- ber of aberrations in primary high-grade osteosarcomas (17.0) was significantly higher than in low- or intermediate- grade osteosarcoma subtypes (3.0) (p 0.038). The most frequent gains included 8q, 1p21-p31 and 1q21-q24, and the most frequent losses were 10q, 5q and 13q. High-level gains were observed on 8q23-q24, 17p13 and 1q21-q24. A gain of 19p (p < 0.001) or loss of 9p (p 0.027) was more frequent in poor responders than in good responders. Univariate anal- ysis revealed that patients with primary metastases (p 0.002), poor histologic responses (p 0.005), high-level gains of 19p (p 0.012) or losses of 13q14 (p 0.042) had signifi- cantly lower event-free survival (EFS), whereas patients with a loss of 5q (p 0.007) or a loss of 10q21-22 (p 0.017) had significantly higher EFS than patients without these aberra- tions. Multivariate analysis demonstrated that primary me- tastasis, loss of 13q14 and loss of 5q were independent prog- nostic factors. The findings of our study seem to be useful for evaluating the prognosis of patients and may finally lead to treatment strategies based on genetic background of osteo- sarcoma. © 2002 Wiley-Liss, Inc. Key words: osteosarcoma; chromosomal aberrations; CGH; progno- sis; genetic pathways Osteosarcoma is the most common primary malignant tumor of bone. In the 1980s and 1990s, the development of more effective imaging techniques, 1,2 chemotherapy 3–6 and surgical techniques 7,8 have improved the prognosis of patients with osteosarcoma. How- ever, the genetic background of osteosarcoma remains poorly understood. Several recent studies attempted to analyze the molecular ge- netics of osteosarcoma. Loss of heterozygosity (LOH) studies have shown a frequent LOH on chromosome arms 3q, 13q, 17p and 18q. 9 LOH of retinoblastoma 1 (RB1; 13q14) has been shown to indicate tumors with poor prognoses. 10,11 Amplification of the human homolog of mouse double minute 2 (MDM2; 12q13-q15) is reported to occur in 30% of recurrent or metastatic osteosarco- mas. 12 Amplification of DNA-damage-inducible transcript 3 (DDIT3; 12q13.1-q13-2) has also been detected in 12% of osteo- sarcomas, 13 and SAS (12q13-q14) amplification has been detected mainly in surface osteosarcomas. 14 EBRBB2 (17q21.1) expression has been detected in 42% of osteosarcomas 15 and was correlated with both histologic response to preoperative chemotherapy and poor prognosis. 16 Comparative genomic hybridization (CGH) is a useful tech- nique for a genome-wide screening of DNA sequence copy num- ber changes. 17,18 Up to now, there have been 5 series of CGH analyses on osteosarcomas. 19 –23 These studies revealed that ge- netic aberrations in osteosarcomas were extensive. 21 However, the patient numbers in previous studies were low or the treatment modality, including chemotherapy or extent of the surgical margin, was different between patients. Therefore, it was difficult to esti- mate the clinical relevance of the aberrations identified. Moreover, the frequencies of the combinations of each aberration, and their influence on clinical course, were not evaluated. In our study, we analyzed the molecular chromosomal aberrations by CGH in 47 osteosarcomas. In 41 primary tumors with detailed follow-up information, the CGH data were statistically compared to clinical data. MATERIAL AND METHODS Patients Of 52 osteosarcoma tissues obtained from 51 patients who were treated from 1996 –2000, 47 biopsy tissues were selected for our study (Table I). Two tissues (one primary tumor and one relapsed tumor) were obtained from one patient with extraskeletal osteo- sarcoma. All samples were obtained from frozen tissues, which were taken before preoperative treatment and preserved at 80°C. Tumor tissues were taken from typical and viable tumor areas with 80% tumor-cell content. The diagnosis was histologically con- firmed as high-grade osteosarcoma. Our study was conducted after human experimentation review by the ethics committee. Of 47 tumors, 41 were primary osteosarcomas (15 osteoblastic including 1 postradiation osteoblastic, 7 fibroblastic, 3 chondro- blastic, 2 parosteal, 2 teleangiectatic, 1 periosteal and 11 unclas- sified) that did not undergo any pretreatment. Two parosteal os- teosarcomas were intermediate grade and 1 periosteal Grant sponsor: Alexander von Humboldt Foundation; Grant sponsor: German Research Foundation DFG; Grant number: PO 529/5-1. The first three authors contributed equally to this work. *Correspondence to: Institute of Pathology, Heinrich-Heine-University, Moorenstrasse 5, 40225 D¨ usseldorf, Germany. Fax: 49-201-81-18353. E-mail: [email protected] Received 21 November 2001; Revised 15 April, 23 July 2002; Accepted 25 July 2002 DOI 10.1002/ijc.10709 Published online 7 October 2002 in Wiley InterScience (www. interscience.wiley.com). Int. J. Cancer: 102, 355–365 (2002) © 2002 Wiley-Liss, Inc. Publication of the International Union Against Cancer

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
GENETIC IMBALANCES REVEALED BY COMPARATIVE GENOMICHYBRIDIZATION IN OSTEOSARCOMASToshifumi OZAKI
1, Karl-Ludwig SCHAEFER2,6, Daniel WAI
2, Horst BUERGER2, Silke FLEGE
3, Norbert LINDNER1, Matthias KEVRIC
3,Raihanatou DIALLO
2,6, Agnes BANKFALVI2, Christian BRINKSCHMIDT
4, Heribert JUERGENS3, Winfried WINKELMANN
1,Barbara DOCKHORN-DWORNICZAK
5, Stefan S. BIELACK3 and Christopher POREMBA
2,6*1Department of Orthopaedics, Westfalische Wilhelms-University, Munster, Germany2Gerhard-Domagk-Institute of Pathology, Westfalische Wilhelms-University, Munster, Germany3Pediatric Hematology and Oncology, Westfalische Wilhelms-University, Munster, Germany4Institute of Pathology, Starnberg, Germany5Institute of Pathology, Kempten, Germany6Institute of Pathology, Heinrich-Heine-University, Dusseldorf, Germany
Osteosarcomas are the most frequent bone sarcomas. Themolecular chromosomal aberrations in osteosarcomas wereanalyzed by comparative genomic hybridization (CGH). Westudied 47 frozen tumors (41 primary samples, 6 relapses) inosteosarcoma patients registered in the Cooperative Osteo-sarcoma Study (COSS) protocol. Genomic imbalances weredetected in 40 of 41 primary tumors and 6 of 6 relapsedtumors. Gains were more frequent than losses (ratio of 1.3:1). The median number of changes was 16 and 12 in primaryand relapsed osteosarcomas, respectively. The median num-ber of aberrations in primary high-grade osteosarcomas(17.0) was significantly higher than in low- or intermediate-grade osteosarcoma subtypes (3.0) (p � 0.038). The mostfrequent gains included 8q, 1p21-p31 and 1q21-q24, and themost frequent losses were 10q, 5q and 13q. High-level gainswere observed on 8q23-q24, 17p13 and 1q21-q24. A gain of19p (p < 0.001) or loss of 9p (p � 0.027) was more frequentin poor responders than in good responders. Univariate anal-ysis revealed that patients with primary metastases (p �0.002), poor histologic responses (p � 0.005), high-level gainsof 19p (p � 0.012) or losses of 13q14 (p � 0.042) had signifi-cantly lower event-free survival (EFS), whereas patients witha loss of 5q (p � 0.007) or a loss of 10q21-22 (p � 0.017) hadsignificantly higher EFS than patients without these aberra-tions. Multivariate analysis demonstrated that primary me-tastasis, loss of 13q14 and loss of 5q were independent prog-nostic factors. The findings of our study seem to be useful forevaluating the prognosis of patients and may finally lead totreatment strategies based on genetic background of osteo-sarcoma.© 2002 Wiley-Liss, Inc.
Key words: osteosarcoma; chromosomal aberrations; CGH; progno-sis; genetic pathways
Osteosarcoma is the most common primary malignant tumor ofbone. In the 1980s and 1990s, the development of more effectiveimaging techniques,1,2 chemotherapy3–6 and surgical techniques7,8
have improved the prognosis of patients with osteosarcoma. How-ever, the genetic background of osteosarcoma remains poorlyunderstood.
Several recent studies attempted to analyze the molecular ge-netics of osteosarcoma. Loss of heterozygosity (LOH) studies haveshown a frequent LOH on chromosome arms 3q, 13q, 17p and18q.9 LOH of retinoblastoma 1 (RB1; 13q14) has been shown toindicate tumors with poor prognoses.10,11 Amplification of thehuman homolog of mouse double minute 2 (MDM2; 12q13-q15) isreported to occur in 30% of recurrent or metastatic osteosarco-mas.12 Amplification of DNA-damage-inducible transcript 3(DDIT3; 12q13.1-q13-2) has also been detected in 12% of osteo-sarcomas,13 and SAS (12q13-q14) amplification has been detectedmainly in surface osteosarcomas.14 EBRBB2 (17q21.1) expressionhas been detected in 42% of osteosarcomas15 and was correlatedwith both histologic response to preoperative chemotherapy andpoor prognosis.16
Comparative genomic hybridization (CGH) is a useful tech-nique for a genome-wide screening of DNA sequence copy num-ber changes.17,18 Up to now, there have been 5 series of CGHanalyses on osteosarcomas.19–23 These studies revealed that ge-netic aberrations in osteosarcomas were extensive.21 However, thepatient numbers in previous studies were low or the treatmentmodality, including chemotherapy or extent of the surgical margin,was different between patients. Therefore, it was difficult to esti-mate the clinical relevance of the aberrations identified. Moreover,the frequencies of the combinations of each aberration, and theirinfluence on clinical course, were not evaluated. In our study, weanalyzed the molecular chromosomal aberrations by CGH in 47osteosarcomas. In 41 primary tumors with detailed follow-upinformation, the CGH data were statistically compared to clinicaldata.
MATERIAL AND METHODS
PatientsOf 52 osteosarcoma tissues obtained from 51 patients who were
treated from 1996–2000, 47 biopsy tissues were selected for ourstudy (Table I). Two tissues (one primary tumor and one relapsedtumor) were obtained from one patient with extraskeletal osteo-sarcoma. All samples were obtained from frozen tissues, whichwere taken before preoperative treatment and preserved at �80°C.Tumor tissues were taken from typical and viable tumor areas with�80% tumor-cell content. The diagnosis was histologically con-firmed as high-grade osteosarcoma. Our study was conducted afterhuman experimentation review by the ethics committee.
Of 47 tumors, 41 were primary osteosarcomas (15 osteoblasticincluding 1 postradiation osteoblastic, 7 fibroblastic, 3 chondro-blastic, 2 parosteal, 2 teleangiectatic, 1 periosteal and 11 unclas-sified) that did not undergo any pretreatment. Two parosteal os-teosarcomas were intermediate grade and 1 periosteal
Grant sponsor: Alexander von Humboldt Foundation; Grant sponsor:German Research Foundation DFG; Grant number: PO 529/5-1.
The first three authors contributed equally to this work.
*Correspondence to: Institute of Pathology, Heinrich-Heine-University,Moorenstrasse 5, 40225 Dusseldorf, Germany. Fax: �49-201-81-18353.E-mail: [email protected]
Received 21 November 2001; Revised 15 April, 23 July 2002; Accepted25 July 2002
DOI 10.1002/ijc.10709Published online 7 October 2002 in Wiley InterScience (www.
interscience.wiley.com).
Int. J. Cancer: 102, 355–365 (2002)© 2002 Wiley-Liss, Inc.
Publication of the International Union Against Cancer
TA
BL
EI
–PA
TIE
NT
INFO
RM
AT
ION
Cas
enu
mbe
rA
ge(y
ears
)Se
xSi
teT
umor
mat
eria
lsSu
btyp
eT
umor
Vol
ume
(mL
)
Prim
ary
Met
asta
sis
Che
mot
hera
pyO
pera
tion
Surg
ical
mar
gin
His
tolo
gic
regr
essi
ongr
ade
Rel
apse
Rel
apse
-fre
epe
riod
(mon
ths)
Out
com
e(m
onth
s)
135
Fem
ale
Dis
tal
Fem
urPr
imar
yO
steo
blas
tic90
No
CO
SS96
Am
puta
tion
Wid
e4
Yes
1516
DO
D2
17Fe
mal
eD
ista
lul
naPr
imar
yFi
brob
last
ic6
No
CO
SS96
Res
ectio
nW
ide
4N
o27
40C
DF
36
Mal
ePr
oxim
alT
ibia
Prim
ary
?12
No
CO
SS96
Res
ectio
nW
ide
3N
o37
37C
DF
414
Fem
ale
Prox
imal
Hum
erus
Prim
ary
Tel
eang
iect
atic
30N
oC
OSS
96R
esec
tion
Wid
e2
No
3636
CD
F5
15Fe
mal
ePr
oxim
alT
ibia
Prim
ary
?—
No
CO
SS96
Res
ectio
nW
ide
2N
o33
33C
DF
613
Fem
ale
Prox
imal
Tib
iaPr
imar
yO
steo
blas
tic—
Yes
CO
SS96
Am
puta
tion
Rad
ical
—Y
es23
25D
OD
719
Mal
ePr
oxim
alFe
mur
Prim
ary
?34
9N
oC
OSS
96R
esec
tion
Wid
e5
No
3535
CD
F8
42Fe
mal
eIl
ium
-Sac
rum
Prim
ary
Ost
eobl
astic
—Y
esC
OSS
96?
—Y
es16
27D
OD
921
Mal
ePr
oxim
alT
ibia
Prim
ary
Fibr
obla
stic
209
No
CO
SS96
Res
ectio
nW
ide
5Y
es3
33N
ED
1013
Fem
ale
Dis
tal
Fem
urPr
imar
y?
80N
oC
OSS
96R
esec
tion
Wid
e2
No
3434
CD
F11
20M
ale
Dis
tal
Fem
urPr
imar
yO
steo
blas
tic—
Yes
CO
SS96
Am
puta
tion
Wid
e4
Yes
724
NE
D12
16Fe
mal
ePr
oxim
alFe
mur
Prim
ary
Ost
eobl
astic
1—
No
CO
SS96
Res
ectio
nW
ide
5Y
es34
36N
ED
1320
Fem
ale
Iliu
m-S
acru
mPr
imar
y?
—Y
esC
OSS
96R
esec
tion
Wid
e5
Yes
1220
DO
D14
33M
ale
Dis
tal
Fem
urPr
imar
yO
steo
blas
tic44
9N
oC
OSS
96R
esec
tion
Wid
e5
Yes
1834
NE
D15
24M
ale
Dis
tal
Rad
ius
Prim
ary
?39
No
CO
SS96
Res
ectio
nW
ide
2N
o30
30C
DF
1653
Mal
ePr
oxim
alFe
mur
Prim
ary
Tel
eang
iect
atic
—N
oC
OSS
96A
mpu
tatio
nW
ide
4Y
es18
30D
OD
1715
Fem
ale
Dis
tal
Fem
urPr
imar
yPa
rost
eal
—Y
esC
OSS
96R
esec
tion
Wid
e—
No
2929
CD
F18
13Fe
mal
eD
ista
lFe
mur
Prim
ary
?73
4N
oC
OSS
96R
otat
ionp
last
yW
ide
2N
o28
28C
DF
1916
Mal
eD
ista
lFe
mur
Prim
ary
Fibr
obla
stic
266
No
CO
SS96
Res
ectio
nW
ide
2N
o27
27C
DF
2028
Mal
ePr
oxim
alH
umer
usPr
imar
yO
steo
blas
tic98
No
CO
SS96
Res
ectio
nW
ide
2N
o25
25C
DF
216
Mal
eD
ista
lFe
mur
Prim
ary
?63
No
CO
SS96
Rot
atio
npla
sty
Wid
e3
Yes
1116
DO
D22
29Fe
mal
eD
ista
lFe
mur
Prim
ary
Pstr
oste
al—
No
CO
SS96
2R
esec
tion
Wid
e—
Yes
2424
AW
D23
16M
ale
Dis
tal
Fem
urPr
imar
y?
316
Yes
CO
SS96
Rot
atio
npla
sty
Wid
e4
Yes
1529
NE
D24
35M
ale
Dis
tal
Tib
iaPr
imar
yO
steo
blas
tic—
No
CO
SS96
Res
ectio
nW
ide
5N
o27
27C
DF
2517
Mal
ePr
oxim
alT
ibia
Prim
ary
Fibr
obla
stic
51N
oC
OSS
96R
esec
tion
Wid
e2
No
2626
CD
F26
16Fe
mal
ePr
oxim
alH
umer
usPr
imar
yO
steo
blas
tic—
Yes
CO
SS96
Res
ectio
nW
ide
3Y
es8
21N
ED
279
Mal
eD
ista
lFe
mur
Prim
ary
Ost
eobl
astic
105
No
CO
SS96
Rot
atio
npla
sty
Wid
e3
No
2424
CD
F28
34M
ale
Dis
tal
Fem
urPr
imar
yO
steo
blas
tic25
1N
oC
OSS
96R
esec
tion
Wid
e1
No
2424
CD
F29
15Fe
mal
ePr
oxim
alT
ibia
Prim
ary
Peri
oste
al-G
3—
No
CO
SS96
Res
ectio
nW
ide
3N
o26
26C
DF
3012
Mal
eD
ista
lFe
mur
Prim
ary
Ost
eobl
astic
92N
oC
OSS
96A
mpu
tatio
nW
ide
5N
o26
26C
DF
3118
Mal
ePr
oxim
alH
umer
usPr
imar
yO
steo
blas
tic—
No
CO
SS96
Res
ectio
nW
ide
3N
o24
24C
DF
3214
Fem
ale
Dis
tal
Tib
iaPr
imar
yFi
brob
last
ic—
No
CO
SS96
Res
ectio
nW
ide
1N
o24
24C
DF
3322
Mal
ePr
oxim
alT
ibia
Prim
ary
Fibr
obla
stic
—Y
esC
OSS
96R
esec
tion
Wid
e4
No
2424
CD
F34
15M
ale
Dis
tal
Fem
urPr
imar
y?
—N
oC
OSS
96R
esec
tion
Wid
e—
No
1919
CD
F35
10Fe
mal
ePe
lvis
-Sac
rum
-L5
Prim
ary
Cho
ndro
blas
tic—
No
CO
SS96
Res
ectio
nIn
tral
esio
nal
—Y
es0
40N
ED
3614
Mal
ePr
oxim
alFi
bula
Prim
ary
Ost
eobl
astic
—N
oC
OSS
96R
esec
tion
Wid
e2
No
3636
CD
F37
11M
ale
Prox
imal
Fibu
laPr
imar
yC
hond
robl
astic
39N
oC
OSS
96R
esec
tion
Wid
e—
No
3132
CD
F38
5M
ale
Prox
imal
Tib
iaPr
imar
yFi
brob
last
ic—
No
CO
SS96
Res
ectio
nW
ide
1N
o15
15C
DF
3916
Fem
ale
Prox
imal
Hum
erus
Prim
ary
Cho
ndro
blas
tic47
No
CO
SS96
Res
ectio
nW
ide
3N
o12
12C
DF
4012
Fem
ale
Prox
imal
Fem
urPr
imar
yO
steo
blas
tic—
No
CO
SS96
Res
ectio
nW
ide
5N
o10
10C
DF
4115
Mal
ePr
oxim
alT
ibia
Prim
ary
?49
No
CO
SS96
Res
ectio
nW
ide
3N
o43
43C
DF
4240
Mal
eD
ista
lFe
mur
Met
asta
sis
Cho
ndro
blas
tic—
Yes
CO
SS86
Res
ectio
nW
ide
5Y
es24
153
NE
D43
51Fe
mal
eD
ista
lFe
mur
Met
asta
sis
Ost
eobl
astic
—N
oC
OSS
96A
mpu
tatio
nW
ide
5Y
es23
36N
ED
4433
Fem
ale
Prox
imal
Tib
iaM
etas
tasi
s?
—N
oC
OSS
86A
mpu
tatio
nR
adic
al4
Yes
127
149
DO
D45
53M
ale
Dis
tal
Fem
urR
ecur
renc
eE
xtra
skel
etal
—N
oC
OSS
96R
esec
tion
Wid
e—
Yes
1324
DO
D46
55M
ale
Pubi
sM
etas
tasi
sO
steo
blas
tic—
No
PIA
**R
esec
tion
Wid
e4
Yes
2640
NE
D47
9M
ale
Dis
tal
Fem
urR
ecur
renc
eO
steo
blas
tic—
No
CO
SS86
CR
esec
tion
Wid
e2
Yes
3454
DO
D
CD
F:C
ontin
uous
dise
ase-
free
;N
ED
:N
oev
iden
ceof
dise
ase;
AW
D:
Aliv
ew
ithdi
seas
e;D
OD
:de
ath
ofdi
seas
e.–1
Post
radi
atio
nos
teos
arco
ma.
–2N
otre
gist
ered
inC
OSS
.
356 OZAKI ET AL.
osteosarcoma was a low-grade tumor; however, all 3 tumors ex-hibited intramedullary infiltration). The 1 postradiation osteoblas-tic osteosarcoma developed in the femur 2 years after 45-Grayirradiation to a Ewing sarcoma that originated in the ilium. Sixrelapsed tumors (3 osteoblastic, 1 chondroblastic, 1 extraskeletaland 1 unclassified osteosarcoma) included 4 metastases and 2 localrecurrences. The relapsed tumors developed from 8–141 months(median � 27 months) after surgical excision of the primarytumors.
Among the 41 patients with primary tumors, the male to femaleratio was 23 to 18 (1.3) and their ages ranged from 5–53 years(median age 16 years). Twelve tumors were proximally located (5proximal humerus, 3 pelvis, 3 proximal femur and 1 proximalthigh) and 29 were distally located (14 distal femur, 11 tibia, 2fibula, 1 radius and 1 ulna). In 21 of 41 patients with primarytumors, tumor volume could be evaluated according to the methodby Gobel et al.24 Tumor volume ranged from 6–734 mL; themedian volume was 90 mL. Eight patients had tumors with largevolumes (�100 mL) and 13 patients had tumors with small vol-umes (�100 mL). Eight of 41 patients with primary tumors hadmetastases at diagnosis.
In 41 patients with primary tumors, 40 patients were registeredin the Cooperative Osteosarcoma Study (COSS); however, all 41patients were reported to have received chemotherapy according tothe current COSS 96 protocol. In this protocol, both pre- andpostoperative chemotherapy were scheduled. All patients weredesignated to receive high-dose methotraxate with leucovorin res-cue, doxorubicin, cisplatinum and ifosfamide. In a high-risk group,etoposide and carboplatin were added as a risk-stratified therapy.
As a local treatment, 40 patients underwent tumor excision andno information was available for one patient. The surgical marginwas classified according to the method by Enneking et al.7:1radical, 38 wide, 0 marginal, and 1 intralesional. The surgicalspecimens were examined histologically and classified into 6 cat-egories of regression grades according to the criteria published bySalzer-Kuntschik et al.25 Grades 1–3 were classified as goodresponses and grades 4–6 were designated poor responses. Infor-mation on the histologic effect of preoperative chemotherapy wasavailable for 35 patients: 20 were classified as good (grades 1–3)and 15 as poor responders (grades 4–6). The follow-up periodranged from 8–43 months (median 24 months).
Comparative genomic hybridization (CGH)Reference DNA from healthy blood donors and tumor DNA
were labeled by the nick translation method with digoxigenin-11-dUTP (Boehringer Mannheim, Mannheim, Germany) and biotin-14-dATP (Gibco BRL, Gaithersburg, MD), respectively. The hy-bridization was performed as described by Kallioniemi et al.26
with some modifications.27,28
Normal lymphocyte metaphase preparations were denatured at73°C for 5 min in formamide sodium (70% formamide/2� SSC,pH 7), dehydrated and treated with proteinase K (0.1 �gml-1 in 20mM tris-HCl/2 mM calcium chloride, pH 7) at 37°C for 6 min anddehydrated again. The probe mixture, after ethanol precipitationand resuspension in 10 �l of 50% formamide/10% dextran sul-phate/2� saline sodium citrate (SSC), was denatured at 75°C for5 min, applied to the slides and hybridized for 3 days at 37°C. Thehybridization was analyzed using an Olympus fluorescence micro-scope and the ISIS digital image analysis system (MetaSystems,Altlussheim, Germany) based on a high-sensitivity interratingmonochrome CCD camera and an automated CGH analysis soft-ware package. Ratio profiles were averaged from 10 metaphasesper sample (up to 20 chromosome homologues). Gains of DNAsequences were defined as chromosomal regions with a fluores-cence ratio above 1.25 and losses as regions with a ratio below0.75. A positive control with known aberrations and a negativecontrol were included in each CGH experiment as quality controls.Overrepresentations were considered to be high-level gains whenthe fluorescence ratio exceeded 1.5. Heterochromatic regions near
the centromeres and the entire X and Y chromosomes were ex-cluded from the analysis. Judgement was based on a consensus ofat least 2 of 3 authors in all cases without reference to the patient’sclinical information.
Interphase cytogenetics by 2-color FISH analysisParaffin-embedded osteosarcoma cell line pellets (cell lines:
OST, MG-63, U2OS) and paraffin-embedded specimens from 2primary high-grade osteosarcomas with losses of chromosome 13qand 19p in CGH analysis were analyzed by 2-color FISH with achromosome13q32-q33-specific DNA probe in combination with achromosome 2 �-satellite probe, a chromosome 13q14-specificDNA probe in combination with a chromosome 13,21 satelliteprobe and a TEL 19p DNA probe in combination with a chromo-some 1,15,19 satellite probe. In brief, slides were pretreated with2� SSC, pH 7.0 at 37°C for 30 min, dehydrated in 70%, 80% and95% ethanol for 2 min each and denatured in 70% formamide/2�SSC, pH 7.0 at 70°C for 2 min, followed by dehydration in cold(�20°C) 70%, 80% and 95% ethanol for 2 min each. For hybrid-ization, the prewarmed probes were incubated o/n at 37°C in ahumidified chamber. After posthybridization washes according tothe manufacturer’s description, detection was carried out as fol-lows: for the chromosome 13q probe (Quint-Essential 13-specificDNA probe, human chromosome assignment: 13q32-q33, locus:D13S585; Appligene Oncor, Illkirch, France) 60 �l of FITC-labeled anti-Digoxenin were used for incubation at 37°C. For thealpha-satellite chromosome 2 probe (chromosome 2 �-satellite[D2Z], Appligene Oncor) 30 �l of Texas Red labeled Avidin wereused for incubation at 37°C. The chromosome 13q14 (13S272)probe and the chromosome 13,21 satellite probe (QBiogene, Hei-delberg, Germany) are each direct-labeled with Rhodamine (Exc.max. 565 nm, Em. max. 590 nm); therefore, hybridization wascarried out on separate slides of consecutive tissue cuttings fromthe paraffin blocks. The TEL 19p DNA probe is direct-labeled(green, FITC spectrum) (QBiogene). The chromosome 1,5,19 sat-ellite probe is direct-labeled with Fluorescein (Exc. 495 nm, Em.520 nm) (QBiogene). For microscopy, the cells were counter-stained with DAPI (final concentration 0.1 �g/ml in Antifade).
Immunohistochemistry for p53-protein accumulationA mouse monoclonal antibody directed against the p53 nuclear
phosphoprotein (Clone Bp53-11; Ventana Medical Systems, Tuc-son, AZ) was used for immunohistochemistry of formalin-fixedand paraffin-embedded osteosarcoma tissues on a Ventana auto-mated slide-staining device. In brief, deparaffinized 4 �m sectionson poly-L-lysine compound coated slides were incubated with theanti-p53 primary antibody for 25 min at 37°C. Afterwards, abiotin-conjugated secondary antibody was first added followed byan avidin/streptavidin-enzyme conjugate. The specific antibody-secondary antibody-avidin/streptavidin-enzyme complex was vi-sualized using a precipitating enzyme reaction product readilydetected by light microscopy. For verification of the immunohis-tochemical results, all samples were incubated with anti-p53 pro-tein monoclonal antibody, clone DO7; (Novocastra Laboratories,Newcastle, UK). The presence of nuclear staining was counted aspositive.
Statistical analysisSignificance of differences in the ratios between or among
groups was evaluated by the 2 test with/without Fisher’s correc-tion. The Mann-Whitney U-test evaluated differences of the meanrank between 2 groups. Event-free survival (EFS) was defined asthe time from diagnosis until the first occurrence of an event(death, relapse or disease progression). One patient treated byintralesional surgery (residual tumor tissue) was counted as arelapsed case in further follow-up. The cumulative probability ofEFS was calculated by univariate analysis with the Kaplan-Meiermethod. Tests of the difference between survival curves werecarried out using the log-rank test. All values, which were signif-icant in the univariate analysis, were entered and reanalyzed in the
357CHROMOSOMAL INSTABILITIES IN OSTEOSARCOMAS
TABLE II – CHROMOSOMAL ABERRATIONS DETECTED BY CGH IN 47 OSTEOSARCOMAS
Case Material CGH results1
1 Primary G 11 �1p31-q25/��1p31-p22/��1q22-q25, ��4p15.3-q32, �5p, �5q21-5q22, �6p21.3-6q16/��6pcent-p21.2, �8q12-q22, ��9p22-pter, �10pter-10q11.2, �10q24-q25, �13q14-q22/��q21, �20p12
L 16 �1p33-pter, �1q32-qter, �2qcent-q21, �3p14-p21, �3q21, �6q24-qter, �7q34-qter, �11p15-pter,�11q13, �12q23-qter, �13q33, �15q23, �16, �17, �19, �22q
2 Primary G 9 �1p22-p31, �4qcent-q22/��q12, �6p, ��8q, ��11p11.1-p12, �14q23-qter, ��17p,��19q13.1,�20q12-qter
L 7 �1q41-qter, �2q34-qter, �3p22-pter, �5q, �6q16-21, �8p, �13q3 Primary G 6 �1p,�5pter-q21, ��8q, �9q21, �17q22-qter, �21q
L 4 �3q1, �10q21, �12, �17p4 Primary G 4 ��6p12-21.3, �8q, ��12q13-15, �14q
L 1 �2q24-qter5 Primary G 11 �1p31-p32, �1q21-25, �2p24-pter, �2q31, �3p22-pter, �3q24-qter, �5p, �8q11.2-q21.3, �9q21-31,
�18, �21qL 7 �5q32-qter, �6, �7pcent-p15, �10, �13q, �16, �19p
6 Primary G 12 �1p31-q25/��1p22-p31, �2pcent-p13, �4q12-q24, �5p, �6p21, �6q23-qter, �7q11.2-q21, �8q, �12p,�13q12, �16q, �17p
L 7 �4p15.3-p16, �5q32-qter, �7p12-p15, �9q31-qter, �10q, �11q23-qter, �16p7 Primary G 18 �1, �2, �3q13.1-qter, �4, ��5p, �6p, �7q11.2-q21, �8, �11q, �12, �13q21-q31, �14q23-qter,
��15q23-qter, �16q22-qter, �17p12, �19, �20, ��21q22,L 10 �3p21-23, �5q12-q23, �6q12-q24, �7p, �9, �10, �16p, �17q, �18qcent-q21.3, �22q
8 Primary G 7 �2q22-qter, �5, �7, �8, �19q, �20, �21qL 5 �3qcent-q13.2, �12p, �13q12-q13, �14q, �18q
9 Primary G 14 �2p14-q32/��q12-q31, �4, ��6p2, �6q23-qter, �7q/��7q11.2-32, �8, �9q21-33, �11pter-q13,��12p, �12q15-qter, �14q, �17, ��20p, �22q13
L 11 �2p23-pter, �3, �5, �6q12-16, �7p, �9p22-pter, �10, �11q21-qter, �13, �15q, �1810 Primary G 13 �1, �2, �4q13-31.2, �6p, ��7p21-pter, ��7q22-q31, �8p, �9p, ��11q14-qter, �12p, ��13q13,
��17p, �20L 10 �3pter-p14, �5q, �6q16-qter, �7p14-q11.2, �10, �11p12-p13, �12q, �13q21-q22, �16p, �17q
11 Primary G 12 �3q22-qter, �4q, �5pter-q15-/��p14, �6p12-p21.2, �7p, �8q, �9q21-qter, ��17p, �18q, ��19p,��20, �21q
L 10 �1q41-qter, �3p, �4p, �6q, �9p21-pter, �10, �12q, �13q, �16, �19q13.212 Primary G 5 �8q, ��12p13, �13q31-qter, ��19p13, �21q
L 2 �8p12-pter, �1013 Primary G 9 �1p34-pter, �1q41-qter, �2qcent-q32, �8q, �12/��12q15, �14q, �17, ��19p, �20p
L 8 �1p13-p22, �3p21-pter, �4qcent-q24, �5q, �10q, �13q, �15qcent-q15, �16q22-qter14 Primary G 3 �5q31, �20q12-13.2, �22q12
L 015 Primary G 10 �1p32-q24/p21-p31, �3q, �4/��4q12-21/��4q32-qter, ��6p12-p21.3, �8q, �13q31 33, �14q, �17p,
�18q, �21q22L 10 �1q32-qter, �2q, �3q13.1-qter, �5q3, �8p, �10, �11p15-pter, �15q21-22, �16q, �19q
16 Primary G 0L 0
17 Primary G 3 �12q12-q22/��12q14, ��18p, �19pL 0
18 Primary G 7 �1p31-q24, �6, �7q22, �8/��q12-21, �9p23-pter, �12p, �13q21-qterL 9 �1q31-32, �2q34-qter, �5q11.2-q33, �9q, �10cent-q23, �1ip, �16, �19, �22q
19 Primary G 11 �1p22-qter, �2p, �5p13, ��7p15-pter, ��8q, ��9p22-pter, �12p, �13q14-31, �14q11.2-q2 2,��17p, ��19qcent-q13.3
L 8 �2q34-qter, �3, �5, �6q21-qter, �10, �11p, �16p, �17q12-2120 Primary G 4 �1p31-qter, �4q13-q27, �5p, �8q22
L 6 �2q34-qter, �5q32-qter, �6q24-qter, �9q33-34, �16p, �19p21 Primary G 5 �1p31-q31, �2q14.1-q32, �5q21, �8q22-qter/��8q23, �17q22-q23
L 5 �10q24-qter, �16, �17p, �19, �22q22 Primary G 5 �1p31-pter, �4q, ��8q, ��11q13-22, ��12q12-q15
L 5 �1p13-p22, �3p, �8p, �13q, �15qcent-q2423 Primary G 13 �1pter-q25/��1q22-q24, �2p, �2q24, �3pcent-p14, �4q, �7, �8qcent-q23, ��9p21-p23, �11q13-22,
�12p11.2-12, ��17p, �19p, �20q12-qterL 8 �1q43-qter, �2q34-qter, �3p22-pter, �4p15.1-pter, �6qcent-q22, �10q23-qter, �13q, �19q13.3-q13.4
24 Primary G 10 �1p33-pter, �1q22-qter, �7p15-p21, �7q32-qter, �9q, �12q13, �12q23-q24.1, �19p, �20q,�22qL 8 �2q21-qter, �5q, �6pter-q16, �10qcent-q22, �11, �12p12-pter, �13q, �18q22-q23
25 Primary G 2 �1p33-pter, �8qL 1 �10pter-q21
26 Primary G 10 �1p33-q25/��q21-24, �3p23-pter, �4qcent-q22, �6p12-p21.3, �7p21-pter, �8q, ��9p, �16q22-q23,�20, �22q11.2-q12
L 7 �2q14.3-qter, �6q14-23, �7q21-qter, �11q13-q22, �12q, �13qcent-q22, �18q21.1-qter27 Primary G 10 �1pter-q21/��1p22, �3q25, �4q, �5p13-p14, �6p21.1-p21.2, �7, �8q/��q21.3-qter, �13q31-qter,
�17p, ��17q23-q24L 6 �1q32-qter, �2q33-qter, �3q13.3-22, �4p15.3-pter, �10, �20p12-pter
28 Primary G 11 �1pter-q25, �2pter-q34, �3q24-qter, �4, �5p, ��8q11.2, �9p21-pter, ��13q14, �14q, ��17p,��21qcent-q22
L 11 �1q32-qter, �5q, �6p22-pter, �8p, �9q31-qter, �10q, �12p, �13q31-qter, �16q12.2-qter, �19q13.2-q13.3, �22q
29 Primary G 1 �20q12-13.2
358 OZAKI ET AL.
multivariate method with a stepwise regression test. Only p-values�0.05 are given in the text. The statistical software used was StatView version 5.0.
RESULTS
Incidence of genomic imbalances in 47 casesGenomic imbalances were detected in 40 of 41 (98%) primary
tumors and 6 of 6 (100%) relapsed tumors (Table II). Gains(median 9.0, range 0–18) were more frequent than losses (median7.1, range 0–16). Many chromosomal aberrations were observed,especially gains of chromosome 1, 4q, 5p, 6p, 7, 8q, 12p, 17, 20qor 21q and losses of chromosome 2q, 5q, 6q, 10, 13q or 16.High-level gains were frequently noted in 1p21-p31, 1q22-q24,8q23-q24, 17p13 or 19p (Fig. 1).
To confirm the CGH data by interphase cytogenetics, FISH forcentromere 2 and 13q32-q33 as well as 13q14 and chromosomes13,21 satellite and 19pter and chromosome 1,5,19 satellite wereperformed on 2 primary osteosarcomas with loss of 13q and gain
of 19p in CGH analysis: both tumors revealed aneuploidy com-bined with loss of 13q32-q33 and 13q14 and gain of 19pter. Basedon 50 tumor cell nuclei in each case, the following results wereobtained: case 11, centromere 2 signals: 2.6 1.0573 (range 0–5),13q32-q33 signals: 1.075 0.7642 (range 0–3), chromosomes13,21 satellite signals: 5.7 1.943 (range 1–9), chromosome13q14 signals: 1.167 0.617 (range 0–3), chromosome 1,5,19satellite signals: 7.977 4.014 (range 3–15), chromosome 19ptersignals: 8.741 2.322 (range 2–12); case 13: centromere 2 sig-nals: 3.0 1.2609 (range 1–7), 13q32-q33 signals: 1.225 0.6597 (range 0–2), chromosomes 13,21 satellite signals: 6.1 1.893 (range 2–9), chromosome 13q14 signals: 1.404 0.433(range 0–3), chromosome 1,5,19 satellite signals: 8.235 3.873(range 0–13), chromosome 19pter signals: �10 with high unspe-cific background in 2 independent hybridizations. The results werefurther confirmed by FISH in 3 osteosarcoma cell lines (OST,MG-63, U2OS) for which we previously obtained data from CGH,24-color multiplex fluorescence in situ hybridization (M-FISH)and conventional cytogenetics (data not shown).
TABLE II – CHROMOSOMAL ABERRATIONS DETECTED BY CGH IN 47 OSTEOSARCOMAS (CONTINUED)
Case Material CGH results1
30 Primary G 4 �12p, �17p, �19p, �22q12L 3 �2q32-34, �6, �9p21-pter
31 Primary G 17 �1pter-1q25/��p34/��q21-24, ��2p, �4pter-q12/��pcent-p14, ��4q28-31.2, �6p21, ��7p21-pter,�7q31, �9q22-qter, ��10p12-15, �11p, �13q12-13, ��15q22-24, �17q/��q24-25, �18p, �19q,�20q, �21q
L 16 �1q32, �2q22-qter, �3p, �5q15-qter, �6p23-pter, �6q16-21, �7p14-q11.2, �8p, �9p, �10q, �11q,�13q21-qter, �14q21-qter, �16p13, �16q12.2-qter, �17p
32 Primary G 9 ��1pter-q23, �3p13-p21, �5pter-p14, �6p2, �9pter-q22/��pter-p22/��cent-q22, ��12p13-pter,��13q31-qter, �19q31-32,�22q12
L 9 �1q31-qter, �2q31-qter, �5q22-qter, �9q31-qter, �10q23-qter, �11q23-qter, �13q12-q21, �17p, �20p33 Primary G 9 �1pter-q24, �5p13, �6p21, �8q/��q21.3-qter, �9q13-qter, �18p, �19q, �20q, �22q
L 5 �2q32-qter, �5q14-qter, �6q22, �13q13-qter, �18q234 Primary G 12 �1pter-q24, �3p, ��5p, �8, �9, �11q, �12p, �14q24-qter, �17/��p/��q24-qter, �18p,
��19p,��21qL 9 �1q31-qter, �2q24-q36, �3q22-qter, �5q, �6, �10, �1ip, �13q12-q33, �16q12.2-qter
35 Primary G 4 �1q21-22, �8q/��q22-qter, ��17p, ��19pL 4 �1q25-qter, �3p13-pter, �13q, �15qcent-q23
36 Primary G 15 �1p36.1-q25/��1q21-23, �3q, �4qcent-27, ��6p, �7/��q22-qter, �8q, �11pter-q22, �12, ��13q31-qter, �14q, �17/��17p, �18q, �19q, �20q, �21q
L 10 �2, �3p21-pter, �4p, �4q31.2-qter, �5, �6q, �8p, �9q31-33, �10, �13qcent-1437 Primary G 12 �1pter-q25/��p21/��q23-24, �5pter-q12, �6p, �7p, �8pter-q21, ��9q21-31, ��13q32-qter,
��15q25-qter, �17p, �19p, �20p, �21qL 9 �2q, �3p21-23, �4p15.1-pter, �5q32-qter, �6q, �10q, �12q, �13q12-14, �22q
38 Primary G 9 �1p31-q31, �3q24, �5p, �6q, �7p, ��8q, �15q25-qter, �17q22-qter, �21qL 8 �1q41-qter, �2q, �4p, �5q, �10, �13q, �16, �22q
39 Primary G 15 �1p32-q25/��p31-22/��q21-q25, �2p, �3p, �3q13, �3q26.3-qter, 4q/��q24-28, �5p �6p, �6q24-qter, �7pter-q22, �8pter-q23, �12q21, �14q, �17q, �18q
L 12 �1q32-qter, �2q, �3q23, �5q, �6q14-21, �7q35-36, �10, �11, �12p13, �13q, �15, �1640 Primary G 15 �1, �2q31-q32, �4q, �5q2, �6p, �7p, ��8q23-q24.1, �10qcent-q11.2, �11,��17p, �17q22-qter,
�18, ��19pter-q13.1, �20q, �21q,L 15 �2p22-pter, �3pter-q13.1, �5q12-q14, �5q31-qter, �6q, �7q, �8pter-q13, �9p22-pter, �10p15,
�10q23-qter, �12q23-qter, �13q, �15q15-21, �16, �22q1341 Primary G 13 �1p13-q23, �3p/��p21-pter, �4q, �5p, �7q, ��8q, �9q13-33, �12qcent-q22, �14q11.2-13, �15q,
�18p, �21q21-qter, �22qL 8 �3q13-q21, �5q32-qter, �6q15-21, �8p, �10q22-qter, �11p15, �11q22-qter, �16p
42 Metasasis G 3 �8q23-qter, �12/��12pter-12q21, �20pL 5 �10q, �11p, �13q, �15qcent-q15, �18q
43 Metastasis G 14 ��1p12-31, �4pter-q13/��4qpter-q12, �4q28-q31.3,��6p12-22, �6q23-24, �7pter-q21/��7p, �8q22-23, �12q12-15, ��14q, �16q22-qter, �17q22, �18p, ��20q �22q
L 13 �1q41-qter, �2q34-qter, �3p21-q24, �8p, �9p21-pter, �10pter-p13, �10q, �11q14-qter, �12q23-q24.1,�13q, �16p12-pter, �18q21.1-q21.3, �19q13.2-q13.3
44 Metastasis G 5 �4q21-33/��4q28-31, �7q21-qter, �12/��12q12-q21, �17q22-qter, �22q13L 4 �5q12-23, �6q15-qter, �8p22-pter, �13q21-qter
45 Recurrence G 7 �4, �6, �7p, ��8q23-q24.1, �16, �20,�22q12-qterL 6 �2, �5q, �8 pter-q21.3, �9p2, �10p11.2-qter, �14q
46 Metastasis G 7 �1p21-p22, �3p25-pter, �3q28-qter, �4p12-p14, �6p21-q16, �14q21, �20qL 4 �6p23-pter, �9q, �12q23-qter, �19q13.1-q13.2
47 Recurrence G 11 �1p33-pter, �1q22-24, �4q31, �6p21-pter, �10p, �15q24-qter, �16p13, �17, �19q, ��20q12-q13.2,�21q22
L 4 �2q22-32, �3p13-q26.1, �5q14-21, �6qcent-q231G, gain, number of aberrations; L, loss, number of abberations; �, gain of DNA sequence copy number; ��, high-level gain; �, loss.
359CHROMOSOMAL INSTABILITIES IN OSTEOSARCOMAS
Number of aberrations in 41 primary and 6 relapsed tumorsIn 41 primary tumors, 370 gains and 290 losses were noted. The
median number of changes was 16.1 per tumor (range 0–33, 9.0gains and 7.1 losses). In 6 relapsed tumors, 47 gains and 36 losseswere detected. The median number of changes was 12.0 per tumor(range 8–27), which was not significantly different from that inprimary tumors. The frequent aberrations that developed in 41primary osteosarcomas are shown in Table III. The most frequentgains in primary tumors involved 8q (33 cases, 80%) including8qcen-q13 (30 cases, 73%), 8q21.3-q22 (29 cases, 71%) or 8q23-q24 (28 cases, 68%). The most frequent losses in primary tumorsinvolved 10q (26 cases, 63%) including 10q21-23 (26 cases, 63%).A total of 107 high-level gains were noted in 32 of 41 primarytumors. The most frequent high-level gain included 8q23-q24 (10cases, 24%).
Comparison of number of aberrations between primary andrelapsed tumors
Table IV shows aberrations with at least a 20% difference inoccurrence rates between 41 primary and 6 relapsed tumors. Gainsof either 8qcen-q13 (p � 0.002) or 8q21.3-q22 (p � 0.013) weresignificantly more frequent in primary tumors than in relapsedtumors. However, there was no aberration that occurred at asignificantly higher rate in relapsed tumors vs. primary tumors.
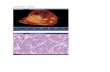
p53-protein accumulationTen cases, 7 of which displayed high-level gains of 17p13 and
3 exhibited gains of 17p, were analyzed for accumulation of p53protein by immunohistochemistry (Fig. 2). Homogenous nuclearstaining in almost 100% of the tumor-cell nuclei was observed in6 cases (5 with high-level gains of 17p13 and 1 with gain of 17p).In 4 cases, no p53 immunostaining was detectable. Both antibodies(Clone Bp53-11, Clone DO7) revealed similar staining results. In
contrast to the 6 positive cases, however, these 4 negative speci-mens required more extensive decalcification for making the his-tologic sections. This raises the possibility that decalcification mayhave led to false-negative results. Furthermore, paraffin-embeddedosteosarcoma cell line pellets of which the p53 status was known28
were included as positive and negative controls. Both antibodiesrevealed the same staining results (�, nuclear staining; �, nonuclear staining: MNNG �, HOS�, ZK58-, OST- SAOS-).
Univariate analysis of clinical data
There were no significant differences regarding the mediannumber of aberrations between men (19.0) and women (16.5),adolescents (�16 years of age; 7.0) and adults (�16 years of age;7.5), proximal (14.5) and distal (18.0) tumors, large (20.0) andsmall (16.0) tumors or tumors with (18.0) and without (17.0)primary metastases. The median numbers of aberrations among thesubtypes were 18.0 in 15 osteoblastic osteosarcomas (including 1postradiation osteoblastic osteosarcoma), 17.0 in 7 fibroblasticosteosarcomas, 21.0 in 3 chondroblastic osteosarcomas, 6.5 in 2parosteal osteosarcomas, 2.5 in 2 teleangiectatic osteosarcoma, 1.0in 1 periosteal osteosarcomas. There was a significant differencebetween the median numbers of aberrations comparing 27 osteo-blastic, fibroblastic, chondroblastic, teleangiectatic osteosarcomas(17.0) against 3 parosteal or periosteal osteosarcomas (3.0) (p �0.038). Although there were no significant differences regardingthe median numbers of aberrations between good (grades 1–3;17.5) and poor (grades 4–6; 17.5) responders to chemotherapy, again of 19p (8 of 14 poor responders, 0 of 20 good responders; p �0.001) or a loss of 9p (6 of 14 poor responders, 1 of 20 goodresponders; p � 0.012) was more frequent in poor responders thanin good responders.
FIGURE 1 – Chromosomal aberrations of osteosarcomas by comparative genomic hybridization. Areas on the left side of the chromosomeindicate losses and areas on the right show gains. Dark areas highlight high-level gains.
360 OZAKI ET AL.
Combinations of chromosomal aberrations in 41 primary tumorsAmong the 41 primary tumors, 21 aberrations that appeared in
�13 cases were selected for combination analysis (Fig. 3) includ-ing loss of 5q, loss of 13q14 and gain of 4q13-q21 (as opposed toloss of 5q32-q33, loss of 13q and gain of 4q, respectively) and 3aberrations within the 8q region. Of 210 combinations, 65 com-binations occurred �13 times. Excluding combinations composedexclusively of 8q aberrations, the 3 most frequent combinationswere a gain of 1p21-p31 along with a gain of either 1q21-q24 (21cases) or 8qcen-q13 (21 cases) and a gain of 8qcen-q13 with a lossof 10q21-22 (21 cases).
Event-free survival (EFS) in 41 patients with primary tumorOur analysis of clinical factors revealed no difference in the
cumulative EFS according to tumor site or tumor size (Table III).Nevertheless, a lower EFS was observed in 8 patients with primarymetastasis at diagnosis (p � 0.002) or in 15 patients with poorhistologic response (p � 0.005). The event-free survival was notassociated with number of aberrations.
Chromosomal aberrations (gains or losses) that occurred in 13or more cases and high-level gains in 6 or more patients wereselected for univariate analysis. Patients with a loss of 13q14 (p �0.042) or a high-level gain of 19p (p � 0.012) had a significantlylower EFS than those without these aberrations. Surprisingly, a
loss of 10q21-22 (p � 0.017) or a loss of 5q (p � 0.007) wasassociated with a better prognosis (Fig. 4).
Furthermore, we analyzed the relationship between combina-tions of chromosomal aberrations and EFS in 41 primary tumors(Table V). A significantly higher EFS was observed for patientswith a loss of 5q32-q33 and either gains of 1p21-p31 (n � 17, p �0.009) or loss of 10q21-q22 (n � 16, p � 0.018). Moreover, ahigher EFS was noted for patients with a loss of 10q21-q22 andeither a gain of 1p21-p31 (n � 16, p � 0.001), a gain of 1q21-q24(n � 15, p � 0.002) or a loss of 2q34-qter (n � 12, p � 0.043).A significantly lower EFS was associated with combinations of aloss of 13q14 and a gain of either 8qcen-q13 (n � 14, p � 0.005),8q21.3-q22 (n � 12, p � 0.001), 8q23-q24 (n � 13, p � 0.001),17p (n � 9, p � 0.001) or 4q13-q21 (n � 8, p � 0.001) (Fig. 5).
The most important prognostic factors were identified by astepwise regression test among several covariates (including pri-mary metastasis; histologic response; gains of 1p21-p31, 1q21-q24, 8qcen-q13, 8q21.3-q22, 8q23-q24, 17p and 21q; high-levelgain of 19p; and losses of 5q, 10q21-q22 and 13q14) by multivar-iate analysis. By the Cox proportional hazard model (includingprimary metastasis), primary metastasis (relative risk [RR] �3.774; p � 0.017) and no loss of 5q (RR � 3.639; p � 0.029) wereidentified as being independent poor prognostic factors (Table VI).When applying the Cox proportional hazard model (and excludingprimary metastasis), no loss of 5q (RR �7.840; p � 0.001) andloss of 13q14 (RR � 5.988; p � 0.002) were shown to beindependent prognostic factors.
DISCUSSION
CGH analysis revealed a high number of unbalanced geneticalterations in nearly all our osteosarcoma cases. Despite this fact,we identified clear tendencies, or clustering, of aberrations. Up tonow, there have been 5 other reports on CGH on osteosarco-mas.19–23 In comparison with these studies (Table VII), we ob-served the highest number of aberrations per tumor (16.1). Ahigher frequency of gains over losses was reported by 4 groupsincluding our study, and gains of chromosome 1 or 8q werefrequently observed (Table VII). Tarkkanen et al.21 reported that
TABLE III – FREQUENT ABBERATIONS IN 41 PRIMARY TUMORS ANDEVENT-FREE SURVIVAL BY UNIVARIATE ANALYSIS
Variable Positive(n � 41)
Univariate analysis ofevent-free survival (p)
Gains (n � 13 cases)8q 33 NS2
8qcent-q13 30 NS8q21.3-q22 29 NS8q23-q24 28 NS
1p21-p31 26 NS1q21-q24 24 NS4q 19 NS
4q13-q21 18 NS6p12-p21 19 NS17p 18 NS5p13-p14 16 NS21q 15 NS7p21 14 NS20q 14 NS12p 13 NS
Losses (n � 13 cases)10q 26 NS10q21-221 23 0.0175q1 23 0.007
5q32-q331 22 0.01213q 23 NS
13q14 17 0.0422q34-qter 19 NS6q16-q22 18 NS10p 15 NS16p 15 NS16q 13 NS
High-level gains (n � 6 cases)8q23-q24 10 NS17p13 10 NS1q21-q24 8 NS1p21-31 6 NS19p 6 0.012
Clinical findingsTumor site (proximal) 12 NSSize (large) 8 NSPrimary metastasis (yes) 8 0.002Histologic response (poor) 15 0.005
Number of aberrations (CGH)�15 26 NS�20 15 NS
1Associated with a better prognosis.–2NS, not significant.
TABLE IV – FREQUENT ABERRATIONS IN 41 PRIMARY AND 6 RELAPSEDOSTEOSARCOMAS: ABERRATIONS WITH A DIFFERENCE OF �20% IN
OCCURRENCE RATES BETWEEN PRIMARY AND RELAPSED CASES
Aberrations Positive in primarycases (n � 41)
Positive in relapsedcases (n � 6)
2 test1
(p)
Frequent aberrations inprimary cases
Gains8qcent-q13 30 (73%) 0 (0%) 0.0018q21.3-q22 29 (71%) 1 (17%) 0.0188q23-24 28 (68%) 2 (33%) NS2
1p21-p31 26 (63%) 2 (33%) NS1q21-q24 24 (59%) 1 (17%) NS17p 18 (44%) 1 (17%) NS5p13-p14 16 (39%) 0 (0%) NS21q 15 (37%) 1 (17%) NS19p 12 (29%) 0 (0%) NS9q21-22 11 (27%) 0 (0%) NS2p 10 (24%) 0 (0%) NS2q31-32 9 (22%) 0 (0%) NS
Frequent aberrations inrelapsed cases
Gains4q13-q21 18(46%) 4 (67%) NS6p12-p21.2 19 (46%) 4 (67%) NS20q13 14 (34%) 4 (67%) NS12q13-q15 8 (20%) 3 (50%) NS17q22-q23 11 (27%) 3 (50%) NS
Losses8p2 9 (22%) 3 (50%) NS
1With Fischer’s correction.–2NS, not significant.
361CHROMOSOMAL INSTABILITIES IN OSTEOSARCOMAS
patients with copy number increase at either 1q21, 8cen-q13 or8q21.3-q22 had a poorer prognosis than patients without theseaberrations. Although the reported chromosomal losses variedamong studies, losses of 10q, 10p and 2q were frequently observedby Tarkkanen et al.19,21 and Zielenska et al.23 Excluding 1 study byTarkkanen et al.,21 however, these previous reports do not includea statistical analysis correlating chromosomal aberrations withpatient outcome. In general, our data on the distribution pattern ofchromosomal aberrations (Fig. 1) was similar to that obtained byTarkkanen et al.;21 however, we observed differences in the fre-quencies of losses of 5q and 16 and gain of 11q. The number oflosses reported by Stock et al.22 is too small to be compared tothose from the other studies. Our observations of frequent high-level gains of chromosome 1p, 8q or 17p support those by Tark-kanen et al.21 and Zielenska et al.23 The most frequent combina-tions of chromosomal aberrations were a gain of 1p21-p31 alongwith a gain of either 1q21-q24 or 8qcen-q13 and a gain of 8qcen-q13 with a loss of 10q21-22. In our study, the relapsed tumors didnot exhibit more aberrations than the primary tumors. This may bedue to the small number of relapsed cases (n � 6) or the milderbiologic character associated with late relapse dates (average 58months). A similar observation was made by Forus et al.,20 whodescribe an average of 3.7 and 1.8 aberrations in 10 primary and 4relapsed tumors, respectively.
In relating the identified chromosomal aberrations to clinico-pathologic results, we found that 27 high grade osteosarcomas (15osteoblastic, including 1 postradiation osteoblastic, 7 fibroblastic,3 chondroblastic, 2 teleangiectatic) had a significantly higherfreqeuncy of abbreviations (17.0) than the 3 tumors of otherosteosarcoma subtypes (3.0 per tumor, including 2 parosteal os-teosarcomas and 1 periosteal osteosarcoma. Similarly, Tarkkanenet al.21 observed an average of 3.5 aberrations in 2 cases ofhigh-grade periosteal osteosarcoma, whereas Szymanska et al.29
described a mean of 6 aberrations per tumor (range 1–13) in theirprimary parosteal osteosarcomas. We also observed that a gain of19p or a loss of 9p was more frequent in poor responders than ingood responders. Moreover, a high-level gain of 19p was associ-ated with a lower EFS. There are several candidate targets on 19psuch as the RAS-oncogene family member RAB3A (19p13.2) andthe insulin-receptor gene (INSR; 19p13.3-p13.2). The cyclin-de-pendent kinase inhibitor 2A (CDKN2A) tumor-suppressor gene islocated at 9p21,18 and its deletion has been associated with a poorprognosis in osteosarcomas.30
Losses of 5q and losses of 13q14 were the most importantprognostic factors among chromosomal aberrations identified in
FIGURE 2 – p53 immunohistochemistry of a high-grade osteoblasticosteosarcoma with a gain of 17p on CGH analysis. Distinct nuclearimmunostaining indicates accumulation/overexpression of p53 nuclearphosphoprotein. Original magnification 200� (a), 630� (b). Anti-p53mouse monoclonal antibody, Clone Bp53-11 (Ventana Medical Sys-tems).
FIGURE 3 – Combinations of aberrations in 41 primary cases. Thenumbers indicate the incidence of each aberration combination, andnumbers in parentheses denote the incidence of each individual aber-ration. Among 210 combinations, 65 combinations occurred �13times. Five combinations (blue) were associated with a better progno-sis, while 5 other combinations (magenta) were associated with aworse prognosis.
FIGURE 4 – Event-free survival in 41 patients with primary osteo-sarcomas according to the presence or absence of a 5q loss (p �0.007).
362 OZAKI ET AL.
our study. We observed 17 cases (41%) with a loss of 13q14 andcould confirm this finding by FISH in selected cases. Stock et al.22
have reported an incidence of LOH at the RB1 locus from 60–70%in osteosarcomas, which has been associated with a poor progno-sis.10,11 13q14 contains LEU1 (leukemia associated gene 1) andLEU2 (leukemia associated gene 2), which are strong candidates astumor suppressor genes relevant to chronic lymphocytic leuke-mia.18,31 We determined that a loss of 13q14 was a strong poorprognostic factor both alone and in combinations with other aber-rations.
Unexpectedly, a loss of 5q was associated with a better prog-nosis. We hypothesize that osteosarcomas may be predisposed tocarrying extra copies of chromosome 5; therefore, losses of the 5qarm (and its associated oncogenes) may account for the observedbetter survival rate. Preliminary CGH and spectral karyotypingexperiments on osteosarcoma cell lines have, in fact, revealedchromosome 5 gains, although not in every case (data not shown).Whereas additional copies of chromosome 5 may result in gain ofgenetic material on 5q and thus amplification/overexpression ofoncogenes in this region, loss of 5q may reverse this effect.Recently, several malignant glioma cell lines (LN-18, T98G andU251MG) harboring a gain of proximal 5q, where cyclin B1(CCNB1; 5q12) and cyclin H (CCNH; 5q13.3-q14) reside, havebeen shown to exhibit increased growth rates.32 Other candidategenes on 5q include interleukin 3 (IL3), IL4, IL5 and IL9 (all at5q31.1), adhesion proteins such as alpha 1 catenin (CTNNA1;5q31), fibroblast growth factor 1 (FGF1; 5q31), FGF receptor 4(FGFR4; 5q35.1-qter), platelet-derived growth factor receptor beta(PDGFRB; 5q31-q32) and transforming growth factor, beta-in-
duced (TGFBI; 5q31). FGF1 has recently been demonstrated toenhance the expression of cytosolic phospholipase A2 (cPLA2)and (in the presence of IL1) both the expression of prostaglandin-endoperoxide synthase 2 (PTGS2) and the biosynthesis of prosta-glandin E2 in human osteosarcoma cell line, MG-63.33
Univariate analysis, but not multivariate analysis, indicated thata 10q21-23 loss was an important good prognostic factor. Wespeculate that the loss of parentally imprinted genes on the 10qarm may result in a better prognosis if the ubiquitously expressedallele is required for oncogenesis. In fact, imprinting is thought toplay a role in the etiology of Hirschsprung’s disease caused bymutations in the RET proto-oncogene (10q11.2).34 Up to now,genomic imprinting in sarcomas has been reported in insulin-likegrowth factor 2 (IGF2; 11p15.5) in rhabdomyosarcomas35 andEwing tumors.36 In the 10q region, genomic imprinting of FGFR2has been reported in Apert syndrome.37 Another candidate gene inthis region is FGF8 (10q25-26).
The most frequent aberration was a gain of 8q. Chromosome 8gains are also frequent in Ewing tumors.38,39 Although we ob-served a high-level gain of 8q23-q24 in 24% of cases, MYC(8q24.12-q24.13) amplification has been reported to occur in only7–12% of osteosarcomas.40,41 Tarkannen et al.21 reported that acopy-number increase at 8q represents a statistically significantpoor prognostic factor; we arrived at this same conclusion for 8qgains, but only when present in combination with losses of 13q14.
Other frequent gains included 1p21-p31 and 1q21-q22. The 1pgains may implicate the gene for v-jun avian sarcoma virus 17oncogene homolog (JUN; 1p31) or cysteine-rich, angiogenic in-ducer, 61 (CYR61, 1p22-p31), which mediates cell adhesion, mi-gration and angiogenesis.42,43 The 1q21-q22 region includes genesfor the S100 family of calcium-binding proteins (S100A1, A4 andA6 on 1q21), small proline-rich protein 3 (SPRR3; 1q21-q22)17 andcathepsins S and K (CTSS and CTSK, respectively; 1q21). Al-though patients with a 1q21 gain have been reported to exhibit apoor prognosis (p � 0.04),21 we associated good prognosis withthis aberration when it occurred simultaneously with a loss of 5q.Molecular analysis of the 1q amplification in human sarcomasrevealed that FLG, NTRK1 and SPRR3, located in 1q21, were themost frequently amplified genes.17,44 However, the frequent 1qgains appear to have little impact on prognosis.
Seventeen percent of osteosarcomas showed a loss of 17p, andthis may affect the TP53 tumor suppressor gene on 17p13.1.18 Onthe other hand, a 17p gain was seen in 44% of cases in our studyand was determined to be a poor prognostic factor in combinationwith a 13q14 loss. Previous studies have reported an amplificationrate of 20–30% at 17p11-12 in osteosarcomas.20,21 We hypothe-size that 17p gains and high-level gains of 17p13 may result in theoverexpression of mutant p53 protein. Experimental evidence forthis hypothesis comes from our analyses of p53 protein accumu-lation in the tumor-cell nuclei in such cases. As wild-type p53protein is usually not detectable because of its short half-life,alterations in the p53 tumor suppressor gene result in an overpro-duction and accumulation of this protein and permit its detectionby immunohistochemistry. Therefore, p53 accumulation indicatesunderlying mutations or other alterations of TP53.45,46 Overex-pression of mutant p53 protein with the loss of its function as atumor suppressor gene would explain our association of a signif-icantly lower EFS with a combination of a loss of 13q14 (RB1locus) and gain of 17p.
Twenty-two percent and 10% of cases showed gains and high-level gains, respectively, of 12q13-q15. This region is well knownfor the MDM2-SAS oncogene locus (12q12-q13),17 which is am-plified in 14–27% of osteosarcomas.12,47 Unfortunately, the rela-tively low aberration frequency we observed did not permit furtherstatistical analysis.
In summary, genomic imbalances were detected in the majorityof osteosarcomas. The most frequent gains involved 8q, chromo-some 1 or 4q, whereas the most frequent losses included 10q, 5qor 13q. A gain of 19p or loss of 9p was correlated with a poor
TABLE V – EVENT-FREE SURVIVAL BY UNIVARIATE ANALYSISACCORDING TO THE COMBINATIONS OF CHROMOSOMAL ABERRATIONS:
10 COMBINATIONS WERE ASSOCIATED WITH PROGNOSIS
Variable Positive(n � 41)
Univariate analysisof event-free survival
(p)
Good prognosis�5q32-q33 � �1p21-p31 17 0.009
� �10q21-q22 16 0.018�10q21-q22 � �1p21-p31 16 �0.001
� �1q21-q24 15 0.002� �2q34-qter 12 0.043
Poor prognosis�13q14 � �8qcent-q13 14 0.005
� �8q21.3-q22 12 �0.001� �8q23-q24 13 �0.001� �17p 9 0.001� �4q13-q21 8 �0.001
FIGURE 5 – Event-free survival in 41 patients with primary osteo-sarcomas according to the presence or absence of a combination of a13q14 loss and an 8q23-q24 gain (p � 0.001).
363CHROMOSOMAL INSTABILITIES IN OSTEOSARCOMAS
histologic response against chemotherapy. A lower EFS was ob-served for patients with primary metastasis, poor histologic re-sponse, high-level gain of 19p or loss of 13q14 than for patientswithout these aberrations. Losses of 10q21-22 or 5q were corre-lated with good prognosis. Multivariate analysis revealed thatprimary metastasis, loss of 5q or loss of 13q14 were independentprognostic factors. Although there were many genomic aberra-tions, a few chromosomal aberrations were closely related with theprognosis of osteosarcoma patients. These data were obtained froma large series of osteosarcomas, with 40 of 41 primary tumorstreated with the same protocol, and could help to clarify thebiologic role of certain genetic aberrations. We are aware thatthese findings need to be confirmed in prospective studies, but we
believe at the same time that they could turn out to be useful forevaluating prognosis, detecting additional target regions importantfor pathogenesis or biologic behavior and may finally lead totreatments based on the genetic background of osteosarcoma.
ACKNOWLEDGEMENTS
The authors thank Ms. P. Fischer, Ms. L. Grote, Ms. P. Meier,Ms. U. Neubert, Ms. F. Schmidt and Ms. A. Sommer for excellenttechnical support.
REFERENCES
1. Ozaki T, Inoue H, Taguchi K, Sugihara S. Gadolinium-DTPA en-hanced magnetic resonance imaging of bone and soft tissue sarcomasin comparison with pathological findings. Acta Med Okayama 1992;46:471–7.
2. Schima W, Amann G, Stiglbauer R, Windhager R, Kramer J, Nico-lakis M, Farres MT, Imhof H. Preoperative staging of osteosarcoma:efficacy of MR imaging in detecting joint involvement. AJR Am JRoentgenol 1994;163:1171–5.
3. Rosen G, Nirenberg A. Neoadjuvant chemotherapy for osteogenicsarcoma: a five year follow-up (T-10) and preliminary report of newstudies (T-12). Prog Clin Biol Res 1985;201:39–51.
4. Winkler K, Bielack S, Delling G, Salzer-Kuntschik M, Kotz R,Greenshaw C, Jurgens H, Ritter J, Kusnierz-Glaz C, Erttmann R,Gredicke G, Graf N, et al. Effect of intraarterial versus intravenouscisplatin in addition to systemic doxorubicin, high-dose methotrexate,and ifosfamide on histologic tumor response in osteosarcoma (studyCOSS-86). Cancer 1990;66):1703–10.
5. Bielack SS, Wulff B, Delling G, Gobel U, Kotz R, Ritter J, WinklerK. Osteosarcoma of the trunk treated by multimodal therapy: experi-ence of the Cooperative Osteosarcoma study group (COSS). MedPediatr Oncol 1995;24:6–12.
6. Bielack S, Kempf-Bielack B, Schwenzer D, Birkfellner T, Delling G,Ewerbeck V, Exner GU, Fuchs N, Gobel U, Graf N, Heise U, HelmkeK, et al. Neoadjuvant therapy for localized osteosarcoma of extrem-
ities. Results from the Cooperative osteosarcoma study group COSSof 925 patients. Klin Padiat 1999;211:260–70.
7. Enneking WF, Spanier SS, Goodman MA. A system for the surgicalstaging of musculoskeletal sarcoma. Clin Orthop 1980;153:106–20.
8. O’Connor MI, Sim FH. Salvage of the limb in the treatment ofmalignant pelvic tumors. J Bone Joint Surg Am 1989;71:481–94.
9. Yamaguchi T, Toguchida J, Yamamuro T, Kotoura Y, Takada N,Kawaguchi N, Kaneko Y, Nakamura Y, Sasaki MS, Ishizaki K.Allelotype analysis in osteosarcomas: frequent allele loss on 3q, 13q,17p, and 18q. Cancer Res 1992;52:2419–23.
10. Feugeas O, Guriec N, Babin-Boilletot A, Marcellin L, Simon P, Babin S,Thyss A, Hofman P, Terrier P, Kalifa C, Brunat-Mentigny M, PatricotLM, et al. Loss of heterozygosity of the RB gene is a poor prognosticfactor in patients with osteosarcoma. J Clin Oncol 1996;14:467–72.
11. Wadayama B, Toguchida J, Shimizu T, Ishizaki K, Sasaki MS,Kotoura Y, Yamamuro T. Mutation spectrum of the retinoblastomagene in osteosarcomas. Cancer Res 1994;54:3042–48.
12. Ladanyi M, Cha C, Lewis R, Jhanwar SC, Huvos AG, Healey JH.MDM2 gene amplification in metastatic osteosarcoma. Cancer Res1993;53:16–18.
13. Forus A, Florenes VA, Maelandsmo GM, Fodstad O, Myklebost O.The protooncogene CHOP/GADD153, involved in growth arrest andDNA damage response, is amplified in a subset of human sarcomas.Cancer Genet Cytogenet 1994;78:165–71.
14. Noble-Topham SE, Burrow SR, Eppert K, Kandel RA, Meltzer PS,
TABLE VI – MULTIVARIATE ANALYSIS WITH THE COX MODEL, USING 2 SELECTED FACTORS, BY STEPWISE REGRESSION TESTING
Prognostic factor n2 factors including primary metastasis 2 factors excluding primary metastasis
Multivariate p Relative risk 95% Confidenceinterval Multivariate p Relative risk 95% Confidence
interval
Primary metastasis 8 0.017 3.774 1.267–11.236No loss of 5q 23 0.029 3.639 1.141–11.607 0.001 7.840 2.294–26.794Loss of 13q14 18 0.002 5.988 1.912–18.86
TABLE VII – SUMMARY OF THE REPORTS OF CGH IN OSTEOSARCOMA
Diagnosis n No. of caseswith aberration
Aberrationsper tumor Amplification Gain : loss
ratio Frequent gains Frequent losses Prognostic factors
Osteosarcoma1
(primary 9,metastasis, 1)
11 10 (91%) 11.0 8 (73%) 1 : 1.3 8q, Xp, 1q, 11q, Xq 2q, 6q, 8p, 10p –
Osteosarcoma2
(primary 10,metastasis 4)7
14 9 (64%) 3.18 7 (18%) – 1q, 6p, 8q, 17p – –
Osteosarcoma3
(high grade)31 30 (97%) 9.6 21 (68%) 1.9 : 1 1q, 8q, 14q, Xp 6q, 10p, 10q gain of 8q or 1q
Osteosarcoma4
(primary 15,relapse 1)
16 12 (75%) 6.1 – 9.9 : 1 8q, 4q, 7q, 5p, 1p 19q, 7p –
Osteosarcoma5
(primary 17,metastasis 1)
18 18 (100%) 4.7 4 (22%) 2.7 : 1 1p, 19, 17p, 5p, 16p, 8q 2q, 10, 13 –
Osteosarcoma6
(primary,current study)
41 40 (98%) 16.1 32 (78%) 1.3 : 1 8q, 1p, 1q, 4q, 6p 10q, 5q, 13q, 2q loss of 5q or 13q
1(19).–2(20).–3(21).–4(22).–5(23).–6Current study.–7Ten tumors were xenografts in nude mice and 4 were obtained from patients. Seven of 10xenografts were established from primary tumors and 3 of 4 tumors were obtained from primary tumors of patients. The other tumors weremetastases.–8Gains only.
364 OZAKI ET AL.
Bell RS, Andrulus IL. SAS is amplified predominantly in surfaceosteosarcoma. J Orthop Res 1996;14:700–5.
15. Onda M, Matsuda S, Higaki S, Iijima T, Fukushima J, Yokokura A,Kojima T, Horiuchi H, Kurokawa T, Yamamoto T. ErbB-2 expressionis correlated with poor prognosis for patients with osteosarcoma.Cancer 1996;77:71–8.
16. Gorlick R, Huvos AG, Heller G, Aledo A, Beardsley GP, Healey JH,Meyers PA. Expression of HER2/erbB-2 correlates with survival inosteosarcoma. J Clin Oncol 1999;17:2781–8.
17. Knuutila S, Bjorkqvist AM, Autio K, Tarkkanen M, Wolf M, MonniO, Szymanska J, Larramendy ML, Tapper J, Pere H, El-Rifai W,Hemmer S, et al. DNA copy number amplifications in human neo-plasms: review of comparative genomic hybridization studies. Am JPathol 1998;152:1107–23.
18. Knuutila S, Aalto Y, Autio K, Bjorkqvist AM, El-Rifai W, HemmerS, Huhta T, Kettunen E, Kiuru-Kuhlefelt S, Larramendy ML, Lush-nikova T, Monni O, et al. DNA copy number losses in humanneoplasms. Am J Pathol 1999;155:683–94.
19. Tarkkanen M, Karhu R, Kallioniemi A, Elomaa IK, Kivioja AH,Nevalainen J, Bohling T, Karaharju E, Hyytinen E, Knuutila S,Kallioniemi OP. Gains and losses of DNA sequences in osteosarco-mas by comparative genomic hybridization. Cancer Res 1995;55:1334–8.
20. Forus A, Weghuis DO, Smeets D, Fodstad O, Myklebost O, Geurtsvan Kessel A. Comparative genomic hybridization analysis of humansarcomas: II. Identification of novel amplicons at 6p and 17p inosteosarcomas. Genes Chromosomes Cancer 1995;14:15–21.
21. Tarkkanen M, Elomaa I, Blomqvist C, Kivioja AH, Kellokumpu-Lehtinen P, Bohling T, Valle J, Knuutila S. DNA sequence copynumber increase at 8q: a potential new prognostic marker in high-grade osteosarcoma. Int J Cancer 1999;84:114.
22. Stock C, Kager L, Fink FM, Gadner H, Ambros PF. Chromosomalregions involved in the pathogenesis of osteosarcomas. Genes Chro-mosomes Cancer 2000;28:329–36.
23. Zielenska M, Bayani J, Pandita A, Toledo S, Marrano P, Andrade J,Petrilli A, Thorner P, Sorensen P, Squire JA. Comparative genomichybridization analysis identifies gains of 1p35 approximately p36 andchromosome 19 in osteosarcoma. Cancer Genet Cytogenet 2001;130:14–21.
24. Gobel V, Jurgens H, Etspuler G, Kemperdick H, Jungblut RM,Stienen U, Gobel U. Prognostic significance of tumor volume inlocalized Ewing’s sarcoma of bone in children and adolescents. JCancer Res Clin Oncol 1987;113:187–91.
25. Salzer-Kuntschik M, Brand G, Delling G. Determination of the degreeof morphological regression following chemotherapy in malignantbone tumors. Pathologe 1983;4:135–41.
26. Kallioniemi OP, Kallioniemi A, Piper J, Isola J, Waldman FM, GrayJW, Pinkel D. Optimizing comparative genomic hybridization foranalysis of DNA sequence copy number changes in solid tumors.Genes Chromosomes Cancer 1994;10:231–43.
27. Brinkschmidt C, Poremba C, Christiansen H, Simon R, Schafer KL,Terpe HJ, Lampert F, Boecker W, Dockhorn-Dworniczak B. Com-parative genomic hybridization and telomerase activity analysis iden-tify two biologically different groups of 4s neuroblastomas. Br JCancer 1998;77:2223–29.
28. Scheel C, Schaefer KL, Jauch A, Keller M, Wai D, Brinkschmidt C,van Valen F, Boecker W, Dockhorn-Dworniczak B, Poremba C.Alternative lengthening of telomeres is associated with chromosomalinstability in osteosarcomas. Oncogene 2001;20:3835–44.
29. Szymanska J, Mandahl N, Mertens F, Tarkkanen M, Karaharju E,Knuutila S. Ring chromosomes in parosteal osteosarcoma containsequences from 12q13-15: a combined cytogenetic and comparativegenomic hybridization study. Genes Chromosomes Cancer 1996;16:31–34.
30. Patino-Garcia A, Sierrasesumaga L. Analysis of the p16INK4 and
TP53 tumor suppressor genes in bone sarcoma pediatric patients.Cancer Genet Cytogenet 1997;98:50–5.
31. Liu Y, Corcoran M, Rasool O, Ivanova G, Ibbotson R, Grander D,Iyengar A, Baranova A, Kashuba V, Merup M, Wu X, Gardiner A, etal. Cloning of two candidate tumor suppressor genes within a 10 kbregion on chromosome 13q14, frequently deleted in chronic lympho-cytic leukemia. Oncogene 1997;15:2463–73.
32. Weber RG, Rieger J, Naumann U, Lichter P, Weller M. Chromosomalimbalances associated with response to chemotherapy and cytotoxiccytokines in human malignant glioma cell lines. Int J Cancer 2001;91:213–18.
33. Laulederkind SJ, Kirtikara K, Raghow R, Ballou LR. The regulationof PGE(2) biosynthesis in MG-63 osteosarcoma cells by IL-1 andFGF is cell density-dependent. Exp Cell Res 2000;258:409–16.
34. Peretz H, Luboshitsky R, Baron E, Biton A, Gershoni R, Usher S,Grynberg E, Yakobson E, Graff E, Lapidot M. Cys 618 Arg mutationin the RET proto-oncogene associated with familial medullary thyroidcarcinoma and maternally transmitted Hirschsprung’s disease sug-gesting a role for imprinting. Hum Mutat 1997;10:155–59.
35. Anderson J, Gordon A, McManus A, Shipley J, Pritchard-Jones K.Disruption of imprinted genes at chromosome region 11p15.5 inpaediatric rhabdomyosarcoma. Neoplasia 1999;1:340–48.
36. Zhan S, Shapiro DN, Helman LJ. Loss of imprinting of IGF2 inEwing’s sarcoma. Oncogene 1995;11:2503–7.
37. Moloney DM, Slaney SF, Oldridge M, Wall SA, Sahlin P, Stenman G,Wilkie AO. Exclusive paternal origin of new mutations in Apertsyndrome. Nat Genet 1996;13:48–53.
38. Tarkkanen M, Kiuru-Kuhlefelt S, Blomqvist C, Armengol G, BohlingT, Ekfors T, Virolainen M, Lindholm P, Monge O, Picci P, KnuutilaS, Elomaa I. Clinical correlations of genetic changes by comparativegenomic hybridization in Ewing sarcoma and related tumors. CancerGenet Cytogenet 1999;114:35–41.
39. Ozaki T, Paulussen M, Poremba C, Brinkschmidt C, Rerin J, AhrensS, Hoffmann C, Hillmann A, Wai D, Schaefer KL, Boecker W,Juergens H, Winkelmann W, et al. Genetic imbalances revealed bycomparative genomic hybridization in Ewing tumors. Genes Chromo-somes Cancer 2001;32:164–71.
40. Ladanyi M, Park CK, Lewis R, Jhanwar SC, Healey JH, Huvos AG.Sporadic amplification of the MYC gene in human osteosarcomas.Diagn Mol Pathol 1993;2:163–7.
41. Pompetti F, Rizzo P, Simon RM, Freidlin B, Mew DJ, Pass HI, PicciP, Levine AS, Carbone M. Oncogene alterations in primary, recurrent,and metastatic human bone tumors. J Cell Biochem 1996;63:37–50.
42. Babic AM, Kireeva ML, Kolesnikova TV, Lau LF. CYR61, a productof a growth factor-inducible immediate early gene, promotes angio-genesis and tumor growth. Proc Natl Acad Sci USA 1998;95:6355–60.
43. Kireeva ML, Mo FE, Yang GP, Lau LF. Cyr61, a product of a growthfactor-inducible immediate-early gene, promotes cell proliferation,migration, and adhesion. Mol Cell Biol 1996;16:1326–34.
44. Forus A, Weterman M, Van Kessel AG, Berner J-M, Fodstand O,Myklebost O. Characterisation of 1q21-22 amplifications in humansarcomas by CGH and molecular analysis. Cytogenet Cell Genet1996;72:8–14.
45. Poremba C, Yandell DW, Metze D, Kamanabrou D, Bocker W,Dockhorn-Dworniczak B. Immunohistochemical detection of p53 inmelanomas with rare p53 gene mutations is associated with mdm-2overexpression. Oncol Res 1995;7:331–9.
46. Poremba C, Yandell DW, Huang Q, Little JB, Mellin W, Schmid KW,Bocker W, Dockhorn-Dworniczak B. Frequency and spectrum of p53mutations in gastric cancer—a molecular genetic and immunohisto-chemical study. Virchows Arch 1995;426:447–55.
47. Oliner JD, Kinzler KW, Meltzer PS, George DL, Vogelstein B.Amplification of a gene encoding a p53-associated protein in humansarcomas. Nature 1992;358:80–3.
365CHROMOSOMAL INSTABILITIES IN OSTEOSARCOMAS
Related Documents