O riginal A rticle Stem Cells 2005;23:1314–1323 www.StemCells.com DNA Methylation Is Required for Silencing of Ant4 , an Adenine Nucleotide Translocase Selectively Expressed in Mouse Embryonic Stem Cells and Germ Cells Nemanja RodiĆ, a Masahiro Oka, a Takashi Hamazaki, a Matthew R. Murawski, a Marda Jorgensen, b Danielle M. Maatouk, c James L. Resnick, c En Li, d Naohiro Terada a,b a Department of Pathology, b Shands Cancer Center, c Department of Molecular Genetics and Microbiology, University of Florida College of Medicine, Gainesville, Florida, USA; d Cardiovascular Research Center, Massachusetts General Hospital, Harvard Medical School, Charlestown, Massachusetts, USA Key Words. Embryonic stem cell • DNA methylation • Differentiation • Germ cell Adenine nucleotide translocase • Gene repression Correspondence: Naohiro Terada, M.D., Ph.D., Department of Pathology, University of Florida College of Medicine, 1600 SW Archer Road, Gainesville, Florida 32610, USA. Telephone: 352-392-2696; Fax: 352-392-6249; e-mail: [email protected] Received March 16, 2005; accepted for publication May 10, 2005; first published online in Stem Cells EXPRESS July 28, 2005. ©AlphaMed Press 1066-5099/2005/$12.00/0 doi: 10.1634/stemcells.2005-0119 Abstract The capacity for cellular differentiation is governed not only by the repertoire of available transcription factors but by the accessibility of cis-regulatory elements. Studying changes in epigenetic modifications during stem cell differentiation will help us understand how cells maintain or lose differentiation potential. We investigated changes in DNA methylation dur- ing the transition of pluripotent embryonic stem cells (ESCs) into differentiated cell types. Using a methylation-sensitive restriction fingerprinting method, we identified a novel ade- nine nucleotide (ADP/ATP) translocase gene, Ant4, that was selectively hypomethylated and expressed in undifferentiated mouse ESCs. In contrast to other pluripotent stem cell–spe- cific genes such as Oct-4 and Nanog, the Ant4 gene was readily derepressed in differentiated cells after 5-aza-2'-deoxycyti- dine treatment. Moreover, expression of de novo DNA meth- yltransferases Dnmt3a and Dnmt3b was essential for repres- sion and DNA methylation of the Ant4 gene during ESC differ- entiation. Although the deduced amino acid sequence of Ant4 is highly homologous to the previously identified Ant isoforms, the expression of Ant4 was uniquely restricted to developing gametes in adult mice, and its promoter hypomethylation was observed only in testis. Additionally, Ant4 was expressed in primordial germ cells. These data indicate that Ant4 is a plu- ripotent stem cell– and germ cell–specific isoform of adenine nucleotide translocase in mouse and that DNA methylation plays a primary role in its transcriptional silencing in somatic cells. Stem Cells 2005;23:1314–1323 Introduction DNA methylation, or the addition of a methyl group to the 5'- position of cytosine within the CpG dinucleotide, is a heri- table modification that contributes to gene silencing. Most CpG sites in mammalian cells are methylated in a nonrandom fashion. For instance, repetitive and parasitic elements tend to be hypermethylated, whereas CpG island–associated promot- ers are usually hypomethylated [1]. The complex process of DNA methylation has been proven essential for normal devel- opment [2], X-chromosome inactivation [3], imprinting [4], This material is protected by U.S. Copyright law. Unauthorized reproduction is prohibited. For reprints contact: [email protected]

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
Original Article
Stem Cells 2005;23:1314–1323 www.StemCells.com
DNA Methylation Is Required for Silencing of Ant4, an Adenine
Nucleotide Translocase Selectively Expressed in Mouse Embryonic
Stem Cells and Germ Cells
Nemanja RodiĆ,a Masahiro Oka,a Takashi Hamazaki,a Matthew R. Murawski,a Marda Jorgensen,b Danielle M. Maatouk,c James L. Resnick,c En Li,d Naohiro Teradaa,b
aDepartment of Pathology, bShands Cancer Center, cDepartment of Molecular Genetics and Microbiology,
University of Florida College of Medicine, Gainesville, Florida, USA; dCardiovascular Research Center,
Massachusetts General Hospital, Harvard Medical School, Charlestown, Massachusetts, USA
Key Words. Embryonic stem cell • DNA methylation • Differentiation • Germ cellAdenine nucleotide translocase • Gene repression
Correspondence: Naohiro Terada, M.D., Ph.D., Department of Pathology, University of Florida College of Medicine, 1600 SW Archer Road, Gainesville, Florida 32610, USA. Telephone: 352-392-2696; Fax: 352-392-6249; e-mail: [email protected] Received March 16, 2005; accepted for publication May 10, 2005; first published online in Stem Cells EXPRESS July 28, 2005. ©AlphaMed Press 1066-5099/2005/$12.00/0 doi: 10.1634/stemcells.2005-0119
AbstractThe capacity for cellular differentiation is governed not only by the repertoire of available transcription factors but by the accessibility of cis-regulatory elements. Studying changes in epigenetic modifications during stem cell differentiation will help us understand how cells maintain or lose differentiation potential. We investigated changes in DNA methylation dur-ing the transition of pluripotent embryonic stem cells (ESCs) into differentiated cell types. Using a methylation-sensitive restriction fingerprinting method, we identified a novel ade-nine nucleotide (ADP/ATP) translocase gene, Ant4, that was selectively hypomethylated and expressed in undifferentiated mouse ESCs. In contrast to other pluripotent stem cell–spe-cific genes such as Oct-4 and Nanog, the Ant4 gene was readily derepressed in differentiated cells after 5-aza-2'-deoxycyti-
dine treatment. Moreover, expression of de novo DNA meth-yltransferases Dnmt3a and Dnmt3b was essential for repres-sion and DNA methylation of the Ant4 gene during ESC differ-entiation. Although the deduced amino acid sequence of Ant4 is highly homologous to the previously identified Ant isoforms, the expression of Ant4 was uniquely restricted to developing gametes in adult mice, and its promoter hypomethylation was observed only in testis. Additionally, Ant4 was expressed in primordial germ cells. These data indicate that Ant4 is a plu-ripotent stem cell– and germ cell–specific isoform of adenine nucleotide translocase in mouse and that DNA methylation plays a primary role in its transcriptional silencing in somatic cells. Stem Cells 2005;23:1314–1323
IntroductionDNA methylation, or the addition of a methyl group to the 5'-
position of cytosine within the CpG dinucleotide, is a heri-
table modification that contributes to gene silencing. Most
CpG sites in mammalian cells are methylated in a nonrandom
fashion. For instance, repetitive and parasitic elements tend to
be hypermethylated, whereas CpG island–associated promot-
ers are usually hypomethylated [1]. The complex process of
DNA methylation has been proven essential for normal devel-
opment [2], X-chromosome inactivation [3], imprinting [4],
This material is protected by U.S. Copyright law. Unauthorized reproduction is prohibited.
For reprints contact: [email protected]
Rodić, Oka, Hamazaki et al. 1315
and the suppression of parasitic DNA sequences [5]. Although
DNA methylation is a modification that promotes genomic
integrity and ensures proper temporal and spatial gene expres-
sion during development, it also associates with malignancy
when aberrantly controlled. Both hypermethylation and
hypomethylation have been attributed to cancer development
[6, 7]. DNA methylation is proposed to prevent the binding
of transcription factors and recruit repressor complexes that
induce the formation of inactive chromatin complexes [8].
For example, it was recently shown that methylation of a CpG
dinucleotide within the glial fibrillary acidic protein promoter
prevents STAT3 binding [9]. In another instance, DNA meth-
ylation–mediated gene silencing is dependent on the pres-
ence of methyl-CpG binding protein, MeCP2, which forms
a complex with histone deacetylases and a repressor protein,
mSin3A, to repress transcription in a methylation-dependent
manner [10]. It is still unclear in most cases, however, whether
DNA methylation is a causal event in gene silencing or, rather,
a consequence of gene silencing.
It is becoming increasingly clear that epigenetic modifica-
tions play a critical role in the regulation of gene expression in
many cellular processes [8, 11]. Studying changes in epigenetic
modifications during stem cell differentiation will help us under-
stand how cells maintain or lose differentiation potential. In the
present study, we attempted to identify differentially methylated
loci that are hypomethylated in undifferentiated embryonic stem
cells (ESCs) and become hypermethylated after differentiation.
Murine ESCs are originally derived from the inner cell mass of
a developing blastocyst and have the ability to differentiate into
all cell types of an adult animal [12]. Pluripotency of ESCs can be
maintained in vitro when the cells are cultured in a serum-con-
taining medium supplemented with leukemia inhibitory factor
(LIF) [13]. When LIF is removed from the medium, the ESCs
begin to differentiate in vitro into all three embryonic germ lay-
ers. This in vitro ESC differentiation system serves as an excel-
lent model to study the regulation of gene expression required for
stem cell self-renewal and pluripotency [14–16]. Recent studies
on molecules involved in epigenetic modifications have revealed
a unique expression pattern of DNA methyltransferases [17], his-
tone deacetylases [18], and methyl-binding proteins [19] in ESCs.
ESCs also have a differential genome-wide DNA methylation
pattern compared with their descendant differentiated cells [20,
21]. However, exact genomic loci of such differentially methyl-
ated regions remain unknown.
Using methylation-sensitive restriction fingerprinting
(MSRF), we identified a novel gene encoding an adenine nucle-
otide (ADP/ATP) translocase homologue that is specifically
expressed in undifferentiated ESCs and germ cells. Further-
more, we show that DNA methylation, but not the availability
of transcription factors, is the dominant factor restricting the
gene’s expression.
Materials and Methods
In Vitro ESC DifferentiationMouse ESC lines R1, J1, Dnmt3a-null, Dnmt3b-null, and
Dnmt3a-Dnmt3b double-null ESCs [22] were maintained on gel-
atin-coated tissue culture dishes in Dulbecco’s modified Eagle’s
medium optimized for ESCs (Invitrogen, Carlsbad, CA, http://
www.invitrogen.com) containing 1,000 U/ml recombinant mouse
LIF (ESGRO; Chemicon, Temecula, CA, http://www.chemicon.
com), 10% knockout serum replacement (Invitrogen), 1% fetal
calf serum (Atlanta Biologicals, Lawrenceville, GA, http://www.
atlantabio.com), 20 mM HEPES (Invitrogen), 300 μM monothio-
glycerol (Sigma-Aldrich, St. Louis, http://www.sigmaaldrich.
com), 100 U/ml penicillin, 100 μg/ml streptomycin (Invitrogen),
and 2 mM L-glutamine (Invitrogen). In vitro ESC differentiation
was induced in an Iscove’s modified Dulbecco’s medium (Invit-
rogen) containing 20% fetal calf serum, 2 mM L-glutamine, 100
U/ml penicillin, 100 μg/ml streptomycin, and 300 μM monothio-
glycerol. The embryoid body (EB) formation was induced in a
hanging drop containing approximately 2,000 cells in differentia-
tion medium for 2 days as described previously [23].
MSRFMSRF was performed according to the original method estab-
lished by Huang et al. [24]. Briefly, ESCs and EBs were harvested
by gentle cell scraping followed by a 5-minute centrifugation in a
bench-top centrifuge. Genomic DNA was extracted using Wizard
Genomic DNA Purification Kit (Promega, Madison, WI, http://
www.promega.com). Extracted DNA was digested with a methyla-
tion-insensitive restriction enzyme (MseI) either by itself or in com-
bination with a methylation-sensitive restriction enzyme, BstUI, at
10 U per 1 μg of DNA for each enzyme. Digestion with BstUI was
performed for 2 hours at 60°C, followed by overnight incubation
with MseI at 37°C. The polymerase chain reaction (PCR) was per-
formed in 20-μl reaction mixtures containing 2 μl of digested DNA
(50–100 ng), 0.4 μM of primers, 1.25 U of HotMasterTaq DNA
polymerase (Eppendorf, Hamburg, Germany), 200 μM deoxy-
nucleotide triphosphates, 1 mCi/μl [α-32P] dCTP (3,000 Ci/mmol,
Amersham, Piscataway, NJ), and 2 μl of ×10 reaction buffer. The
primers used in this study were Bs-1 (5'-AGCGGCCGCG), Bs-2
(5'-GCCCCCGCGA), Bs-3 (5'-CGGGGCGCGA), and Bs-4 (5'-
ACCCCACCCG). The PCR reaction consisted of an initial dena-
turing step for 5 minutes at 94°C and 30 cycles of the following: 2
minutes at 94°C, 1 minute at 40°C, and 2 minutes at 72°C. A final
step at 72°C was 8 minutes. After PCR amplification, 5 μl of each
sample was mixed with 1 μl of ×6 loading dye solution. Each sample
was then separated in a 4.5% nondenaturing polyacrylamide gel.
All reactions were run as duplicates to account for loading error.
Wet gels were laid on 3M-filter paper, wrapped with plastic wrap,
and exposed to Kodak X-OMAT LS film (Kodak, New Haven, CT,
http://www.kodak.com) for 24–48 hours at −80°C.
1316 DNA Methylation–Dependent Repression of Ant4
Cloning and SequencingDNA segments of interest were excised from polyacrylamide
gels using a sterile scalpel. DNA was eluted by incubation of
gel fragments in 50 μl of sterile deionized water for 10 minutes
at 100°C. Eluted DNA was reamplified by the identical prim-
ers and PCR conditions as used in the MSRF method. Ream-
plified DNA fragments were excised from the gel, ligated into
a TA-cloning vector using pCRII-Topo Cloning Kit (Invit-
rogen), and sequenced. To determine the identity of resulting
DNA sequences, searches were performed against the follow-
ing mouse genomic databases: http://www.ensambl.org, http://
genome.ucsc.edu, and http://www.ncbi.nlm.nih.gov/genome/
seq/MmBlast.html. All of the recombinant DNA experiments
here and below were performed under the National Institutes of
Health guidelines.
Bisulfite Sequencing and Combined Bisulfite Restriction AnalysisDNA was extracted from ESCs, EBs, and adult tissues using
the DNA Wizard Genomic DNA Purification Kit (Promega).
Mouse testes were decapsulated, and seminiferous tubules
were collected for analysis. A bisulfite reaction was performed
using EZ DNA Methylation Kit (Zymo Research, Orange,
CA, http://www.zymoresearch.com). Up to 2 μg of genomic
DNA was used for conversion with the bisulfite reagent.
Approximately 80 ng of bisulfite-converted DNA was used as
a template for each PCR analysis. Primers used for Ant4 com-
bined bisulfite restriction analysis (COBRA) and for bisulfite
sequencing were 5'-TTGTTGTGTATTGATTGAGTATG
and 5'-ACACTAAAAAAAACTAAAAAACC (40 cycles).
Primers used for bisulfite sequencing were 5'-AAGGTGTGT-
GTTATTATTGTTTGATGT and 5'-TACCCCTCATCTAT-
CATATTCCCTA; 5'-TTGAGGAGGAGTTAATAAGTT-
TAGGG and 5'-CCAAAAAACACACTCTAACCAAATAC;
5'-GTAGGTAAATTAATTGTGGATTAAATAGTA and
5'-TACACAACAACTTTTACAAAAAAAC; 5'-AGTTGTTGT-
GTATTGATTGAGTATG and 5'-TCTTTAAAAACTACTTCT-
TAAAAAATTC; and 5'-TTTTTAAGAAGTAGTTTTTA-
AAGAAGG and 5'-AAACAATCCAACATACCCTTATAAC.
PCR fragments were cloned into pCRII-TOPO cloning vector
(Invitrogen), and individual clones were sequenced.
Reverse Transcription–Polymerase Chain ReactionTotal RNA was isolated from ESCs and EBs using RNAque-
ous kit (Ambion, Austin, TX, http://www.ambion.com). Two
micrograms of RNA was used as a template for reverse tran-
scription (RT) reactions using SuperScript Synthesis for First
Strand Synthesis Kit (Invitrogen). Sequences of forward and
reverse primer pairs were as follows: Ant4 (5'-TGGAGCAA-
CATCCTTGTGTG, 5'-AGAAATGGGGTTTCCTTTGG),
Oct-4 (5'-TGGAGACTTTGCAGCCTGAG, 5'-TGAATGCAT-
GGGAGAGCCCA), Nanog (5'-AGGGTCTGCTACTGAGA-
TGCTCTG, 5'-CAACCACTGGTTTTTCTGCCACCG), Ttr
(5'-CTCACCACAGATGAGAAG, 5'-CCGTGAGTCTCT-
CAATTC), Fgf-5 (5'-AAAGTCAATGGCTCCCACGAA, 5'-
CTTCAGTCTGTACTTCACTGG), β-actin (5'-TTCCTTCTT-
GGGTATGGAAT, 5'-GAGCAATGATCTTGATCTTC),
Gapdh (5'-CCCTTCATTGACCTCACTACATGG, 5'-CCT-
GCTTCACCACCTTCTTGATGTC), and Hprt (5'-GCTGGT-
GAAAAGGACCTCT, 5'-CACAGG ACTAGAACACCTGC).
Northern BlottingNorthern blotting was performed using multiple premade blots
(Clontech, Palo Alto, CA, http://www.clontech.com). Briefly,
the membrane was preincubated with 5 ml of ExpressHyb
Hybridization Solution at 68°C for 1 hour. Seventy-five nano-
grams of the full-length Ant4 or β-actin cDNA fragment was
radio-labeled with [α-32P] dCTP nucleotides. Ant1 and Ant2
probes were PCR-amplified using the following primers: Ant1
(5'-GGCGCTACTTTGCTGGTAAC, 5'-GCAATCTTCCTC-
CAGCAGTC), Ant2 (5'-CAGCTGGATGATTGCACAGT,
5'-CAAGCCCAGAGAATCTGTCC). Unincorporated nucleo-
tides were removed using ProbeQuant G-50 Micro Column
(Amersham Biosciences, Piscataway, NJ, http://www.amer-
sham.com). Heat-denatured and briefly chilled probe (~20
ng/ml) was added to the hybridization solution. The membrane
was incubated with gentle shaking for 24 hours at 68°C and
washed twice for half an hour per wash in ×2 standard saline
citrate (SSC)/0.05% SDS at room temperature and twice in
×0.1 SSC/0.1% SDS at 50°C. The membrane was wrapped with
plastic wrap and exposed to X-OMAT Kodak film at −80°C for
18–24 hours.
ImmunohistochemistryRabbit polyclonal antibodies were raised against mouse Ant4
peptide (1-MSNESSKKQSSKKALFD-17) and purif ied
through an affinity column using the same peptide (Sigma
Genosys, The Woodlands, TX, http://www.sigma-genosys.
com/index.asp). Mouse ovary, testis, and liver were harvested
from 3-month-old BALB/c mice and immediately frozen into
optimal cutting temperature blocks (Tissue-Tek, Torrance, CA,
http://www.sakura.com). Frozen sections were cut at 5 μm,
air-dried for 2 hours, and then fixed in acetone for 10 minutes
before being rehydrated and blocked for endogenous peroxidase
activity. After a serum-blocking step, affinity-purified rabbit
polyclonal anti-Ant4 antibodies were applied at 2 μg/ml and
incubated overnight at 4°C. Staining was achieved using the
EnVision+ HRP kit (DakoCytomation, Glostrup, Denmark,
http://www.dakocytomation.com) following the manufactur-
er’s directions. Positive signal was detected with DAB+ (Dako-
Cytomation), and slides were counterstained using Light Green
SF Yellowish (Sigma-Aldrich).
Rodić, Oka, Hamazaki et al. 1317
Primordial Germ Cell PreparationPrimordial germ cells (PGCs) were obtained from timed matings
of B6C3F1 mice purchased from Jackson Laboratory (Bar Har-
bor, ME, http://www.jax.org). Noon of the day on which a mating
plug was first observed was taken to be 0.5 days post coitus (dpc).
PGCs were immunomagnetically purified using anti-SSEA1
antibody from isolated urogenital ridges as described [25]. Geni-
tal ridges at 12.5 dpc were sex-separated by visual inspection
for testis cords. The immunodepleted fraction refers to cells not
retained on the magnetic column.
Results
Identification of a CpG-Rich DNA Sequence That Undergoes De Novo DNA Methylation During ESC DifferentiationWe investigated changes in DNA methylation during the transi-
tion of ESCs into differentiated cell types using an MSRF method.
Mouse ESCs were differentiated in a culture medium without LIF
using a hanging drop method. DNA was extracted from undif-
ferentiated ESCs (day 0) and differentiated EBs (days 5 and
10). The DNA was then digested by a combination of methyla-
tion-sensitive (BstUI) and insensitive (MseI) restriction enzymes
and amplified by PCR using CpG-rich 10-mer primers (Fig. 1A).
Using a combination of four different CpG-rich primers, we iden-
tified a total of eight bands that exhibited differential methylation
patterns during ESC differentiation (data not shown). The bands
were excised from the gels, and their DNA sequences were deter-
mined. Database searches revealed that one of these methylated
fragments (methylated fragment 1, MF1 in Figs. 1B, 1C) mapped
to the exon 1/intron 1 boundary of a previously uncharacterized
gene (RIKEN cDNA 1700034J06). The other seven DNA frag-
ments were localized outside of any CpG islands and were derived
from various genomic regions. Because the MF1 DNA fragment
mapped to a genomic region that partially overlapped with a CpG
island, we decided to further analyze the associated gene.
The associated gene on mouse chromosome 13 did not
contain a classic TATA box sequence at its 5'-flanking region.
Because TATA-less genes are often associated with multiple
transcriptional initiation sites, we performed a 5'RACE (rapid
amplification of cDNA ends) reaction using cDNA isolated from
undifferentiated ESCs. 5'RACE identified five different tran-
scriptional initiation sites, four of which were previously reported
in Expressed Sequence Tag (EST) databases (data not shown).
Because there are multiple transcriptional initiation sites, nucleo-
tide positions are marked relative to the translation initiation site
(+1) throughout the manuscript. The major transcript initiated at
−60 bp, whereas the longest transcript initiated 304 bp upstream
of the translation initiation site.
The MF1-associated gene was predicted to encode a 320-amino-
acid protein and shared high amino acid sequence homology with
the other mouse adenine nucleotide translocase (Ant) proteins pre-
viously identified (70.1% and 69.1% overall amino acid identity to
Ant1 and Ant2 isoform, respectively) (Fig. 2A). The MF1-associated
Figure 1. Identification of differentially methylated CpG-rich frag-
ment in ES cells and EBs. (A): MSRF. Genomic DNA was prepared
from undifferentiated ES cells and differentiating EBs (days 5 and
10). DNA was digested either by MseI alone or MseI and BstUI and
subjected to PCR amplification using CpG-rich 10-mer primers in the
presence of radiolabeled dCTP. Amplified DNA fragments were sep-
arated in 4.5% polyacrylamide gels. In MseI/BstUI double-digested
samples, the intensity of a band indicated by arrowhead (MF1)
increased after ES cell differentiation. (B): Nucleotide sequence of
methylated fragment 1 (MF1). The DNA band was cloned into a PCR
cloning vector and sequenced. Bold letters indicate primers used,
whereas BstUI sites are underlined. MF1 corresponds to the exon
1/intron 1 boundary region of a previously uncharacterized gene on
mouse chromosome 13; black solid boxes represent exons predicted
by Expressed Sequence Tag and Ensemble mouse genome databases.
The translation initiation site is marked as +1. The arrow represents the
major transcription initiation site (−60 bp). Abbreviations: EB, embry-
oid body; ESC, embryonic stem cell; MseI, methylation-insensitive
restriction enzyme; MSRF, methylation-sensitive restriction finger-
printing; PCR, polymerase chain reaction.
1318 DNA Methylation–Dependent Repression of Ant4
gene also contained three tandem repeats of a domain of approxi-
mately 100 residues, each domain containing two transmembrane
regions, a characteristic shared by all members of the Ant family
[26]. Because another isoform of Ant, ANT3, has been reported in
human [27], we tentatively named the present gene adenine nucleo-
tide translocase 4 (Ant4). All known mammalian Ant isoforms
(unlike plant adenine nucleotide translocases) lack an N-terminal
mitochondrial localization sequence, yet they localize to mitochon-
dria [28]. In agreement with this observation, Ant4 also does not
contain a classic mitochondrial localization sequence. However, it
did localize to mitochondria when N-terminal FLAG-tagged Ant4
was expressed in NIH3T3 fibroblast cells (data not shown).
The human orthologue of ANT4 is located on chromosome
4q28.1, and its deduced amino acid sequence shares 85.9% over-
all amino acid identity with mouse Ant4 (Fig. 2B). In addition,
the genomic architecture with six exons is conserved between
mouse and human (data not shown). It is notable that the human
genome contains at least seven ANT pseudogenes on the X chro-
mosome; however, in contrast to ANT4, such pseudogenes do
not reveal conserved exon/intron architecture or translatable
coding regions [29].
Ant4 Promoter Locus Undergoes De Novo DNA Methylation After ESC DifferentiationTo determine DNA methylation patterns across the Ant4 pro-
moter locus during ESC differentiation, genomic DNAs from
ESCs or day-10 EBs were treated with bisulfite and subjected
to sequence analysis. We analyzed the Ant4 promoter region
that encompasses a total of 47 CpG dinucleotides, from 516
bp upstream of the translation initiation site to the 3'-end of
exon 1. Sequencing of individual bisulfite-converted genomic
DNAs revealed that the Ant4 promoter and associated CpG
island region were mostly unmethylated in undifferentiated
ESCs (Fig. 3). In contrast, EBs at day 10 showed significant
hypermethylation around the Ant4 promoter region, indicating
that after ESC differentiation, the Ant4 promoter undergoes de
novo DNA methylation. The further upstream region (>1 kb) of
the Ant4 promoter outside of the CpG island revealed hyper-
methylation regardless of the differentiation status of ESCs
(Fig. 3). Ant4 promoter methylation was confirmed by the
quantitative method of COBRA [30]. The frequency of methyl-
ation at a specific CpG site overlapping an HhaI restriction site
(−169 bp) was evaluated by the COBRA assay. In agreement
with the bisulfite sequencing data, we detected low levels of
DNA methylation in ESCs and high methylation levels (~75%)
in EBs at day 10.
Ant4 mRNA Expression Is Downregulated After ESC DifferentiationWe then examined Ant4 mRNA expression levels in ESCs and
EBs using semiquantitative RT-PCR. Ant4 mRNA was easily
detectable in undifferentiated ESCs, whereas the relative abun-
dance of Ant4 mRNA decreased after ESC differentiation (Fig.
4A). β-Actin expression was relatively unchanged during ESC dif-
ferentiation. We then used the DNA demethylating agent 5-aza-2'-
deoxycytidine (5-aza-dC) to study the effect of demethylation on
transcription of the Ant4 gene. 5-aza-dC induces DNA demeth-
ylation in cells by depleting DNA methyltransferases through its
covalent and irreversible binding to the enzymes. Notably, the
Ant4 gene was readily derepressed by the addition of 5-aza-dC
to differentiated EBs (day 10) (Fig. 4B). In contrast, expression
of Oct-4 and Nanog, transcription factors selectively expressed
in pluripotent ESCs and primordial germ cells, was not affected.
Ant4 (but not Oct-4 or Nanog) was similarly derepressed by addi-
tion of 5-aza-dC in NIH3T3 cells and an immortalized endoderm
precursor cell line (OBAT, unpublished data).
DNA Methylation of Ant4 Gene During ESC Differentiation Requires Dnmt3DNA methyltransferase 3a and 3b (Dnmt3a and Dnmt3b) has
been proposed to play a primary role in de novo methylation dur-
ing murine embryonic development [31]. To investigate the role of
the Dnmt3 in the establishment of DNA methylation of the Ant4
Figure 2. Ant4 encodes a novel isoform of adenine nucleotide trans-
locase. (A): Deduced amino acid sequence of the mouse Ant4 gene
is aligned with previously identified mouse Ant proteins (Ant1 and
Ant2). (B): Deduced amino acid sequence of the mouse Ant4 gene is
aligned with ANT4 human orthologue.
Rodić, Oka, Hamazaki et al. 1319
promoter during ESC differentiation, we used ESCs homozy-
gously deleted for both the Dnmt3a and Dnmt3b genes. Dnmt3a-
Dnmt3b double-null ESCs (earlier passage cell lines) were sub-
jected to the same differentiation protocol as parental (wild-type)
J1 ESCs. Of interest, Ant4 expression was not suppressed but
rather increased in differentiated double-null cells (Fig. 5A). In
contrast, the pluripotent ESC marker Oct-4 was downregulated,
whereas the primitive ectoderm marker fibroblast growth factor,
Fgf-5, and endoderm marker transthyretin, Ttr, were upregulated
after ESC differentiation. These results indicate that Dnmt3a-
Dnmt3b double-null ESCs proceed with a normal differentiation
pattern. COBRA assays confirmed that there was an absence of
DNA methylation at the Ant4 promoter region (−169 bp) in the
double knockout ESCs (Fig. 5B). These results indicate that func-
tional Dnmt3 is required for repression of the Ant4 gene during
ESC differentiation.
In addition, we examined which member of the Dnmt3
gene family, Dnmt3a or Dnmt3b, played a predominant role in
Ant4 methylation using ESCs deleting either gene. As shown in
Figure 5C, downregulation of Ant4 expression was incomplete
when Dnmt3a-null ESCs or Dnmt3b-null ESCs were differenti-
ated. Furthermore, both Dnmt3a-null ESCs and Dnmt3b-null
ESCs demonstrated virtually no DNA methylation on the CpG
site assayed by COBRA. These data indicated that both Dnmt3a
and Dnmt3b are required for DNA methylation and repression
of Ant4.
Figure 4. Ant4 is repressed during ES cell differentiation. (A): Total
RNA was extracted from undifferentiated ES cells and differentiat-
ing EBs at days 5 and 10. RNA expression levels of Ant4 and β-actin
were evaluated by semiquantitative RT-PCR analysis using various
cycles of DNA amplification (20 to 30 cycles). (B): Ant4 derepres-
sion by 5-aza-dC. EBs were treated with various concentrations
(0–10 μM) of 5-aza-dC for 48 hours from day 8 and harvested at day
10. RNA expression levels of Ant4, Oct-4, Nanog, and β-actin genes
were examined by RT-PCR. Abbreviations: 5-aza-dC, 5-aza-2'-deox-
ycytidine; EB, embryoid body; ESC, embryonic stem cell; RT-PCR,
reverse transcription–polymerase chain reaction.
Figure 3. Ant4 promoter region undergoes DNA methylation dur-
ing ES cell differentiation. Summary of Ant4 methylation levels in
ES cells and EBs (day 10) is illustrated. For each primer pair, up to
14 clones were sequenced after bisulfite treatment of either ES cells
or EBs (day 10) to determine the rate of DNA methylation. Closed
circles represent methylated CpG, whereas open circles represent
unmethylated CpG. The y-axis of the graph represents percent meth-
ylation for each CpG residue. Diagram at the bottom represents rela-
tive position of CpG residues within the 5'-end regulatory region of
Ant4 gene. Nucleotide positions are marked relative to the translation
initiation site (+1). Arrow represents a major transcription start site
(−60 bp), black rectangle represents the first exon, and vertical lines
mark the location of individual CpG residues. Abbreviations: EB,
embryoid body; ESC, embryonic stem cell.
1320 DNA Methylation–Dependent Repression of Ant4
Hypomethylation of the Ant4 Promoter and Ant4 Expression Are Restricted to Testicular Germ CellsTo determine if Ant4 is specifically expressed in ESCs, we ini-
tially used a Basic Local Alignment Search Tool (BLAST) search
against EST databases. We used full-length Ant4 cDNA as a query
sequence and found a total of eight cDNA clones (with scores of
>200 bits). All clones identified were from testis. We then inves-
tigated expression patterns of Ant4 in adult mouse organs using
Northern blot analysis. Ant4 mRNA was found specifically in tes-
tes, at the predicted approximately 1.6-kb transcript size, but was
Figure 5. Dnmt3 is required for Ant4 repression during ESC dif-
ferentiation. (A): Expression of the Ant4 gene in WT ES cells and
Dnmt3a-Dnmt3b double-null ESCs. WT J1 ES cells and Dnmt3a-
Dnmt3b double-null ESCs (Dnmt3a−/−, Dnmt3b−/−) were differ-
entiated as described above. RNA expression was evaluated using
RT-PCR as described above. Oct-4, Ttr, and Fgf-5 are markers for
undifferentiated ESCs, early visceral endoderm, and early ecto-
derm, respectively. (B): Ant4 promoter DNA methylation deter-
mined by COBRA assay. DNA was extracted from WT ESCs and
Dnmt3a-Dnmt3b-null ESCs (Dnmt3a−/−, Dnmt3b−/−) and treated
with bisulfite. The Ant4 promoter region was amplified and sub-
jected to overnight digestion with HhaI restriction enzyme, which
cuts GCGC sites at −169 bp. DNA methylation of the CpG protects
the site from bisulfite conversion; thus, the polymerase chain reac-
tion fragments are digested by HhaI only when the template genomic
DNA is methylated at the site. The digested DNA samples were sepa-
rated in 4.5% polyacrylamide gels and visualized using a SyBr-green
dye. (C): Expression of the Ant4 gene and Ant4 promoter methyla-
tion in Dnmt3a-null ESCs and Dnmt3b-null ESCs. Dnmt3a-null
ESCs (Dnmt3a−/−) and Dnmt3b-null ESCs (Dnmt3b−/−) were differ-
entiated, and Ant4 expression and DNA methylation of the Ant4 pro-
moter were examined as described above. Abbreviations: COBRA,
combined bisulfite restriction analysis; EB, embryoid body; ESC,
embryonic stem cell; RT-PCR, reverse transcription–polymerase
chain reaction; WT, wild-type.
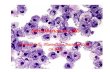
Figure 6. Ant4 expression and promoter DNA methylation in adult
organs. (A): Northern blot analysis of Ant4 mRNA expression in vari-
ous organs from adult mice (8 weeks old). The blot was hybridized to
specific cDNA probes for Ant4, Ant1, Ant2, and β-actin. (B): Immu-
nohistochemical analysis of Ant4 expression in testis, ovary, and liver.
Mouse ovary, testis, and liver were harvested from 3-month-old mice,
and frozen sections were stained with affinity-purified rabbit poly-
clonal anti-Ant4 antibodies and visualized using horse radish peroxi-
dase (brown). Slides were counterstained using Light Green SF Yel-
lowish. (C): Bisulfite analysis of the Ant4 promoter in various organs
from adult mice (8 weeks old). DNA samples were extracted from the
indicated mice organs and subjected to bisulfite conversion. Seven to
eight individual clones were sequenced for each sample. Nucleotide
positions are marked relative to the translation initiation site. Arrow
represents a major transcription start site (−60 bp). (D): Ant4 mRNA
is expressed in purified primordial germ cells. Primordial germ cells
were obtained from E11.5 and 12.5-dpc genital ridges and purified
using anti-SSEA1 magnetic beads. The immunodepleted fraction
(dep) refers to cells not retained on the magnetic column. Individual
samples were subjected to reverse transcription–polymerase chain
reaction analysis.
Rodić, Oka, Hamazaki et al. 1321
undetectable in any other organs examined in Figure 6A. There
was no detectable Ant4 mRNA expression in stomach, small
intestine, skeletal muscle, ovary, thymus, uterus, or placenta (data
not shown). In contrast, the previously identified Ant isoforms,
Ant1 and Ant2, were expressed in many non-germ cell organs at
various levels but were low or undetectable in testes.
We then examined which cells within testis express the Ant4
protein using specific antibodies raised against mouse Ant4 (Fig.
6B). Results obtained from immunohistochemical analyses con-
firmed the presence of Ant4 protein within testicular tissue. The
strong cytoplasmic staining of spermatagonia, spermatocytes,
and spermatids within the seminiferous tubules, coupled with the
lack of signal in interstitial, capillary, and capsular cells, suggests
testicular germ cell–specific expression of Ant4. Of interest, it
seems that mature sperm is absent for Ant4 expression. Although
Northern blot analyses using whole ovaries did not detect any
Ant4 gene expression (described above), immunohistochemical
analyses revealed that Ant4 protein is selectively expressed in the
cytoplasm of oocytes (Fig. 6B, middle panels). In contrast, stain-
ing performed on liver sections was negative. These data indicate
that Ant4 is specifically expressed in developing gametes in testis
and ovary. Similar data were obtained using in situ RNA hybrid-
ization with riboprobes against Ant4 transcript (data not shown).
Further, we determined DNA methylation levels across the
Ant4 promoter region in adult mice organs. Only testis showed
hypomethylation of the Ant4 promoter, whereas other tissues
examined were hypermethylated (Fig. 6C). Additionally, Ant4
was expressed in purified primordial germ cells, obtained from
E11.5 and 12.5-dpc genital ridges when they were purified using
anti-SSEA1 magnetic beads (Fig. 6D). The data indicate that Ant4
is expressed in premeiotic fetal germ cells as well.
DiscussionWe have identified a novel isoform of the adenine nucleotide trans-
locase genes, Ant4, by screening differential DNA methylation
patterns in ESCs and EBs. Selective expression status of Ant4 in
undifferentiated ESCs, primordial germ cells, and developing
gametes in adult testis and ovary indicates that Ant4 is a pluripotent
cell-specific isoform of the adenine nucleotide translocase family
in mouse. This contrasts with the previously identified isoforms
of mouse Ant, Ant1 and Ant2, which are predominately expressed
in somatic cells. BLAST analysis revealed the human orthologue
(ANT4), which shares a high homology in deduced amino acid
sequence with mouse Ant4. Moreover, the mouse Ant4 and human
ANT4 genes have a conserved genomic architecture. While we
were preparing this manuscript, Dolce et al. [32] reported this
human orthologue as AAC4 (ADP/ATP carrier protein 4), which
they identified by screening human genome databases for homol-
ogy to AAC1 (ANT1). Their study demonstrated that AAC4
actively exchanges ADP for ATP by an electrogenic antiport mech-
anism. Although AAC4 expression was highest in testis among
human tissues, as we showed here with mouse Ant4, their real-time
PCR analysis indicated that AAC4 was expressed in other somatic
organs, including liver and brain. This is in contrast to the pattern
of Ant4 expression in mice demonstrated by Northern blotting
and immunohistochemical analyses in the present study, which is
highly restricted to developing gametes among adult tissues.
Ant exchanges mitochondrial ATP for cytosolic ADP and
therefore plays a pivotal role in cellular metabolism in eukaryotic
cells. It is intriguing that germ cell–specific isoforms also exist
in other proteins involved in mitochondrial energy metabolism.
Developing spermatocytes are known to express testis-specific
isoforms of pyruvate dehydrogenase complex E1α subunit,
Pdha-2 [33], testis-specific cytochrome c, Cyt cT, [34], and sub-
unit VIb of cytochrome c oxidase [35]. Interestingly, Cyt cT–null
mice exhibit early testicular atrophy and apoptosis [34]. Ant
has been implicated in the mitochondrial permeability transi-
tion, which is a common feature of apoptosis. It is possible that
this germ cell–specific isoform of Ant, probably interacting with
other germ cell–specific mitochondrial membrane proteins, could
also be critical for germ cell development and maintenance. Fur-
ther, a recent study indicates that an antiapoptotic protein, Bcl2,
supports ESC survival and maintenance in harsh culture condi-
tions such as serum deprivation [36], implying that antiapoptotic
mechanisms of the mitochondrial membrane play a role in ESC
self-renewal. We are now generating Ant4-null ESCs and knock-
out mice, which will elucidate the in vivo function of the protein
in spermatogenesis and ESC maintenance.
The present data indicate that de novo DNA methylation,
mediated by Dnmt3a and Dnmt3b, plays a pivotal role in gene
repression of Ant4 during ESC differentiation. In adult organs, as
well as during in vitro ESC differentiation, Ant4 expression levels
inversely correlated with the DNA methylation status of the gene.
Further, Ant4 was readily derepressed by addition of the demethyl-
ating agent 5-aza-dC. In contrast, Oct-4 and Nanog, genes specifi-
cally expressed in pluripotent stem cells and primordial germ cells,
were not derepressed by 5-aza-dC. This implies that transcription
factors involved in Ant4 gene expression are present and active in
differentiated EBs and that DNA methylation may play a primary
role in the suppression of the Ant4 gene in somatic cells. We con-
firmed derepression of Ant4, but not Oct-4 or Nanog, by addition of
5-aza-dC into two other somatic cell lines. Further, this hypothesis
was supported by the fact that functional deletion of Dnmt3a and
Dnmt3b led to failure of gene suppression of Ant4, but not Oct-4,
during ESC differentiation. Taken together, these data suggest that
DNA methylation is required for the transcriptional repression of
the Ant4 gene. The present data also indicate that both members
of the Dnmt3 gene family are required for complete repression of
the Ant4 gene. At the particular CpG site that we investigated by
COBRA, DNA methylation was hardly detected in single-knock-
out EBs within 10 days of differentiation. Dnmt3a and Dnmt3b
may induce DNA methylation cooperatively at some CpG sites.
1322 DNA Methylation–Dependent Repression of Ant4
Undifferentiated ESCs are known to be transcriptionally
permissive for expression of some germ line cell–specific genes
[37]. This might be due to a conserved mode of transcriptional
regulation between ESCs and germ cells or simply reflect the
fact that ESCs are derived from largely unmethylated preim-
plantation embryos (i.e., blastocyst). Recent reports demon-
strated that DNA methylation is involved in the transcriptional
regulation of several germ cell–specific genes, including pgk2
[38], pdha-2 [39], the MAGE gene family [40], and Tact1/Actl7b
[41]. Because Ant4 is expressed exclusively in the testes of the
adult mouse, the regulation of Ant4 transcription may be closely
related to that of other germ cell–specific genes. Of interest, the
promoter region of Ant4 contains the Coup-Tf binding element
at −586 bp, which has been implicated in selective silencing of
the c-Mos proto-oncogene in somatic cells [42, 43]. In addition,
the Ant4 promoter has multiple elements that potentially bind to
Sox/Sry factors, which have also been implicated in early cell
fate specification in development and germ cell–specific tran-
scription [44, 45]. By elucidating the role of DNA methylation
in gene regulation of pluripotent cells, we may uncover common
gene regulatory pathways underlying the transition between plu-
ripotent stem/germ cells and somatic cells.
AcknowledgmentsThe authors thank Drs. Thomas Yang, Paul Oh, Jorg Bungert,
Keith Robertson, and Michael Rutenberg for helpful discussions
and critical reading of the manuscript. This work was supported
in part by National Institutes of Health grants DK59699 and
RR17001 (to N.T.).
DisclosuresN.T. owns stock in RegenMed, Inc.
References
1 Robertson AK, Geiman TM, Sankpal UT et al. Effects of chromatin
structure on the enzymatic and DNA binding functions of DNA methyl-
transferases DNMT1 and Dnmt3a in vitro. Biochem Biophys Res Com-
mun 2004;322:110–118.
2 Li E, Bestor TH, Jaenisch R. Targeted mutation of the DNA methyltrans-
ferase gene results in embryonic lethality. Cell 1992;69:915–926.
3 Panning B, Jaenisch R. RNA and the epigenetic regulation of X chromo-
some inactivation. Cell 1998;93:305–308.
4 Li E, Beard C, Jaenisch R. Role for DNA methylation in genomic imprint-
ing. Nature 1993;366:362–365.
5 Walsh CP, Chaillet JR, Bestor TH. Transcription of IAP endogenous ret-
roviruses is constrained by cytosine methylation. Nat Genet 1998;20:116–
117.
6 Jones PA, Laird PW. Cancer epigenetics comes of age. Nat Genet
1999;21:163–167.
7 Baylin SB, Esteller M, Rountree MR et al. Aberrant patterns of DNA
methylation, chromatin formation and gene expression in cancer. Hum
Mol Genet 2001;10:687–692.
8 Bird AP, Wolffe AP. Methylation-induced repression: belts, braces, and
chromatin. Cell 1999;99:451–454.
9 Takizawa T, Nakashima K, Namihira M et al. DNA methylation is a
critical cell-intrinsic determinant of astrocyte differentiation in the fetal
brain. Dev Cell 2001;1:749–758.
10 Nan X, Ng HH, Johnson CA et al. Transcriptional repression by the
methyl-CpG-binding protein MeCP2 involves a histone deacetylase com-
plex. Nature 1998;393:386–389.
11 Grewal SI, Moazed D. Heterochromatin and epigenetic control of gene
expression. Science 2003;301:798–802.
12 Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells
from mouse embryos. Nature 1981;292:154–156.
13 Smith AG, Heath JK, Donaldson DD et al. Inhibition of pluripotential
embryonic stem cell differentiation by purified polypeptides. Nature
1988;336:688–690.
14 Chambers I, Colby D, Robertson M et al. Functional expression cloning
of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell
2003;113:643–655.
15 Mitsui K, Tokuzawa Y, Itoh H et al. The homeoprotein Nanog is required
for maintenance of pluripotency in mouse epiblast and ESCs. Cell
2003;113:631–642.
16 Niwa H, Miyazaki J, Smith AG. Quantitative expression of Oct-3/4
defines differentiation, dedifferentiation or self-renewal of ESCs. Nat
Genet 2000;24:372–376.
17 Okano M, Xie S, Li E. Cloning and characterization of a family of
novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet
1998;19:219–220.
18 Lee JH, Hart SRL, Skalnik DG. Histone deacetylase activity is required
for embryonic stem cell differentiation. Genesis 2004;38:32–38.
19 Huntriss J, Hinkins M, Oliver B et al. Expression of mRNAs for DNA
methyltransferases and methyl-CpG-binding proteins in the human
female germ line, preimplantation embryos, and embryonic stem cells.
Mol Reprod Dev 2004;67:323–336.
20 Kremenskoy M, Kremenska Y, Ohgane J et al. Genome-wide analysis of
DNA methylation status of CpG islands in embryoid bodies, teratomas,
and fetuses. Biochem Biophys Res Commun 2003;311:884–890.
21 Shiota K, Kogo Y, Ohgane J et al. Epigenetic marks by DNA methylation
specific to stem, germ and somatic cells in mice. Genes Cells 2002;7:961–
969.
22 Chen T, Ueda Y, Dodge JE et al. Establishment and maintenance of
genomic methylation patterns in mouse embryonic stem cells by Dnmt3a
and Dnmt3b. Mol Cell Biol 2003;23:5594–5605.
23 Hamazaki T, Iiboshi Y, Oka M et al. Hepatic maturation in differentiating
embryonic stem cells in vitro. FEBS Lett 2001;497:15–19.
24 Huang TH, Laux DE, Hamlin BC et al. Identification of DNA methylation
markers for human breast carcinomas using the methylation-sensitive
restriction fingerprinting technique. Cancer Res 1997;57:1030–1034.
25 Pesce M, De Felici M. Purification of mouse primordial germ cells by
MiniMACS magnetic separation system. Dev Biol 1995;170:722–725.
26 Belzacq AS, Vieira HL, Kroemer G et al. The adenine nucleotide translo-
cator in apoptosis. Biochimie 2002;84:167–176.
Rodić, Oka, Hamazaki et al. 1323
27 Slim R, Levilliers J, Ludecke HJ et al. A human pseudoautosomal gene
encodes the ANT3 ADP/ATP translocase and escapes X-inactivation.
Genomics 1993;16:26–33.
28 Mozo T, Fischer K, Flugge UI et al. The N-terminal extension of the ADP/
ATP translocator is not involved in targeting to plant mitochondria in vivo.
Plant J 1995;7:1015–1020.
29 Chen ST, Chang CD, Huebner K et al. A human ADP/ATP translocase
gene has seven pseudogenes and localizes to chromosome X. Somat Cell
Mol Genet 1990;16:143–149.
30 Xiong Z, Laird PW. COBRA: a sensitive and quantitative DNA methyla-
tion assay. Nucleic Acids Res 1997;25:2532–2534.
31 Okano M, Bell DW, Haber DA et al. DNA methyltransferases Dnmt3a and
Dnmt3b are essential for de novo methylation and mammalian develop-
ment. Cell 1999;99:247–257.
32 Dolce V, Scarcia P, Iacopetta D et al. A fourth ADP/ATP carrier isoform in
man: identification, bacterial expression, functional characterization and
tissue distribution. FEBS Lett 2005;579:633–637.
33 Brown RM, Dahl HH, Brown GK. Pyruvate dehydrogenase E1 alpha sub-
unit genes in the mouse: mapping and comparison with human homologs.
Somat Cell Mol Genet 1990;16:487–492.
34 Narisawa S, Hecht NB, Goldberg E et al. Testis-specific cytochrome c-
null mice produce functional sperm but undergo early testicular atrophy.
Mol Cell Biol 2002;22:5554–5562.
35 Huttemann M, Jaradat S, Grossman LI. Cytochrome c oxidase of mam-
mals contains a testes-specific isoform of subunit VIb–the counterpart to
testes-specific cytochrome c? Mol Reprod Dev 2003;66:8–16.
36 Yamane T, Dylla SJ, Muijtjens M et al. Enforced Bcl-2 expression over
rides serum and feeder cell requirements for mouse embryonic stem cell
self-renewal. Proc Natl Acad Sci U S A 2005;102:3312–3317.
37 Geijsen N, Horoschak M, Kim K et al. Derivation of embryonic germ cells
and male gametes from embryonic stem cells. Nature 2004;427:148–154.
38 Zhang LP, Stroud JC, Walter CA et al. A gene-specific promoter in trans-
genic mice directs testis-specific demethylation prior to transcriptional
activation In vivo. Biol Reprod 1998;59:284–292.
39 Iannello RC, Gould JA, Young JC et al. Methylation-dependent silenc-
ing of the testis-specific Pdha-2 basal promoter occurs through selective
targeting of an activating transcription factor/cAMP-responsive element-
binding site. J Biol Chem 2000;275:19603–19608.
40 De Smet C, Lurquin C, Lethe B et al. DNA methylation is the primary
silencing mechanism for a set of germ line- and tumor-specific genes with
a CpG-rich promoter. Mol Cell Biol 1999;19:7327–7335.
41 Hisano M, Ohta H, Nishimune Y et al. Methylation of CpG dinucleotides
in the open reading frame of a testicular germ cell-specific intronless gene,
Tact1/Actl7b, represses its expression in somatic cells. Nucleic Acids Res
2003;31:4797–4804.
42 Xu W, Cooper GM. Identification of a candidate c-mos repressor
that restricts transcription of germ cell-specific genes. Mol Cell Biol
1995;15:5369–5375.
43 Lin HB, Jurk M, Gulick T et al. Identification of COUP-TF as a tran-
scriptional repressor of the c-mos proto-oncogene. J Biol Chem
1999;274:36796–36800.
44 Tchenio T, Casella JF, Heidmann T. Members of the SRY family regulate
the human LINE retrotransposons. Nucleic Acids Res 2000;28:411–415.
45 Clarkson MJ, Harley VR. Sex with two SOX on: SRY and SOX9 in testis
development. Trends Endocrinol Metab 2002;13:106–111.
Related Documents