Control of Eukaryotic Genes (Ch. 19)

Control of Eukaryotic Genes (Ch. 19)
Jan 02, 2016
Control of Eukaryotic Genes (Ch. 19). The BIG Questions…. How are genes turned on & off in eukaryotes? How do cells with the same genes differentiate to perform completely different, specialized functions?. Evolution of gene regulation. Prokaryotes single-celled - PowerPoint PPT Presentation
Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

Control of Eukaryotic Genes
(Ch. 19)

The BIG Questions…
• How are genes turned on & off in eukaryotes?
• How do cells with the same genes differentiate to perform completely different, specialized functions?

Evolution of gene regulation
• Prokaryotes– single-celled– evolved to grow & divide rapidly– must respond quickly to changes in
external environment• exploit transient resources
• Gene regulation- Operons– turn genes on & off rapidly
• flexibility & reversibility– adjust levels of enzymes
for synthesis & digestion

Evolution of gene regulation
• Eukaryotes– multicellular– evolved to maintain homeostasis– regulate body as a whole
• specialization–turn on & off large number of
genes• must coordinate the body as a whole
rather than serve the needs of individual cells

Points of control
• The control of gene expression can occur at any step in the pathway from gene to functional protein

from DNA double helix to condensed chromosome
1. DNA packing

Nucleosomes • “Beads on a string”
– 1st level of DNA packing– histone proteins
• 8 protein molecules• positively charged amino acids • bind tightly to negatively charged DNA
8 histone 8 histone moleculesmolecules

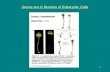
DNA packing as gene control• Degree of packing of DNA regulates
transcription– tightly wrapped around histones
• no transcription• genes turned off
heterochromatindarker DNA (H) = tightly packed
euchromatinlighter DNA (E) = loosely packed

DNA methylation
• Methylation of DNA blocks transcription factors – attachment of methyl groups (–CH3) to cytosine
– nearly permanent inactivation of genes• ex. inactivated mammalian X chromosome =
Barr body

Histone acetylation Acetylation of histones
unwinds DNA Loose histone wrapping
enables transcription genes turned on
attachment of acetyl groups (–COCH3) to histones
conformational change in histone proteins
transcription factors have easier access to genes
Chromatin changes
Transcription
RNA processing
mRNA degradation
Translation
Protein processingand degradation
Histonetails
DNAdouble helix Amino acids
availablefor chemicalmodification
(a) Histone tails protrude outward from a nucleosome
(b) Acetylation of histone tails promotes loose chromatinstructure that permits transcription
Unacetylated histones Acetylated histones

2. Transcription initiation• Control regions on DNA
– promoter• nearby control sequence on DNA• binding of RNA polymerase & transcription
factors– enhancer
• distant control sequences on DNA
• binding of activator proteins

Model for Enhancer action
• Enhancer DNA sequences – distant control sequences
• Activator proteins – bind to enhancer sequence &
stimulates transcription
• Silencer proteins – bind to enhancer sequence &
block gene transcription

Transcription complex
Enhancer
ActivatorActivator
Activator
Coactivator
RNA polymerase II
A
B F E
HTFIID
Core promoterand initiation complex
Activator Proteins• regulatory proteins bind to DNA at
distant enhancer sites• increase the rate of transcription
Coding region
T A T A
Enhancer Sitesregulatory sites on DNA distant from gene
Initiation Complex at Promoter Site binding site of RNA polymerase

14
Many control elements for different genes are the same
It is the combination of control elements
that provides specificity

3. Post-transcriptional control
• Alternative RNA splicing– variable processing of exons creates a
family of proteins

4. Regulation of mRNA degradation
• Life span of mRNA determines amount of protein synthesis– mRNA can last from hours to weeks

RNA interference NEW!
Chromatin changes
Transcription
RNA processing
mRNAdegradation
Translation
Protein processingand degradation
Degradation of mRNAOR
Blockage of translation
Target mRNA
miRNA
Proteincomplex
Dicer
Hydrogenbond
The micro-RNA (miRNA)precursor foldsback on itself,held togetherby hydrogenbonds.
1
2 An enzymecalled Dicer movesalong the double-stranded RNA, cutting it intoshorter segments.
2
One strand ofeach short double-stranded RNA isdegraded; the otherstrand (miRNA) thenassociates with acomplex of proteins.
3 The boundmiRNA can base-pairwith any targetmRNA that containsthe complementarysequence.
4 The miRNA-proteincomplex prevents geneexpression either bydegrading the targetmRNA or by blockingits translation.
5

19
miRNA’s• Scientists discovery video• Animation of miRNA function

RNA interference 1990s | 2006
Andrew FireAndrew FireStanfordStanford
Craig MelloCraig MelloU MassU Mass
“for their discovery of RNA interference —gene silencing by
double-stranded RNA”

5. Control of translation
• Block initiation of translation stage – regulatory proteins attach to 5' end of mRNA
• prevent attachment of ribosomal subunits & initiator tRNA
• block translation of mRNA to protein

6-7. Protein processing & degradation
• Protein processing– folding, cleaving, adding sugar groups,
targeting for transport• Protein degradation
– ubiquitin tagging– proteasome degradation

Ubiquitin
• “Death tag”– mark unwanted proteins with a label – 76 amino acid polypeptide, ubiquitin– labeled proteins are broken down rapidly
in "waste disposers"• proteasomes
1980s | 2004
Aaron CiechanoverAaron CiechanoverIsraelIsrael
Avram HershkoAvram HershkoIsraelIsrael
Irwin RoseIrwin RoseUC RiversideUC Riverside

Proteasome • Protein-degrading “machine”
– cell’s waste disposer– breaks down any proteins
into 7-9 amino acid fragments• cellular recycling

initiation of transcription
1
mRNA splicing
2
mRNA protection3
initiation of translation
6
mRNAprocessing
5
1 & 2. transcription - DNA packing - transcription factors
3 & 4. post-transcription - mRNA processing
- splicing- 5’ cap & poly-A tail- breakdown by siRNA
5. translation - block start of translation
6 & 7. post-translation - protein processing - protein degradation
7protein processing & degradation
4
4
Gene Regulation

26
Concept Check
1. Do miRNA’s increase or decrease gene expression?
2. Does histone de-acetylation increase or decrease gene expression?

Genome Sizes and Estimated Numbers of Genes*

Types of DNA sequences in the human genome
Exons (regions of genes codingfor protein, rRNA, tRNA) (1.5%)
RepetitiveDNA thatincludes
transposableelements
and relatedsequences
(44%)
Introns andregulatorysequences
(24%)
UniquenoncodingDNA (15%)
RepetitiveDNA
unrelated totransposable
elements(about 15%)
Alu elements(10%)
Simple sequenceDNA (3%)
Large-segmentduplications (5–6%)

Just how little of out Genome Codes for Genes?
QuickTime™ and aH.264 decompressor
are needed to see this picture.

30
Cancer results from genetic changes in cell
cycle control

31
Proto-oncogene --> oncogene

32

33
Multistep Model of cancer

34
Genome Evolution
• How did our genome get so big?

35

36

37
• The similarity in the amino acid sequences of the various globin proteins– Supports this model of gene duplication and
mutation
Table 19.1

38
Evolution of Genes with Novel Functions
• The copies of some duplicated genes– Have diverged so much during
evolutionary time that the functions of their encoded proteins are now substantially different

39

40
Concept check
1. How much of the human genome consists of exons?
2. How can exon shuffling lead to the evolution of a new gene

Turn yourQuestion Genes on!

42
A larger portion of the DNA in a eukaryotic cell is transcribed than would be predicted by the proteins made by the cell. What is being transcribed and what is its function?
1. Multiple enhancer regions are being transcribed to amplify the transcription of protein-coding genes.
2. These transcriptions are of non-coding “junk” DNA that is a remnant of mutated protein coding segments. The transcripts are degraded by nuclear enzymes.
3. The additional DNA that is transcribed is introns that are excised fro the primary transcript in the produciton of mRNA.
4. Many RNA coding genes are transcribed. Precursor RNAs fold into hairpin structures, which are cut and processed into miRNAs that regulate translation of mRNAs.

43
How is the coordinated transcription of genes involved in the same pathway regulated?
1. The genes are transcribed in one transcription unit, although each gene has its own promoter.
2. The genes are located in the same region of the chromosome, and enzymes de-acetylate the entire region so that transcription may begin.
3. A steroid hormone selectively binds to the promoters for all the genes.
4. The genes have the same combination of control elements in the enhancer that bind with the particular activators present in the cell.

44
Histones are
1. Small positively charged proteins that bind tightly to DNA
2. Small bodies in the nucleus involved in rRNA synthesis
3. Basic units of DNA packing consisting of DNA wound around a protein core.
4. DNA bending proteins that facilitate formation of transcription initiation complexes.
5. Proteins responsible for producing repeating sequences at telomeres.

45
Heterochromatin
1. Has a higher degree of packing than does euchromatin.
2. Is visible with a light microscope during interphase.
3. Is not actively involved in transcription
4. Makes up metaphase chromosomes
5. Is all of the above.

46
What is the main reason that prokaryotic genes average 1,000 nucleotride base pairs whereas human genes average about 27.000 base pairs?
1. Prokaryotes have smaller, bue many more individual genes.
2. Prokaryotes are more ancient organisms, longer genes arose later in evolution.
3. Prokaryotes are much simpler organism humans have many types of differentiated cells.
4. Prokaryotic genes do not have introns, human genes have many
5. Human proteins are much larger and more complex than prokaryotic proteins.

47
A eukaryotic gene typically has all of the following associated with it except
1. A promoter
2. An operator
3. Enhancers
4. Introns and exons
5. Control elements.
Related Documents