-1- Evaluation of four single-locus markers for Leishmania species discrimination by 1 sequencing 2 3 Gert Van der Auwera a# , Christophe Ravel b , Jaco J. Verweij c , Aldert Bart d , Gabriele Schönian e , 4 Ingrid Felger f 5 6 a Institute of Tropical Medicine, Antwerp, Belgium. 7 b University of Montpellier, UMR5290 MiVEGEC, and French Reference Centre on 8 Leishmaniasis, Montpellier, France. 9 c Department of Parasitology and Department of Medical Microbiology, Leiden University 10 Medical Center, Leiden, The Netherlands. Current address: Laboratory for Medical 11 Microbiology and Immunology, St. Elisabeth Hospital, Tilburg, The Netherlands. 12 d Department of Medical Microbiology, Center for Infection and Immunity Amsterdam, 13 Academic Medical Center, Amsterdam, The Netherlands. 14 e Institute of Microbiology and Hygiene, Charité University Medicine, Berlin, Germany. 15 f Swiss Tropical and Public Health Institute and University of Basel, Basel, Switzerland. 16 17 Running title: Leishmania species typing by 4 markers 18 19 Word count abstract: 249 20 Word count text: 3336 21 JCM Accepts, published online ahead of print on 22 January 2014 J. Clin. Microbiol. doi:10.1128/JCM.02936-13 Copyright © 2014, American Society for Microbiology. All Rights Reserved. on March 18, 2018 by guest http://jcm.asm.org/ Downloaded from

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

-1-
Evaluation of four single-locus markers for Leishmania species discrimination by 1
sequencing 2
3
Gert Van der Auwera a#, Christophe Ravel b, Jaco J. Verweij c, Aldert Bart d, Gabriele Schönian e, 4
Ingrid Felger f 5
6
a Institute of Tropical Medicine, Antwerp, Belgium. 7
b University of Montpellier, UMR5290 MiVEGEC, and French Reference Centre on 8
Leishmaniasis, Montpellier, France. 9
c Department of Parasitology and Department of Medical Microbiology, Leiden University 10
Medical Center, Leiden, The Netherlands. Current address: Laboratory for Medical 11
Microbiology and Immunology, St. Elisabeth Hospital, Tilburg, The Netherlands. 12
d Department of Medical Microbiology, Center for Infection and Immunity Amsterdam, 13
Academic Medical Center, Amsterdam, The Netherlands. 14
e Institute of Microbiology and Hygiene, Charité University Medicine, Berlin, Germany. 15
f Swiss Tropical and Public Health Institute and University of Basel, Basel, Switzerland. 16
17
Running title: Leishmania species typing by 4 markers 18
19
Word count abstract: 249 20
Word count text: 3336 21
JCM Accepts, published online ahead of print on 22 January 2014J. Clin. Microbiol. doi:10.1128/JCM.02936-13Copyright © 2014, American Society for Microbiology. All Rights Reserved.
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-2-
# Corresponding author: 22
Gert Van der Auwera 23
Department of Biomedical Sciences, Institute of Tropical Medicine 24
Nationalestraat 155, 2000 Antwerp, Belgium 25
Tel. +32 32476586; fax +32 32476359; e-mail [email protected] 26
27
Key Words: Leishmania; Multilocus sequence typing; ribosomal DNA; mini-exon; heat-shock 28
protein 70; 7SL-RNA. 29
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-3-
Abstract 30
Several genetic markers have been described for discriminating Leishmania species. In 31
most reported cases, one or a few polymorphisms are the basis of species identification, and 32
the methods were validated on a limited number of strains from a particular geographical 33
region. Thereby most techniques may underestimate the global intra-species variability, and are 34
applicable only in a certain area. In addition, inter-laboratory standardization is mostly absent, 35
complicating comparison among different studies. In this paper, we compared species typing 36
results from all sequence polymorphisms found in four popular markers applicable directly on 37
clinical samples: the mini-exon or spliced leader, the internal transcribed spacer of the 38
ribosomal DNA array; the 7SL-RNA gene; and the heat-shock protein 70 gene. Clustering was 39
evaluated among 74 Leishmania strains, selected to represent a wide geographic distribution 40
and genetic variability of the medically relevant species of the genus. Results were compared 41
with a multilocus sequence typing (MLST) approach using 7 single-copy household genes, and 42
with multilocus enzyme electrophoresis (MLEE), by some still considered the gold standard. We 43
show that strain groupings are highly congruent across the four different single-locus markers, 44
MLST, and MLEE. Overall, the heat-shock protein 70 gene and the mini-exon present the best 45
resolution for separating medically relevant species. As gene sequence analysis is validated here 46
on a global scale, it is advocated as the method of choice for use in genetic, clinical, and 47
epidemiological studies, and for managing patients with unknown origin of infection, especially 48
in Western infectious disease clinics dealing with imported leishmaniasis. 49
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-4-
Introduction 50
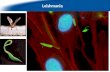
The parasitic protozoa of the genus Leishmania cause a spectrum of diseases in humans, 51
collectively called the leishmaniases. In its most benign form, referred to as cutaneous 52
leishmaniasis, the disease manifests itself as a localized skin ulcer at the site of infection by the 53
bite of a female infectious sandfly. Sometimes the parasites spread to other parts of the body, 54
causing secondary lesions. More severely, when the mucosa is infected the disease leads to 55
disfiguring lesions of nose and mouth, a condition known as mucosal leishmaniasis. Finally, 56
when the parasite colonizes internal organs such as the spleen, liver, and bone marrow, a 57
condition referred to as visceral leishmaniasis, the disease becomes lethal. As the manifestation 58
of disease to a large extent depends on the infecting species, so do the treatment options (1,2). 59
60
According to a recent estimate (3), leishmaniasis is endemic in 98 countries and 3 61
territories. Besides the endogenous population being at risk of infection, many active 62
transmission areas are frequently visited by tourists, military personnel, expats, and people 63
visiting friends and relatives. They can potentially import leishmaniasis into their home country, 64
and managing such cases calls for a globally applicable reliable species typing approach, as often 65
the time and place of infection is difficult to assess. Also in clinical and epidemiological studies 66
at the species level, accurate typing tools are required that have been validated on a global 67
scale, in order to deal with changing epidemiology and uncharted genetic mutations. 68
69
Several molecular assays have been described for discriminating Leishmania species, 70
based on various genomic loci. Four of these targets are quite widespread in literature: the 71
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-5-
mini-exon or spliced leader (ME)(4,5), the internal transcribed spacer of the ribosomal DNA 72
array (ITS1)(6-10); the 7SL-RNA (11,12); and the heat-shock protein 70 gene (hsp70)(13-16). In 73
this paper we set out to compare typing results on the basis of all sequence information present 74
from these targets, rather than using size and single-nucleotide polymorphism techniques such 75
as restriction-fragment length polymorphism analysis, specific PCRs, and oligonucleotide 76
hybridization. Even though sequencing is not available in primary health centers, it can 77
nowadays routinely be applied in Western clinics, clinical trials, and epidemiological surveys, 78
settings where a globally applicable typing strategy is most relevant. 79
80
Apart from a comparison of these four marker genes, we also compared the results to 81
those from multi-locus enzyme electrophoresis (MLEE)(17) and multi-locus sequence typing 82
(MLST)(18-20). MLEE has for many years been the gold standard in species typing of Leishmania, 83
but has now been surpassed by molecular techniques such as the high resolution MLST, which is 84
based upon sequence analysis of several household genes. We believe that our study will aid in 85
interpreting and comparing reported species differentiation by different genes and methods; 86
and contributes to a more reliable distinction of species, both in endemic and non-endemic 87
settings, and both in laboratory and clinical applications. 88
89
Materials and Methods 90
Strains: A total of 74 clinical Leishmania isolates was selected to represent the genetic 91
and geographical variability of the medically relevant species (Table 1 and Supplemental 92
Table S1). The species of most of these strains was determined by MLEE, while the others were 93
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-6-
initially typed using AFLP (amplified fragment length polymorphisms) (21) or other molecular 94
markers (22). DNA was obtained from promastigote cultures, either from the French Reference 95
Centre on Leishmaniasis (Montpellier, France), or from the culture collections of the Institute of 96
Tropical Medicine Alexander von Humboldt (Lima, Peru) and the Institute of Tropical Medicine 97
(Antwerp, Belgium). Various DNA extraction kits were used, such as the High Pure PCR Template 98
Preparation Kit (Roche, Basel, Switzerland) and the QIAamp DNA mini kit (Qiagen, Hilden, 99
Germany). 100
101
Multi-locus sequence typing (MLST): Seven genes were amplified from each strain, as 102
described in El Baidouri et al. (20): the putative elongation initiation factor 2 alpha subunit; the 103
putative spermidine synthase 1; a zinc binding dehydrogenase-like protein; a putative 104
translation initiation factor alpha subunit; a putative nucleoside hydrolase-like protein; a 105
conserved hypothetical protein; and the largest subunit of RNA polymerase II. After removing 106
low-quality sequence reads and both PCR primers, the sequences from these genes were 107
concatenated in a global alignment of 4677 nucleotides, from which a neighbor-joining 108
dendrogram was built on the basis of p-distances, with pairwise gap exclusion. Bootstrap 109
support was calculated from 2000 resamplings. All analyses were performed using the software 110
package MEGA5 (23, www.megasoftware.net). 111
112
Heat-shock protein 70 gene (hsp70): The 1280 bp fragment PCR-F was amplified as 113
described (14,16), and subsequently sequenced. A global sequence alignment was produced, 114
which was facilitated as no size variation was observed. After elimination of the PCR primer 115
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-7-
sequences, a dendrogram was produced on the basis of the remaining 1245 nucleotides, as 116
described for MLST above. 117
118
7SL-RNA gene (7SL-RNA): A PCR fragment was amplified as in Zelazny et al. (12), and 119
sequenced. As size variation was minimal (184-187 nucleotides were amplified), a global 120
sequence alignment was produced. After stripping the primer sequences, a dendrogram was 121
constructed as described for MLST. 122
123
rDNA-ITS1 (ITS1): A PCR fragment was amplified as in Schönian et al. (8). After 124
sequencing of the PCR amplicon, the primers were stripped from the sequences, resulting in 125
sequences between 257 and 302 nucleotides. Because of this size variation, no reliable global 126
sequence alignment could be constructed, and a neighbor-joining dendrogram was built with 127
the MEGA5 package, based on a guide tree produced from ClustalW version 1.7 (24). 128
129
Mini-exon (ME): A PCR fragment was amplified as in Marfurt et al. (4). After sequencing 130
of the PCR amplicon, the primers were stripped from the sequences, resulting in sequences 131
between 176 and 397 nucleotides in size. Because of this excessive size variation, no reliable 132
global sequence alignment could be constructed, and a dendrogram was built as described 133
above for ITS1. As some nucleotides were missing from the 5’ start of the sequence in 6 isolates 134
(indicated with ° in Fig. 1), the stretch corresponding to the first 11 nucleotides following the 135
forward primer was deleted from all sequences. Some isolates could only be partially sequenced 136
because of irresolvable sequence reads caused by an extensive homopolymer stretch that varies 137
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-8-
in size between the tandem repeated copies in the same genome, and these were omitted from 138
the analysis (indicated with * in Fig. 1). 139
140
Results 141
All GenBank accession numbers are listed in Supplemental Table S1. Fig. 1 shows the 142
dendrogram obtained on the basis of the 7 genes used in MLST analysis. Clusters corresponding 143
to the recognized species are indicated on the nodes, with their respective bootstrap support. 144
The MLEE-defined species L. major, L. aethiopica, L. tropica, L. lainsoni, and L. naiffi are all 145
identified as separate clusters supported by a 99 or 100% bootstrap value, while L. mexicana is 146
backed up by 71%. Also L. amazonensis and L. guyanensis both form 100% supported groups, 147
but with inclusion of L. garnhami and L. panamensis respectively. As for L. donovani, this 148
together with L. infantum and L. archibaldi constitutes a 100% supported cluster. Within this 149
cluster, L. infantum isolates derive from a common point, and the species is supported with 75% 150
provided strain MHOM/SU/84/MARZ-KRIM is considered L. infantum, as opposed to L. donovani 151
as determined by MLEE (25). Finally, L. braziliensis deserves some special attention. 152
L. braziliensis and L. peruviana jointly form a cluster with a 100% bootstrap value. In this cluster, 153
isolates referred to as L. braziliensis type 2 separate as a 100% supported subgroup that was 154
previously identified in AFLP analysis (group 3 in Odiwuor et al., 21). Its 97% supported sister 155
clade named L. braziliensis type 1 comprises also the L. peruviana isolates, which group with a 156
91% bootstrap value. 157
158
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-9-
The view from hsp70 is largely congruent with MLST analysis, even though bootstrap 159
values are somewhat lower (Fig. 1, Supplemental Fig. S1). On three fronts the hsp70 160
dendrogram deviates. First, L. mexicana isolates do not derive from a common point, but rather 161
group with L. amazonensis (including L. garnhami). Nevertheless, within this joint cluster 162
L. amazonensis is clearly recognizable (95% bootstrap value). Second, one L. naiffi strain 163
(MHOM/--/94/CRE58) does not group with the 2 others, but rather diverges between L. lainsoni 164
and the rest of the L. (Viannia) species (not shown). Third, L. braziliensis types 1 and 2 do not 165
form sister clades. 166
167
The 7SL-RNA dendrogram on the other hand shows a lower resolution, as it identifies far 168
less groups (Fig. 1, Supplemental Fig. S2), and generally with lower bootstrap support as do 169
MLST and hsp70. L. tropica and L. aethiopica constitute a joint cluster, but the species cannot be 170
identified separately. L. infantum cannot be distinguished from L. donovani. L. mexicana, 171
L. amazonensis, and L. garnhami too are not recognizable as separate taxa. In the L. (Viannia) 172
subgenus, L. braziliensis type 2 and L. naiffi form one group (92% bootstrap support). Finally, 173
L. braziliensis type 1, L. peruviana, L. guyanensis, and L. panamensis form a clade in which these 174
4 species cannot be distinguished. 175
176
The ITS1 dendrogram shows the same groups as does MLST for the L. (Leishmania) 177
subgenus (Fig. 1, Supplemental Fig. S3). In the L. (Viannia) subgenus, L. lainsoni forms a sister 178
clade of the remaining species, from which only L. naiffi and L. braziliensis type 2 separate as 179
defined entities. L. braziliensis type 1, L. peruviana, and L. guyanensis (including L. panamensis) 180
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-10-
cannot be discriminated based on ITS1. Bootstrap analysis could not be performed as no global 181
alignment was constructed. Sequence reads are often problematic in this region because of 182
homopolymer tracts, requiring a manual time-consuming read-out for the subsequent 183
stretches. Using this technique most sequences could be resolved, but still some are lacking 184
from the set (indicated with ~ in Fig. 1). PCR amplicon size differences between the species are 185
generally of no use for typing, as the ranges are largely overlapping (Fig. 1, Fig. S3). 186
187
The mini-exon separates the same species as does MLST in the L. (Leishmania) subgenus 188
(Fig. 1, Supplemental Fig. S4). Several L. infantum sequences are however lacking because of 189
sequencing difficulties, not allowing to fully assess the clustering of all strains of this species. In 190
the L. (Viannia) subgenus, no full-length sequences could be obtained from L. lainsoni, and 191
L. peruviana could not be separated from type 1 L. braziliensis strains. Sequencing of the mini-192
exon proved challenging in many instances because of intra-genome variation of different 193
tandem repeated copies, which is why several sequences are lacking from the analysis 194
(indicated with * Fig. 1). Also this marker does not allow bootstrap analysis as no global 195
sequence alignment is possible. In the mini-exon, size differences can be used to separate large 196
species groups: the Old World L. (Leishmania) samples have PCR amplicon sizes of 350 bp and 197
up, the New World L. (Leishmania) of about 300-330 bp, L. (Viannia) lainsoni approximately 198
between 300 and 350 bp, and the remaining L. (Viannia) around 225 bp. Within these groups, 199
size ranges for the different species are for the most part overlapping, not allowing species 200
discrimination by length alone. As for L. lainsoni, this species can be identified on the basis of 201
partial sequences (data not shown). 202
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-11-
203
Discussion 204
When comparing results from MLST, the four single-gene markers, and MLEE, the 205
following observations can be made for the species depicted in Fig. 1: 206
- L. major can clearly be separated in every analysis, hence its identification poses no 207
problem. 208
- L. turanica and L. gerbilli do not group with any species of human medical importance, and 209
generally they form sister taxa. One exception is the 7SL-RNA analysis, where L. turanica 210
clusters with the L. tropica-L.aethiopica group (result not shown). 211
- L. aethiopica and L. tropica: both species stand out as separate entities in all analyses 212
except 7SL-RNA, where they are identified as one group. 213
- L. donovani poses a more complicated problem. All markers clearly distinguish the 214
L. donovani species complex, which includes strains identified by MLEE as L. infantum and 215
L. archibaldi, from the other complexes. Within the complex, L. infantum can essentially be 216
recognized as a separate subgroup, at least from MLST, hsp70, ITS1, and the ME analysis, be 217
it with moderate bootstrap support. Strain MHOM/SU/84/MARZ-KRIM also belongs to this 218
group, even though it was classified as L. donovani by MLEE. The single L. chagasi isolate 219
MHOM/PA/78/WR285 is found in this cluster as well, in line with the current consensus 220
that this species is in fact New World L. infantum (26-28). The remaining marker 7SL-RNA 221
does not allow identification of L. infantum, and as several ME sequences are lacking some 222
reservations can be made for the mini-exon as well. For clinical case management, 223
distinguishing between both species is not always needed, as treatment is often identical 224
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-12-
(29,30). L. archibaldi and L. donovani form a mixed group, and neither one can be identified 225
as a distinguished species, which agrees perfectly with previous analyses (18,19,31,32,32-226
34). 227
- L. amazonensis can easily be identified in all analyses except 7SL-RNA, but it cannot be 228
distinguished from L. garnhami, which in our dataset is represented by one isolate only. 229
- L. mexicana is generally a sister clade of L. amazonensis, except in hsp70 where the latter 230
species diverges within the L. mexicana group, still allowing a clear identification. Using 231
7SL-RNA no distinction of both species is possible. 232
- L. lainsoni is easily recognized in all assays, except from the mini-exon of which no full-233
length sequences could be obtained, even though partial sequences allow identification of 234
the species. 235
- L. naiffi separates as a group in MLST, and can be identified using ITS1 and the mini-exon. In 236
hsp70, one L. naiffi strain (MHOM/--/94/CRE58) is separate from the two remaining ones, 237
and does not cluster with any species. 238
- L. braziliensis type 2 is clearly a recognizable group separate from L. braziliensis type 1, and 239
it can be identified by MLST, hsp70, ITS1, and the mini-exon. MLEE does not recognize this 240
is a separate taxonomic unit and classifies it with other L. braziliensis, even though the 241
MON141 zymodeme of MCAN/PE/91/LEM2222 – the only isolate of the group of which 242
MLEE data are available – is quite distinct. Also, a previous genome-wide AFLP analysis 243
clearly demonstrated the group to be a separate entity (21). Even though parasites 244
belonging to this group have been isolated from mucosal lesions (results not shown), the 245
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-13-
clinical relevance is not clear because L. braziliensis type 2 is not recognized by currently 246
deployed diagnostic assays, and hence clinical records are not properly documented. 247
- L. peruviana separates as a unit from within the L. braziliensis type 1 cluster. It can be 248
identified only by MLST and hsp70. Nevertheless, the boundary between L. peruviana and 249
L. braziliensis type 1 is a matter of debate, as intermediate forms carrying signatures of 250
both species do exist (21). Also, in MLEE only one protein separates both species (mannose 251
phosphate isomerase), making its species status at least dubious (35). In this analysis, we 252
have considered L. peruviana as in Odiwuor et al. (21). 253
- L. braziliensis type 1 forms a clearly defined group together with L. peruviana in MLST, 254
hsp70, and the mini-exon, but only in the former two L. peruviana can be separated from 255
the remaining isolates of the cluster. Contrary to L. peruviana, L. braziliensis is known to 256
frequently cause mucocutaneous leishmaniasis, which makes the distinction between both 257
species highly relevant (36). 258
- L. guyanensis is clearly recognized on the basis of MLST, hsp70, and the mini-exon. As only 1 259
L. panamensis strain was included in our dendrograms, we cannot confirm the correct 260
identification of this species on the basis of the markers studied here. 261
262
All four genetic markers give a highly congruent typing result, in line with the species as 263
identified by MLST, which is considered one of the highest resolution methods apart from full-264
genome analysis. Nevertheless, all genes analyzed are located on different chromosomes, 265
except 2 of the MLST genes that both map on chromosome 31 (Fig. 2). Also, when the obtained 266
species groups were compared with those from a protein characterization through MLEE, a 267
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-14-
nearly perfect agreement was observed. This general concordance of all markers is in line with 268
the apparent absence of recent inter-species recombination evidence as pointed out by 269
multilocus sequence analysis (20), and the observed strong linkage disequilibrium in several 270
Leishmania species giving rise to stable multi-locus genotypes and the existence of near-clades, 271
which can persist despite frequent recombination events (37). Hence, given combinations of 272
gene variants are generally found in the same genomes, rather than being shuffled across 273
different genomes. 274
275
Even though sequencing is nowadays a common technique in reference laboratories and 276
Western diagnostic settings, our approach remains limited to such environments, as it is not yet 277
feasible to apply it on a routine basis elsewhere. For limited-resource labs technically less 278
demanding species-typing tools such as those mentioned in the Introduction are available, even 279
though in these settings treatment choice is often guided by economical rather than diagnostic 280
determinants (30). Global typing methods that allow identification to the species level are 281
primarily needed in epidemiological monitoring, genetic and clinical studies, in view of the 282
changing global ecology and related alterations in species distribution. Also infectious disease 283
clinics dealing with import leishmaniasis, the origin of which is not always clear as travelers 284
frequently visit various endemic areas or countries, will benefit from such techniques. 285
286
All PCRs from the individual markers used in this study have proven applicable directly 287
on clinical specimen (4,8,11-14,16), and allow species identification on the basis of 1 PCR 288
amplicon instead of 7 used in MLST, and without the need for parasite culturing, a technique 289
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-15-
that is time-consuming and often not successful. The hsp70 target was identified as the best 290
choice for high resolution species discrimination, as it identifies the same species and species 291
complexes as does the here applied gold standard MLST. A downside of hsp70 sequencing is the 292
size of the PCR product, which is 1286 base pairs. In our experience, some samples require 293
amplification of 2 overlapping PCR fragments, either in a single round or by nested PCR (16), but 294
in most cases the 1286 bp fragment can be amplified from a single PCR. In order to obtain a 295
reliable Sanger sequence read from both strands, 4 primers are needed. Noteworthy, three 296
shorter hsp70 fragments have been described and evaluated in clinical samples (13,14,16). 297
These are almost equally effective for sequence-based typing as the here used 1286 bp region 298
(Supplemental Figs. 6-8), although the most suitable fragment is governed by the required 299
discrimination. The mini-exon provides nearly the same resolution as does hsp70, but has as an 300
advantage that the amplicon is much smaller and more copies are present in the genome, which 301
may facilitate amplification of samples with low parasite load (5). Sequencing can be completed 302
with 2 primers, but mainly due to variations between the different copies in the same genome 303
(38), it is often technically challenging or impossible. ITS1 is equally advantageous in copy 304
number and size (8), but provides poor resolution in the L. (Viannia) subgenus. Also for this 305
marker, sequencing is challenging mainly because of homopolymer tracks. Finally, the 7SL-RNA 306
marker provides poor typing resolution both in the Old and New World, and is not 307
recommended for accurate species identification. It should be noted that an extended 7SL-RNA 308
fragment (11) could improve the resolution obtained with this marker. 309
310
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-16-
Finally, the full sequence typing presented in this paper does not necessarily agree with 311
an approach based on discrimination by single-nucleotide polymorphism (SNP) in the genes. As 312
a single mutation at the site of diagnostic species-specific SNP can lead to an erroneous typing 313
result, this is less so for full sequence analysis, which takes into account the sum of all sequence 314
variations in the genes, and is therefore less prone to errors. 315
316
In conclusion, we have shown that sequencing of a single gene is an accurate method for 317
discriminating medically important Leishmania species. The highest resolution is obtained from 318
hsp70 and the mini-exon, but other markers can be used depending on the origin of the sample 319
or in case typing is not necessary to the species level. Genetic markers essentially result in the 320
same species discrimination as does MLEE, allowing comparison of studies using various typing 321
methods. 322
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-17-
Acknowledgements 323
We thank Ilse Maes (Institute of Tropical Medicine, Antwerp, Belgium); Yolaine Wernet (Leiden 324
University Medical Center, Leiden, The Netherlands); Carla J.A. Wassenaar (Academic Medical 325
Center, Amsterdam, The Netherlands); Carola Schweynoch (Charité University Medicine, Berlin, 326
Germany); Christoph Stalder and Mark Finlayson (Swiss Tropical and Public Health Institute, 327
Basel, Switzerland) for technical assistance. Jean-Claude Dujardin is acknowledged for providing 328
feedback on the manuscript and the study design, and for financial support. Several parasite 329
isolates were kindly provided by the Institute of Tropical Medicine Alexander von Humboldt 330
(Lima, Peru). The study was performed in the context of the LeishMan study group on European 331
leishmaniasis (www.leishman.eu). 332
333
References 334
1. Romero GA, Guerra MV, Paes MG, Macedo VO. 2001. Comparison of cutaneous 335
leishmaniasis due to Leishmania (Viannia) braziliensis and L. (V.) guyanensis in Brazil: 336
therapeutic response to meglumine antimoniate. Am. J. Trop. Med. Hyg. 65:456-465. 337
2. Arevalo J, Ramirez L, Adaui V, Zimic M, Tulliano G, Miranda-Verastegui C, Lazo M, Loayza-338
Muro R, De Doncker S, Maurer A, Chappuis F, Dujardin JC, Llanos-Cuentas A. 2007. 339
Influence of Leishmania (Viannia) species on the response to antimonial treatment in 340
patients with American tegumentary leishmaniasis. J. Infect. Dis. 195:1846-1851. 341
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-18-
3. Alvar J, Velez ID, Bern C, Herrero M, Desjeux P, Cano J, Jannin J, den Boer M, WHO 342
Leishmaniasis control team. 2012. Leishmaniasis Worldwide and Global Estimates of Its 343
Incidence. PLoS. One. 7:e35671. 344
4. Marfurt J, Nasereddin A, Niederwieser I, Jaffe CL, Beck HP, Felger I. 2003. Identification and 345
differentiation of Leishmania species in clinical samples by PCR amplification of the 346
miniexon sequence and subsequent restriction fragment length polymorphism analysis. J. 347
Clin. Microbiol. 41:3147-3153. 348
5. Marfurt J, Niederwieser I, Makia ND, Beck HP, Felger I. 2003. Diagnostic genotyping of Old 349
and New World Leishmania species by PCR-RFLP. Diagn. Microbiol. Infect. Dis. 46:115-124. 350
6. el Tai NO, Osman OF, El Fari M, Presber W, Schönian G. 2000. Genetic heterogeneity of 351
ribosomal internal transcribed spacer in clinical samples of Leishmania donovani spotted 352
on filter paper as revealed by single-strand conformation polymorphisms and sequencing. 353
Trans. R. Soc. Trop. Med. Hyg. 94:575-579. 354
7. Nasereddin A, Bensoussan-Hermano E, Schönian G, Baneth G, Jaffe CL. 2008. Molecular 355
diagnosis of Old World cutaneous leishmaniasis and species identification by use of a 356
reverse line blot hybridization assay. J. Clin. Microbiol. 46:2848-2855. 357
8. Schönian G, Nasereddin A, Dinse N, Schweynoch C, Schallig HD, Presber W, Jaffe CL. 2003. 358
PCR diagnosis and characterization of Leishmania in local and imported clinical samples. 359
Diagn. Microbiol. Infect. Dis. 47:349-358. 360
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-19-
9. Talmi-Frank D, Nasereddin A, Schnur LF, Schönian G, Özensoy Töz S, Jaffe CL, Baneth G. 361
2010. Detection and identification of Old World Leishmania by high resolution melt 362
analysis. PLoS. Negl. Trop. Dis. 4:e581. 363
10. Odiwuor SO, Saad AA, De Doncker S., Maes I, Laurent T, El Safi S, Mbuchi M, Buscher P, 364
Dujardin JC, Van der Auwera G. 2011. Universal PCR assays for the differential detection 365
of all Old World Leishmania species. Eur. J. Clin. Microbiol. Infect. Dis. 30:209-218. 366
11. Stevenson LG, Fedorko DP, Zelazny AM. 2010. An enhanced method for the identification of 367
Leishmania spp. using real-time polymerase chain reaction and sequence analysis of the 368
7SL RNA gene region. Diagn. Microbiol. Infect. Dis. 66:432-435. 369
12. Zelazny AM, Fedorko DP, Li L, Neva FA, Fischer SH. 2005. Evaluation of 7SL RNA gene 370
sequences for the identification of Leishmania spp. Am. J. Trop. Med. Hyg. 72:415-420. 371
13. Fraga J, Veland N, Montalvo AM, Praet N, Boggild AK, Valencia BM, Arevalo J, Llanos-372
Cuentas A, Dujardin JC, Van der Auwera G. 2012. Accurate and rapid species typing from 373
cutaneous and mucocutaneous leishmaniasis lesions of the New World. Diagn. Microbiol. 374
Infect. Dis. 74:142-150. 375
14. Montalvo AM, Fraga J, Maes I, Dujardin JC, Van der Auwera G. 2012. Three new sensitive 376
and specific heat-shock protein 70 PCRs for global Leishmania species identification. Eur. J. 377
Clin. Microbiol. Infect. Dis. 31:1453-1461. 378
15. Garcia L, Kindt A, Bermudez H, Llanos-Cuentas A, De Doncker S, Arévalo J, Quispe Tintaya 379
KW, Dujardin JC. 2004. Culture-independent species typing of neotropical Leishmania for 380
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-20-
clinical validation of a PCR-based assay targeting heat shock protein 70 genes. J. Clin. 381
Microbiol. 42:2294-2297. 382
16. Van der Auwera G, Maes I, De Doncker S, Ravel C, Cnops L, Van Esbroeck M, Van Gompel 383
A, Clerinx J, Dujardin JC. 2013. Heat-shock protein 70 gene sequencing for Leishmania 384
species typing in European tropical infectious disease clinics. Euro. Surveill 18:pii=20543. 385
17. Rioux JA, Lanotte G, Serres E, Pratlong F, Bastien P, Perieres J. 1990. Taxonomy of 386
Leishmania. Use of isoenzymes. Suggestions for a new classification. Ann. Parasitol. Hum. 387
Comp. 65:111-125. 388
18. Mauricio IL, Yeo M, Baghaei M, Doto D, Pratlong F, Zemanová E, Dedet JP, Lukeš J, Miles 389
MA. 2006. Towards multilocus sequence typing of the Leishmania donovani complex: 390
resolving genotypes and haplotypes for five polymorphic metabolic enzymes (ASAT, GPI, 391
NH1, NH2, PGD). Int. J. Parasitol. 36:757-769. 392
19. Zemanová E, Jirku M, Mauricio IL, Horák A, Miles MA, Lukeš J. 2007. The Leishmania 393
donovani complex: genotypes of five metabolic enzymes (ICD, ME, MPI, G6PDH, and FH), 394
new targets for multilocus sequence typing. Int. J. Parasitol. 37:149-160. 395
20. El Baidouri F, Diancourt L, Berry V, Chevenet F, Pratlong F, Marty P, Ravel C. 2013. Genetic 396
structure and evolution of the Leishmania genus in Africa and Eurasia: what does MLSA 397
tell us. PLoS. Negl. Trop. Dis. 7:e2255. 398
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-21-
21. Odiwuor S, Veland N, Maes I, Arévalo J, Dujardin JC, Van der Auwera G. 2012. Evolution of 399
the Leishmania braziliensis species complex from amplified fragment length 400
polymorphisms, and clinical implications. Infect. Genet. Evol. 12:1994-2002. 401
22. Veland N, Boggild AK, Valencia C, Valencia BM, Llanos-Cuentas A, Van der Auwera G, 402
Dujardin JC, Arevalo J. 2012. Leishmania (Viannia) species identification on clinical 403
samples from cutaneous leishmaniasis patients in Peru: assessment of a molecular 404
stepwise approach. J. Clin. Microbiol. 50:495-498. 405
23. Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5: molecular 406
evolutionary genetics analysis using maximum likelihood, evolutionary distance, and 407
maximum parsimony methods. Mol. Biol. Evol. 28:2731-2739. 408
24. Thompson JD, Higgins DG, Gibson TJ. 1994. CLUSTAL W: improving the sensitivity of 409
progressive multiple sequence alignment through sequence weighting, position-specific 410
gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. 411
25. Kellina OI, Shurkhal AV, Strelkova MV, Passova OM, Rakitskaia TA. 1985. [Isoenzyme 412
characteristics of strains of the causative agent of visceral leishmaniasis in the territory of 413
the USSR]. Med. Parazitol. (Mosk) 1985:11-15. 414
26. Leblois R, Kuhls K, Francois O, Schönian G, Wirth T. 2011. Guns, germs and dogs: On the 415
origin of Leishmania chagasi. Infect. Genet. Evol. 11:1091-1095. 416
27. Mauricio IL, Stothard JR, Miles MA. 2000. The strange case of Leishmania chagasi. Parasitol. 417
Today 16:188-189. 418
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-22-
28. Kuhls K, Alam MZ, Cupolillo E, Ferreira GE, Mauricio IL, Oddone R, Feliciangeli MD, Wirth T, 419
Miles MA, Schönian G. 2011. Comparative microsatellite typing of new world Leishmania 420
infantum reveals low heterogeneity among populations and its recent old world origin. 421
PLoS. Negl. Trop. Dis. 5:e1155. 422
29. World Health Organization. 2010. Control of the leishmaniasis: report of a meeting of the 423
WHO Expert Committee on the Control of Leishmaniases, Geneva, 22-26 March 2010. 424
World Health Organization, Geneva, Switzerland. 425
30. Gradoni L, Soteriadou K, Louzir H, Dakkak A, Toz SO, Jaffe C, Dedet JP, Campino L, 426
Cañavate C, Dujardin JC. 2008. Drug regimens for visceral leishmaniasis in Mediterranean 427
countries. Trop. Med. Int. Health 13:1272-1276. 428
31. Odiwuor S, Vuylsteke M, De Doncker S, Maes I, Mbuchi M, Dujardin JC, Van der Auwera G. 429
2011. Leishmania AFLP: paving the way towards improved molecular assays and markers 430
of diversity. Infect. Genet. Evol. 11:960-967. 431
32. Lukeš J, Mauricio IL, Schönian G, Dujardin JC, Soteriadou K, Dedet JP, Kuhls K, Quispe-432
Tintaya KW, Jirku M, Chocholová E, Haralambous C, Pratlong F, Oborník M, Horák A, 433
Ayala FJ, Miles MA. 2007. Evolutionary and geographical history of the Leishmania 434
donovani complex with a revision of current taxonomy. Proc. Natl. Acad. Sci. USA 435
104:9375-9380. 436
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-23-
33. Zemanová E, Jirku M, Mauricio IL, Miles MA, Lukeš J. 2004. Genetic polymorphism within 437
the Leishmania donovani complex: correlation with geographic origin. Am. J. Trop. Med. 438
Hyg. 70:613-617. 439
34. Kuhls K, Keilonat L, Ochsenreither S, Schaar M, Schweynoch C, Presber W, Schönian G. 440
2007. Multilocus microsatellite typing (MLMT) reveals genetically isolated populations 441
between and within the main endemic regions of visceral leishmaniasis. Microbes. Infect. 442
9:334-343. 443
35. Bañuls AL, Dujardin JC, Guerrini F, De Doncker S, Jacquet D, Arevalo J, Noel S, Le Ray D, 444
Tibayrenc M. 2000. Is Leishmania (Viannia) peruviana a distinct species? A MLEE/RAPD 445
evolutionary genetics answer. J. Eukaryot. Microbiol. 47:197-207. 446
36. Nolder D, Roncal N, Davies CR, Llanos-Cuentas A, Miles MA. 2007. Multiple hybrid 447
genotypes of Leishmania (Viannia) in a focus of mucocutaneous leishmaniasis. Am. J. Trop. 448
Med. Hyg. 76:573-578. 449
37. Tibayrenc M, Ayala FJ. 2013. How clonal are Trypanosoma and Leishmania? Trends 450
Parasitol. 29:264-269. 451
38. van Thiel PP, van Gool T, Faber WR, Leenstra T, Kager PA, Bart A. 2011. Variation in clinical 452
presentation and genotype of causative Leishmania major strain in cutaneous 453
leishmaniasis in north and south Afghanistan. Am. J. Trop. Med. Hyg. 85:60-63. 454
455
456
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-24-
Table 1: Species designation and strain origin
Species a Origin b
L. aethiopica (4) Ethiopia (3); Kenya
L. tropica (4) Egypt; Morocco (2); Yemen
L. archibaldi (4) Italy; Kenya; Sudan (2)
L. donovani (7) China; Ethiopia; India; Morocco; Sudan (2); former USSR
L. infantum (9) Algeria; Egypt; Israel; Panama c ; Portugal; Spain (4)
L. major (11) Algeria; Burkina Faso (2); Cameroon; India; Israel; Jordan;
Mali; Senegal; Sudan; former USSR
L. mexicana (3) Belize; Ecuador; Mexico
L. amazonensis (4) Brazil; Colombia; Panama; Peru
L. garnhami (1) Venezuela
L. gerbilli (1) Former USSR
L. turanica (1) Former USSR
L. lainsoni (3) Bolivia; Peru (2)
L. naiffi (3) Brazil; French Guiana; Unknown origin
L. braziliensis (9) Bolivia; Brazil (2); Colombia; Peru (5)
L. guyanensis (6) Colombia; Ecuador (2); French Guiana (3)
L. peruviana (3) Peru (3)
L. panamensis (1) Costa Rica
a Most species were determined by MLEE, except when otherwise mentioned
in Supplemental Table S1. The number given between brackets is the total
number of isolates tested (74 in total). b The number of isolates from each country is given between brackets, except
when only 1 isolate was used. Details of all the strains with their WHO code
are listed in Supplemental Table S1. This table also lists the GenBank
accession numbers of all sequences. c New World Leishmania infantum is synonym for Leishmania chagasi.
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-25-
Figure legends 457
Fig. 1. MLST analysis and comparison with individual markers. The Neighbor-Joining dendrogram was 458
built from a concatenated alignment of 7 household genes (20). The numbers shown at the internodes 459
indicate the bootstrap support in percentages, whereby values lower than 70% are not shown. The 460
recognized species are indicated by their bootstrap value, and separated by dotted horizontal lines. The 461
dissimilarity scale is depicted in the top left corner, in substitutions per nucleotide. The vertical black 462
lines on the right hand side depict the clusters as seen in dendrograms from the 4 typing markers 463
analyzed in this paper, which are indicated on top. For hsp70 and 7SL-RNA, the numbers accompanying 464
these lines indicate the bootstrap support of the respective clades. For ITS1 and ME, the size range of the 465
PCR products is given for each species, as determined from the complete nucleotide sequences. As for 466
L. lainsoni no complete sequences were obtained from the mini-exon, the size range is an estimate from 467
agarose gel analysis of the PCR amplicons. Strains are identified using their WHO code, as in 468
Supplemental Table S1, followed by the species as determined by MLEE, if available (maj: L. major; ger: 469
L. gerbilli; tur: L. turanica; aet: L. aethiopica; tro: L. tropica; don: L. donovani; arc: L. archibaldi; inf: 470
L. infantum; mex: L. mexicana; ama: L. amazonensis; gar: L. garnhami; nai: L. naiffi; bra: L. braziliensis; 471
guy: L. guyanensis; pan: L. panamensis). Strains indicated with # were not included in the 7SL-RNA 472
analysis because no sequences were obtained; ~ indicates a lacking ITS1 sequence; * means no complete 473
ME sequence could be determined; ° indicates some 5’ nucleotides were lacking from the ME sequence. 474
OWL: Old World L. (Leishmania) subgenus; NWL: New World L. (Leishmania) subgenus; NWV: New World 475
L. (Viannia) subgenus, indicated on the branch leading to these taxa. 476
477
478
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from

-26-
Fig. 2. Chromosomal location of the genetic typing markers. The location of the 7 MLST markers and the 479
4 single gene species typing markers analyzed in this paper is shown relative to the L. major 480
chromosomes. The 7 MLST genes are depicted in grey below the indicated mapped positions, the 4 481
targets used as individual typing markers are in black. The size of the PCR products without the primers is 482
indicated after the gene names (nt = nucleotides). 483
484 485
on March 18, 2018 by guest
http://jcm.asm
.org/D
ownloaded from
Related Documents






![Tagging the juvenile locus in soybean [Glycine max (L.) Merr.] with molecular markers](https://static.cupdf.com/doc/110x72/63325e504e0143040300a382/tagging-the-juvenile-locus-in-soybean-glycine-max-l-merr-with-molecular-markers.jpg)