A phylogeny and revised classification of Squamata, including 4161 species of lizards and snakes Pyron et al. Pyron et al. BMC Evolutionary Biology 2013, 13:93 http://www.biomedcentral.com/1471-2148/13/93

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

A phylogeny and revised classification ofSquamata, including 4161 species of lizardsand snakesPyron et al.
Pyron et al. BMC Evolutionary Biology 2013, 13:93http://www.biomedcentral.com/1471-2148/13/93

Pyron et al. BMC Evolutionary Biology 2013, 13:93http://www.biomedcentral.com/1471-2148/13/93
RESEARCH ARTICLE Open Access
A phylogeny and revised classification ofSquamata, including 4161 species of lizardsand snakesR Alexander Pyron1*, Frank T Burbrink2,3 and John J Wiens4
Abstract
Background: The extant squamates (>9400 known species of lizards and snakes) are one of the most diverse andconspicuous radiations of terrestrial vertebrates, but no studies have attempted to reconstruct a phylogeny for thegroup with large-scale taxon sampling. Such an estimate is invaluable for comparative evolutionary studies, and toaddress their classification. Here, we present the first large-scale phylogenetic estimate for Squamata.
Results: The estimated phylogeny contains 4161 species, representing all currently recognized families andsubfamilies. The analysis is based on up to 12896 base pairs of sequence data per species (average = 2497 bp) from12 genes, including seven nuclear loci (BDNF, c-mos, NT3, PDC, R35, RAG-1, and RAG-2), and five mitochondrialgenes (12S, 16S, cytochrome b, ND2, and ND4). The tree provides important confirmation for recent estimates ofhigher-level squamate phylogeny based on molecular data (but with more limited taxon sampling), estimates thatare very different from previous morphology-based hypotheses. The tree also includes many relationships that differfrom previous molecular estimates and many that differ from traditional taxonomy.
Conclusions: We present a new large-scale phylogeny of squamate reptiles that should be a valuable resource forfuture comparative studies. We also present a revised classification of squamates at the family and subfamily levelto bring the taxonomy more in line with the new phylogenetic hypothesis. This classification includes new,resurrected, and modified subfamilies within gymnophthalmid and scincid lizards, and boid, colubrid, andlamprophiid snakes.
Keywords: Amphisbaenia, Lacertilia, Likelihood support measures, Missing data, Serpentes, Squamata,Phylogenetics, Reptilia, Supermatrices, Systematics
BackgroundSquamate reptiles (lizards, snakes, and amphisbaenians["worm lizards"]) are among the most diverse radiationsof terrestrial vertebrates. Squamata includes more than9400 species as of December 2012 [1]. The rate of newspecies descriptions shows no signs of slowing, with arecord 168 new species described in 2012 [1], greaterthan the highest yearly rates of the 18th and 19th cen-turies (e.g. 1758, 118 species; 1854, 144 species [1]).Squamates are presently found on every continent ex-cept Antarctica, and in the Indian and Pacific Oceans,and span many diverse ecologies and body forms,
* Correspondence: [email protected] of Biological Sciences, The George Washington University,2023 G St. NW, Washington, DC 20052, USAFull list of author information is available at the end of the article
© 2013 Pyron et al.; licensee BioMed Central LCommons Attribution License (http://creativecreproduction in any medium, provided the or
from limbless burrowers to arboreal gliders (summa-rized in [2-4]).Squamates are key study organisms in numerous fields,
from evolution, ecology, and behavior [3] to medicine [5,6]and applied physics [7]. They have also been the focus ofmany pioneering studies using phylogenies to address ques-tions about trait evolution (e.g. [8,9]). Phylogenies are nowrecognized as being integral to all comparative studies ofsquamate biology (e.g. [10,11]). However, hypotheses aboutsquamate phylogeny have changed radically in recent years[12], especially when comparing trees generated from mor-phological [13-15] and molecular data [16-20]. Further-more, despite extensive work on squamate phylogeny at alltaxonomic levels, a large-scale phylogeny (i.e. includingthousands of species and multiple genes) has never beenattempted using morphological or molecular data.
td. This is an Open Access article distributed under the terms of the Creativeommons.org/licenses/by/2.0), which permits unrestricted use, distribution, andiginal work is properly cited.

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 2 of 53http://www.biomedcentral.com/1471-2148/13/93
Squamate phylogenetics has changed radically in thelast 10 years, revealing major conflicts between the re-sults of morphological and molecular analyses [12]. Earlyestimates of squamate phylogeny [21] and recent studiesbased on morphological data [13-15,22] consistentlysupported a basal division between Iguania (includingchameleons, agamids, and iguanids, sensu lato), andScleroglossa, which comprises all other squamates (in-cluding skinks, geckos, snakes, and amphisbaenians).Within Scleroglossa, many phylogenetic analyses of mor-phological data have also supported a clade containinglimb-reduced taxa, including various combinations ofsnakes, dibamids, amphisbaenians, and (in some ana-lyses) limb-reduced skinks and anguids [13-15,19,22],though some of these authors also acknowledged thatthis clade was likely erroneous.In contrast, recent molecular analyses have estimated
very different relationships. Novel arrangements includeplacement of dibamids and gekkotans near the root ofthe squamate tree, a sister-group relationship betweenamphisbaenians and lacertids, and a clade (Toxicofera)uniting Iguania with snakes and anguimorphs withinScleroglossa [16-20,23,24]. These molecular results (andthe results of combined morphological and molecularanalyses) suggest that some estimates of squamate phy-logeny based on morphology may have been misled, es-pecially by convergence associated with adaptations toburrowing [19]. However, there have also been disagree-ments among molecular studies, such as placement ofdibamids relative to gekkotans and other squamates, andrelationships among snakes, iguanians, and anguimorphs(e.g. [17,20]).Analyses of higher-level squamate relationships based
on molecular data have so far included relatively few(less than 200) species, and none have included repre-sentatives from all described families and subfamilies[17-20,23,24]. This limited taxon sampling makesexisting molecular phylogenies difficult to use for broad-scale comparative studies, with some exceptions basedon supertrees [10,11]. In addition, limited taxon sam-pling is potentially a serious issue for phylogenetic ac-curacy [25-28]. Thus, an analysis with extensive taxonsampling is critically important to test hypotheses basedon molecular datasets with more limited sampling, andto provide a framework for comparative analyses.Despite the lack of a large-scale phylogeny across
squamates, recent molecular studies have producedphylogenetic estimates for many of the major groups ofsquamates, including iguanian lizards [29-34], higher-level snake groups [35-37], typhlopoid snakes [38,39],colubroid snakes [40-46], booid snakes [47,48], scincidlizards [49-52], gekkotan lizards [53-60], teiioid lizards[61-64], lacertid lizards [65-69], and amphisbaenians[70,71]. These studies have done an outstanding job of
clarifying the phylogeny and taxonomy of these groups,but many were limited in some ways by the number ofcharacters and taxa that they sampled (and which wereavailable at the time for sequencing).Here, we present a phylogenetic estimate for Squamata
based on combining much of the existing sequence data forsquamate reptiles, using the increasingly well-establishedsupermatrix approach [41,72-77]. We present a newphylogenetic estimate including 4161 squamate species.The dataset includes up to 12896 bp per species from12 genes (7 nuclear, 5 mitochondrial). We include speciesfrom all currently described families and subfamilies.In terms of species sampled, this is 5 times larger than anyprevious phylogeny for any one squamate group [30,41],3 times larger than the largest supertree estimate [11],and 25 times larger than the largest molecular study ofhigher-level squamate relationships [20]. While we didnot sequence any new taxa specifically for this project,much of the data in the combined matrix were gene-rated in our labs or from our previous collaborative pro-jects [16,19,20,34,36,37,41,44,78-82], including thousandsof gene sequences from hundreds of species (>550 spe-cies; ~13% of the total).The supermatrix approach can provide a relatively
comprehensive phylogeny, and uncover novel relation-ships not seen in any of the separate analyses in whichthe data were generated. Such novel relationships can berevealed via three primary mechanisms. First, differentstudies may have each sampled different species from agiven group for the same genes, and combining thesedata may reveal novel relationships not apparent in theseparate analyses. Second, different studies may haveused different genetic markers for the same taxa, andcombining these markers can dramatically increase char-acter sampling, potentially revealing new relationshipsand providing stronger support for previous hypotheses.Third, even for clades that were previously studied usingcomplete taxon sampling and multiple loci, novel rela-tionships may be revealed by including these lineageswith other related groups in a large-scale phylogeny.The estimated tree and branch-lengths should be use-
ful for comparative studies of squamate biology. How-ever, this phylogeny is based on a supermatrix withextensive missing data (mean = 81% per species). Someauthors have suggested that matrices with missing cellsmay yield misleading estimates of topology, support, andbranch lengths [83]. Nevertheless, most empirical andsimulation studies have not found this to be the case, atleast for topology and support [41,73,84,85]. Thoughfewer studies have examined the effects of missing dataon branch lengths [44,86,87], these also suggest thatmissing data do not strongly impact estimates. Here, wetest whether branch lengths for terminal taxa are relatedto their completeness.

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 3 of 53http://www.biomedcentral.com/1471-2148/13/93
In general, our results corroborate those of many recentmolecular studies with regard to higher-level relationships,species-level relationships, and the monophyly, compo-sition, and relationships of most families, subfamilies, andgenera. However, our results differ from previous estimatesfor some groups, and reveal (or corroborate) numerousproblems in the existing classification of squamates. Wetherefore provide a conservative, updated classification ofextant squamates at the family and subfamily level basedon the new phylogeny, while highlighting problematic ta-xonomy at the genus level, without making changes. Thegeneric composition of all families and subfamilies underour revised taxonomy are provided in Appendix I.We note dozens of problems in the genus-level tax-
onomy suggested by our tree, but we acknowledge in ad-vance that we do not provide a comprehensive review ofthe previous literature dealing with all these taxonomic is-sues (this would require a monographic treatment). Simi-larly, we do not attempt to fix these genus-level problemshere, as most will require more extensive taxon (and po-tentially character) sampling to adequately resolve.Throughout the paper, we address only extant squa-
mates. Squamata also includes numerous extinct spe-cies classified in both extant and extinct families,subfamilies, and genera. Relationships and classificationof extinct squamates based on morphological datafrom fossils have been addressed by numerous authors(e.g. [14,15,19,22,88-93]). A classification based only onliving taxa may create some problems for classifyingfossil taxa, but these can be addressed in future studiesthat integrate molecular and fossil data [19,86].
ResultsSupermatrix phylogenyWe generated the final tree (lnL = −2609551.07) usingMaximum Likelihood (ML) in RAxMLv7.2.8. Supportwas assessed using the non-parametric Shimodaira-Hasegawa-Like (SHL) implementation of the approximatelikelihood-ratio test (aLRT; see [94]). The tree and datamatrix are available in NEXUS format in DataDryad re-pository 10.5061/dryad.82h0m and as Additional file 1:Data File S1. A skeletal representation of the tree (exclud-ing several species which are incertae sedis) is shown inFigure 1. The full species-level phylogeny (minus theoutgroup Sphenodon) is shown in Figures 2, 3, 4, 5, 6, 7, 8,9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25,26, 27, 28. The analysis yields a generally well-supportedphylogenetic estimate for squamates (i.e. 70% of nodes haveSHL values >85, indicating they are strongly supported).There is no relationship between proportional complete-ness (bp of non-missing data in species / 12896 bp ofcomplete data) and branch length (r = −0.29, P = 0.14) forterminal taxa, strongly suggesting that the estimatedbranch lengths are not consistently biased by missing data.
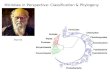
Higher-level relationshipsOur tree (Figure 1) is broadly congruent with most previ-ous molecular studies of higher-level squamate phylogenyusing both nuclear data and combined nuclear and mito-chondrial data (e.g. [16-20]), providing important confirm-ation of previous molecular studies based on more limitedtaxon sampling. Specifically we support (Figure 1): (i) theplacement of dibamids and gekkotans near the base of thetree (Figure 1A); (ii) a sister-group relationship betweenScincoidea (scincids, cordylids, gerrhosaurids, and xantu-siids; Figure 1B) and a clade (Episquamata; Figure 1C)containing the rest of the squamates excluding dibamidsand gekkotans; (iii) Lacertoidea (lacertids, amphisbaenians,teiids, and gymnophthalmids; Figure 1D), and (iv) a clade(Toxicofera; Figure 1E) containing anguimorphs (Figure 1F),iguanians (Figure 1G), and snakes (Figure 1H) as the sistertaxon to Lacertoidea.These relationships are strongly supported in general
(Figure 1), but differ sharply from most trees based onmorphological data [13-15,19,22,95]. Nevertheless, manyclades found in previous morphological taxonomies andphylogenies are also present in this tree in someform, including Amphisbaenia, Anguimorpha, Gekkota,Iguania, Lacertoidea (but including amphisbaenians),Scincoidea, Serpentes, and many families and subfamilies.In contrast, the relationships among these groups differstrongly between molecular analyses [17-20] and morpho-logical analyses [14,15]. Our results demonstrate that thisincongruence is not explained by limited taxon sampling inthe molecular data sets. In fact, our species-level samplingis far more extensive than in any morphological analyses(e.g. [14,15]), by an order of magnitude.We find that the basal squamate relationships are strong-
ly supported in our tree. The family Dibamidae is the sistergroup to all other squamates, and Gekkota is the sistergroup to all squamates excluding Dibamidae (Figure 1), asin some previous studies (e.g. [16,18]). Other recent mo-lecular analyses have also placed Dibamidae near the squa-mate root, but differed in placing it as either the sistertaxon to all squamates excluding Gekkota [17], or thesister- group of Gekkota [19,20]. Our results also corrobor-ate that the New World genus Anelytropsis is nested withinthe Old World genus Dibamus [96], but the associatedbranches are weakly supported (Figure 2).
GekkotaWithin Gekkota, we corroborate both earlier morpho-logical [97] and recent molecular estimates [55,56,59,98]in supporting a clade containing the Australian radiation of"diplodactylid" geckos (Carphodactylidae and Diplodac-tylidae) and the snakelike pygopodids (Figures 1, 2).As in previous studies [55], Carphodactylidae is theweakly supported sister group to Pygopodidae, andthis clade is the sister group of Diplodactylidae

0.2 subst./site
Scincidae
Xantusiidae
Gerrhosauridae
Teiidae
Gymnophthalmidae
Lacertidae
Anguidae
Chamaeleonidae
Agamidae
Leiosauridae
Boidae
Viperidae
Colubridae
Lamprophiidae
A
B
C
D
E
F
G
A) GekkotaB) ScincoideaC) EpisquamataD) LacertoideaE) ToxicoferaF) AnguimorphaG) IguaniaH) Serpentes
H
Dipsadinae
Gekkonidae
Cercosaurinae
Acontiinae
Lacertinae
Hydrosaurinae
Xenodermatidae
Anniellidae
Crotaphytidae
Hoplocercidae
Sphenodontidae
Leiolepidinae
Polychrotidae
Pseudoxyrhophiinae
Loxocemidae
Dibamidae
Prosymninae
Lygosominae
Natricinae
Xantusiinae
Tropiduridae
Corytophanidae
Ecpleopinae
Anomochilidae
Amphisbaenidae
Leiosaurinae
Xenophiidae
Scincinae
Draconinae
Bipedidae
Gerrhosaurinae
Rhineuridae
Anomalepididae
Trogonophiidae
Aniliidae
Azemiopinae
Alopoglossinae
Lanthanotidae
Pseudaspidinae
Tupinambinae
Xenotyphlopidae
Blanidae
Gerrhopilidae
Liolaemidae
Candoiinae
Gallotiinae
Leptotyphlopidae
Platysaurinae
Pythonidae
Boinae
Colubrinae
Opluridae
Elapidae
Typhlopidae
Pareatidae
Cadeidae
Crotalinae
Sibynophiinae
Chamaeleoninae
Rhachisaurinae
Grayiinae
Ungaliophiinae
Erycinae
Phyllodactylidae
Anguinae
Helodermatidae
Phrynosomatidae
Sanziniinae
Tropidophiidae
Viperinae
Cricosaurinae
Shinisauridae
Teiinae
Diplodactylidae
Cylindrophiidae
Brookesiinae
Lepidophyminae
Enyaliinae
Varanidae
Bachiinae
Aparallactinae
Sphaerodactylidae
Atractaspidinae
Uropeltidae
LeiocephalidaeIguanidae
Homalopsidae
Amphibolurinae
Xenopeltidae
Diploglossinae
Calamariinae
Xenosauridae
Uromastycinae
Agaminae
Acrochordidae
Pygopodidae
Calabariidae
Psammophiinae
Carphodactylidae
Cordylinae
Gymnophthalminae
Pseudoxenodontinae
Bolyeriidae
Dactyloidae
Gerrhonotinae
Eublepharidae
Lamprophiinae
Zonosaurinae
83
98
100
100
98
99
100
99
100
99
83
54
98
99
84
100
68
100
100
95
100
94
63
84
71
90
100
100
100
100
100
81
100
100
100
100
87
9969
90
100
100
88
65
97
72
89
100
90
100
100
100
100
96
100
100
96
100
94
100
95
100
97
100
100
100
100
100
100
100
100
58
100
77
100
74
100
95
95
100
99
95
54
100
95
96
79
95
100
99
87
100
100
100
Squamata
Cordylidae
Figure 1 Higher-level squamate phylogeny. Skeletal representation of the 4161-species tree from maximum-likelihood analysis of 12 genes,with tips representing families and subfamilies (following our taxonomic revision; species considered incertae sedis are not shown). Numbers atnodes are SHL values greater than 50%. The full tree is presented in Figures 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23,24, 25, 26, 27, 28.
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 4 of 53http://www.biomedcentral.com/1471-2148/13/93

A
Figure 2 (See legend on next page.)
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 5 of 53http://www.biomedcentral.com/1471-2148/13/93

(See figure on previous page.)Figure 2 Species-level squamate phylogeny. Large-scale maximum likelihood estimate of squamate phylogeny, containing 4161 species.Numbers at nodes are SHL values greater than 50%. A skeletal version of this tree is presented in Figure 1. Bold italic letters indicate figure panels(A-AA). Within panels, branch lengths are proportional to expected substitutions per site, but the relative scale differs between panels.
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 6 of 53http://www.biomedcentral.com/1471-2148/13/93
(Figures 1, 2). We recover clades within the formerGekkonidae that correspond to the strongly supportedfamilies Eublepharidae, Sphaerodactylidae, Phyllodactylidae,and Gekkonidae as in previous studies, and similar rela-tionships among these groups [55-57,59,60,98-100].Within Gekkota, we find evidence for non-monophyly
of many genera. Many relationships among the NewCaledonian diplodactylids are weakly supported (Figure 2),and there is apparent non-monophyly of the generaRhacodactylus, Bavayia, and Eurydactylodes with respectto each other and Oedodera, Dierogekko, Paniegekko,Correlophus, and Mniarogekko [101]. In the Australiandiplodactylids, Strophurus taenicauda is strongly sup-ported as belonging to a clade that is only distantly relatedto the other sampled Strophurus species (Figure 2). Thetwo species of the North African sphaerodactylid genusSaurodactylus are divided between the two majorsphaerodactylid clades (Figure 3), but the associatedbranches are weakly supported. The South Americanphyllodactylid genus Homonota is strongly supportedas being paraphyletic with respect to Phyllodactylus(Figure 3).A number of gekkonid genera (Figure 4) also appear
to be non-monophyletic, including the Asian generaCnemaspis (sampled species divided into two non-sisterclades), Lepidodactylus (with respect to Pseudogekko andsome Luperosaurus), Gekko (with respect to Ptychozoonand Lu. iskandari), Luperosaurus (with respect toLepidodactylus and Gekko), Mediodactylus (with respectto Pseudoceramodactylus, Tropiocolotes, Stenodactylus,Cyrtopodion, Bunopus, Crossobamon, and Agamura), andBunopus (with respect to Crossobamon), and the AfricanAfrogecko (with respect to Afroedura, Christinus, Cryp-tactites, and Matoatoa), Afroedura (with respect toAfrogecko, Blaesodactylus, Christinus, Geckolepis, Pachy-dactylus, Rhoptropus, and numerous other genera),Chondrodactylus (with respect to Pachydactylus laevi-gatus), and Pachydactylus (with respect to Chondro-dactylus and Colopus). Many of these taxonomicproblems in gekkotan families have been identified inprevious studies (e.g. [59,99,102]), and extensive changeswill likely be required to fix them.
ScincoideaWe strongly support (SHL = 100; Figures 1, 5, 6, 7, 8, 9,10) the monophyly of Scincoidea (Scincidae, Xantusiidae,Gerrhosauridae, and Cordylidae), as in other recent stu-dies [16-20]. All four families are strongly supported(Figures 5, 6, 7, 8, 9, 10). A similar clade is also re-
cognized in morphological phylogenies [14], thoughwithout Xantusiidae in some [13].Within the New World family Xantusiidae, we corrob-
orate previous analyses [103,104] that found strong sup-port for a sister-group relationship between Xantusia andLepidophyma, excluding Cricosaura (Figure 5). These re-lationships support the subfamily Cricosaurinae forCricosaura [105]. We also recognize Xantusiinae for theNorth American genus Xantusia and Lepidophyminae forthe Central American genus Lepidophyma [106,107].Within the African and Madagascan family Gerrho-
sauridae (Figure 5), the genus Gerrhosaurus is weakly sup-ported as being paraphyletic with respect to the cladecomprising Tetradactylus + Cordylosaurus, with G. majorplaced as the sister group to all other gerrhosaurids.Within Cordylidae (Figure 5), we use the generic ta-xonomy from a recent phylogenetic analysis and re-classification based on multiple nuclear and mitochondrialgenes [108]. This classification broke up the non-monophyletic Cordylus [109] into several smaller genera,and we corroborate the non-monophyly of the formerCordylus and support the monophyly of the newly recog-nized genera (Figure 5). We support the distinctiveness ofPlatysaurus (Figure 5) and recognition of the subfamilyPlatysaurinae [108].We strong support (SHL = 100) for the monophyly of
Scincidae (Figure 6) as in previous studies (e.g.[20,50,51]). We strongly support the basal placement ofthe monophyletic subfamily Acontiinae (Figure 6), asfound in some previous studies (e.g. [20,51]) but notothers (e.g. [50]). Similar to earlier studies, we find thatthe subfamily Scincinae (sensu [110]) is non-monophyletic,as Feylininae is nested within Scincinae (also found in[20,50,51,111]). Based on these results, synonymizingFeylininae with Scincinae produces a monophyleticScincinae (SHL = 97), which is then sister to a monophy-letic Lygosominae (SHL = 100 excluding Ateuchosaurus;see below) with 94% SHL support (Figures 6, 7, 8, 9,10). This yields a new classification in which all threesubfamilies (Acontiinae, Lygosominae, Scincinae) arestrongly supported. Importantly, these definitions approxi\mate the traditional content of the three subfamilies[50,110], except for recognition of Feylininae.We note that a recent revision of the New World
genus Mabuya introduced a nontraditional family-levelclassification for Scincidae [112]. These authors dividedScincidae into seven families: Acontiidae, Egerniidae,Eugongylidae, Lygosomidae, Mabuyidae, Scincidae andSphenomorphidae. However, there was no phylogenetic

BTarentola gigas
Tarentola caboverdianaTarentola rudis
Tarentola darwiniTarentola chazaliae
Tarentola delalandiiTarentola gomerensis
Tarentola annularisTarentola ephippiata
Tarentola boettgeriTarentola mindiae
Tarentola neglectaTarentola angustimentalisTarentola mauritanica
Tarentola desertiTarentola boehmei
Tarentola americanaPhyllopezus maranjonensis
Phyllopezus lutzaePhyllopezus pollicaris
Phyllopezus periosusPhyllodactylus homolepidurus
Phyllodactylus davisiPhyllodactylus duellmani
Phyllodactylus delcampoiPhyllodactylus paucituberculatus
Phyllodactylus nocticolusPhyllodactylus bugastrolepisPhyllodactylus unctus
Phyllodactylus xantiPhyllodactylus bordai
Phyllodactylus laneiPhyllodactylus tuberculosus
Phyllodactylus reissiiPhyllodactylus wirshingi
Homonota andicolaHomonota darwinii
Homonota borelliiHomonota underwoodi
Homonota fasciataHomonota gaudichaudii
Ptyodactylus guttatusPtyodactylus hasselquistii
Ptyodactylus oudriiPtyodactylus ragazzii
Asaccus platyrhynchusHaemodracon riebeckii
Thecadactylus rapicaudaThecadactylus solimoensis
Sphaerodactylus nicholsiSphaerodactylus ocoae
Sphaerodactylus gaigeaeSphaerodactylus klauberi
Sphaerodactylus semasiopsSphaerodactylus goniorhynchus
Sphaerodactylus townsendiSphaerodactylus armstrongi
Sphaerodactylus macrolepisSphaerodactylus argus
Sphaerodactylus rooseveltiSphaerodactylus altavelensis
Sphaerodactylus notatusSphaerodactylus darlingtoni
Sphaerodactylus cryphiusSphaerodactylus shrevei
Sphaerodactylus elegansSphaerodactylus leucaster
Sphaerodactylus copeiSphaerodactylus nigropunctatus
Sphaerodactylus intermediusSphaerodactylus torrei
Sphaerodactylus cinereusSphaerodactylus thompsoni
Sphaerodactylus glaucusSphaerodactylus molei
Sphaerodactylus oliveriSphaerodactylus richardi
Sphaerodactylus cricoderusSphaerodactylus ramsdeni
Sphaerodactylus schwartziSphaerodactylus kirbyiSphaerodactylus vincenti
Sphaerodactylus microlepisSphaerodactylus parvus
Sphaerodactylus sabanusSphaerodactylus elegantulus
Sphaerodactylus sputatorSphaerodactylus fantasticus
Pseudogonatodes guianensisPseudogonatodes lunulatus
Pseudogonatodes manessiColeodactylus natalensis
Coleodactylus meridionalisColeodactylus brachystoma
Coleodactylus septentrionalisGonatodes ocellatusGonatodes ceciliaeGonatodes seigliei
Gonatodes concinnatusGonatodes antillensis
Gonatodes humeralisGonatodes falconensis
Gonatodes taniaeGonatodes purpurogularis
Gonatodes alexandermendesiGonatodes superciliaris
Gonatodes annularisGonatodes hasemani
Gonatodes infernalisGonatodes vittatus
Gonatodes petersiGonatodes albogularis
Gonatodes daudiniGonatodes caudiscutatus
Gonatodes eladioiLepidoblepharis festae
Lepidoblepharis xanthostigmaChatogekko amazonicus
Saurodactylus mauritanicusTeratoscincus roborowskii
Teratoscincus przewalskiiTeratoscincus scincusTeratoscincus microlepisEuleptes europaea
Saurodactylus fasciatusAristelliger georgeensis
Aristelliger praesignisAristelliger lar
Quedenfeldtia moerensQuedenfeldtia trachyblepharus
Pristurus somalicusPristurus crucifer
Pristurus carteriPristurus minimusPristurus rupestrisPristurus flavipunctatus
Pristurus abdelkuriPristurus guichardi
Pristurus sokotranusPristurus insignis
Pristurus celerrimusGoniurosaurus araneus
Goniurosaurus luiiGoniurosaurus catbaensis
Goniurosaurus lichtenfelderiGoniurosaurus kuroiwae
Hemitheconyx tayloriHemitheconyx caudicinctus
Holodactylus africanusEublepharis turcmenicusEublepharis macularius
Coleonyx elegansColeonyx mitratus
Coleonyx variegatusColeonyx brevis
Aeluroscalabotes felinus
100
100
92
73100
99
9376
9162
100 99
73
100
100
87100
99
10090
92
93
100
10065
100
99
100
6775
96 57
97
97 100
86
10085
92
65 100100
98
100
100
83
100
79
82
87
100
83
84
69
93
8590
859384
8366
9499
79
52
99
83
81
100 100
91
84
100
84
98
81
82 100
87
10096
10099
100
99
100
60
97
89
99100
10093
10095
98
7060
100
89100
96
97
100
60
78100
9798
75
100100
91
100
100
76
10063
85 98
92100
100
100
100
97100
9690
100
100 100
100
79
100100
100
100
Phyllodactylidae
Eublepharidae
Sphaerodactylidae
C
Figure 3 Species-level squamate phylogeny continued (B).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 7 of 53http://www.biomedcentral.com/1471-2148/13/93

CPhelsuma kely
Phelsuma lineataPhelsuma comorensis
Phelsuma pusillaPhelsuma quadriocellata
Phelsuma antanosyPhelsuma klemmeri
Phelsuma robertmertensiPhelsuma laticaudaPhelsuma serraticaudaPhelsuma malamakibo
Phelsuma berghofiPhelsuma hielscheriPhelsuma flavigularis
Phelsuma ravenalaPhelsuma dubiaPhelsuma modesta
Phelsuma nigristriataPhelsuma inexpectata
Phelsuma ornataPhelsuma guimbeaui
Phelsuma cepedianaPhelsuma borbonica
Phelsuma guentheriPhelsuma gigas
Phelsuma edwardnewtoniPhelsuma standingi
Phelsuma andamanensePhelsuma mutabilis
Phelsuma brevicepsPhelsuma pronkiPhelsuma barbouri
Phelsuma seippiPhelsuma parkeri
Phelsuma abbottiPhelsuma madagascariensis
Phelsuma guttataPhelsuma sundbergiPhelsuma astriata
Phelsuma vanheygeniLygodactylus keniensis
Lygodactylus kimhowelliLygodactylus luteopicturatusLygodactylus picturatus
Lygodactylus williamsiLygodactylus chobiensis
Lygodactylus gutturalisLygodactylus angularis
Lygodactylus thomensisLygodactylus conraui
Lygodactylus klugeiLygodactylus bradfieldiLygodactylus capensisLygodactylus stevensoni
Lygodactylus lawrenceiLygodactylus pictus
Lygodactylus mirabilisLygodactylus tuberosus
Lygodactylus montanusLygodactylus gravis
Lygodactylus paulianiLygodactylus arnoulti
Lygodactylus blancaeLygodactylus verticillatusLygodactylus heterurus
Lygodactylus tolampyaeLygodactylus guibei
Lygodactylus miopsLygodactylus madagascariensis
Lygodactylus rarusLygodactylus expectatus
Rhoptropella ocellataCnemaspis tropidogaster
Cnemaspis kandianaCnemaspis podihuna
Pachydactylus affinisPachydactylus vansoni
Pachydactylus capensisPachydactylus oshaughnessyi
Pachydactylus tigrinusPachydactylus montanus
Pachydactylus vanzyliPachydactylus rangei
Pachydactylus austeniPachydactylus mariquensis
Pachydactylus mclachlaniPachydactylus monicaePachydactylus weberi
Pachydactylus servalPachydactylus griffini
Pachydactylus carinatusPachydactylus fasciatus
Pachydactylus tsodiloensisPachydactylus waterbergensis
Pachydactylus purcelliPachydactylus oculatus
Pachydactylus maculatusPachydactylus geitje
Pachydactylus labialisPachydactylus barnardi
Pachydactylus formosusPachydactylus rugosus
Pachydactylus punctatusPachydactylus scherzi
Pachydactylus sansteynaePachydactylus caraculicus
Pachydactylus bicolorPachydactylus oreophilus
Pachydactylus gaiasensisPachydactylus reconditus
Pachydactylus parascutatusPachydactylus scutatus
Pachydactylus namaquensisPachydactylus kladaroderma
Pachydactylus haackeiColopus kochii
Colopus wahlbergiiPachydactylus robertsi
Chondrodactylus bibroniiChondrodactylus fitzsimonsi
Chondrodactylus turneriPachydactylus laevigatus
Chondrodactylus anguliferElasmodactylus tuberculosus
Elasmodactylus tetensisRhoptropus barnardi
Rhoptropus biporosusRhoptropus boultoni
Rhoptropus bradfieldiRhoptropus afer
Goggia lineataBlaesodactylus sakalava
Blaesodactylus antongilensisBlaesodactylus boiviniHomopho lis mulleri
Homopholis walbergiiHomopholis fasciataGeckolepis maculataGeckolepis typica
Afroedura karroicaAfroedura pondolia
Afrogecko porphyreusMatoatoa brevipes
Cryptactites peringueyiAfrogecko swartbergensis
Christinus marmoratusParagehyra gabriellae
Uroplatus fimbriatusUroplatus giganteus
Uroplatus henkeliUroplatus sikorae
Uroplatus lineatusUroplatus pietschmanni
Uroplatus alluaudiUroplatus phantasticus
Uroplatus ebenauiUroplatus malama
Uroplatus malaheloUroplatus guentheri
Cnemaspis africanaCnemaspis dickersonae
Cnemaspis uzungwaeNarudasia festiva
Ptenopus carpiCalodactylodes aureusCalodactylodes illingworthorum
Ailuronyx trachygasterAiluronyx seychellensis
Ailuronyx tachyscopaeusParoedura bastardi
Paroedura tanjakaParoedura vazimba
Paroedura androyensisParoedura picta
Paroedura sanctijohannisParoedura stumpffi
Paroedura lohatsaraParoedura karstophila
Paroedura ovicepsParoedura homalorhina
Paroedura gracilisParoedura masobe
Ebenavia inunguisUrocotyledon inexpectata
Perochirus ateles
69
89
95
90
100
100
100
959987
99100
5794
96
95
100
9894
99
100
55
85
9499
95100
100100
99
8292
96 100
100
100100
87
92
100
100
99
68
9778
6794
9468
9793
100
81
899384
8273
98
10081
91 85
100
7198 92
100 99
52
98
99
100
96
100
89
70
85
61
74
72100 100
100
99
100
100
85
69
98
68
97
100
84
52 93
99
68
100
6386
96
91
74
95 90
100
98
100
93
100100 86
100
86
10098 98
79
93
10099 100
99 100100
56
10052 100
97
100
65
53
92
10085
50
10077
100100
100
100
100
87100
98 100
100
100
93
98
100
99
9261
99100
100
85
83100
99
73
(ii)
Hemidactylus homoeolepisHemidactylus forbesii
Hemidactylus oxyrhinusHemidactylus macropholis
Hemidactylus mindiaeHemidactylus lemurinus
Hemidactylus turcicusHemidactylus robustus
Hemidactylus yerburiiHemidactylus persicus
Hemidactylus grantiHemidactylus dracaenacolus
Hemidactylus pumilioHemidactylus foudaii
Hemidactylus citerniiHemidactylus modestus
Hemidactylus agriusHemidactylus palaichthusHemidactylus bouvieri
Hemidactylus brasilianusHemidactylus greeffiiHemidactylus platycephalus
Hemidactylus longicephalusHemidactylus mercatoriusHemidactylus mabouia
Hemidactylus haitianusHemidactylus angulatus
Hemidactylus gracilisHemidactylus reticulatus
Hemidactylus albofasciatusHemidactylus imbricatus
Hemidactylus sataraensisHemidactylus brookii
Hemidactylus frenatusHemidactylus flaviviridis
Hemidactylus leschenaultiiHemidactylus maculatu s
Hemidactylus prashadiHemidactylus triedrus
Hemidactylus depressusHemidactylus giganteus
Hemidactylus aaronbaueriHemidactylus fasciatus
Hemidactylus platyurusHemidactylus karenorum
Hemidactylus garnotiiHemidactylus bowringii
Cyrtodactylus louisiadensisCyrtodactylus epiroticusCyrtodactylus tripartitus
Cyrtodactylus robustusCyrtodactylus klugei
Cyrtodactylus tuberculatusCyrtodactylus novaeguineae
Cyrtodactylus loriaeCyrtodactylus sermowaiensis
Cyrtodactylus pulchellusCyrtodactylus intermedius
Cyrtodactylus agusanensisCyrtodactylus philippinicus
Cyrtodactylus annulatusCyrtodactylus consobrinus
Cyrtodactylus irregularisCyrtodactylus marmoratus
Cyrtodactylus triedrusCyrtodactylus jarujini
Cyrtodactylus angularisCyrtodactylus ayeyarwadyensis
Cyrtodactylus oldhamiCyrtopodion gastropholeCyrtopodion agamuroides
Cyrtopodion caspiumCyrtopodion longipes
Cyrtopodion scabrumCyrtopodion sistanensis
Agamura persicaBunopus crassicauda
Crossobamon orientalisBunopus tuberculatus
Mediodactylus heteropholisMediodactylus heterocercum
Mediodactylus sagittiferumMediodactylus kotschyi
Stenodactylus sthenodactylusStenodactylus yemenensis
Stenodactylus doriaeStenodactylus leptocosymbotus
Stenodactylus petriiStenodactylus arabicus
Tropiocolotes tripolitanusPseudoceramodactylus khobarensis
Mediodactylus russowiiMediodactylus spinicaudum
Cnemaspis kendalliiCnemaspis limi
Tropiocolotes helenaeAlsophylax pipiens
Gehyra minutaGehyra montium
Gehyra pilbaraGehyra punctata
Gehyra variegataGehyra purpurascensGehyra nana
Gehyra xenopusGehyra australis
Gehyra occidentalisGehyra koira
Gehyra robustaGehyra borroloola
Gehyra pamelaGehyra catenata
Gehyra dubiaGehyra membranacruralis
Gehyra oceanicaGehyra marginata
Gehyra bareaGehyra baliola
Gehyra brevipalmataGehyra lacerata
Gehyra mutilataGehyra fehlmanni
Hemiphyllodactylus aurantiacusHemiphyllodactylus yunnanensis
Hemiphyllodactylus typusNactus pelagicusNactus multicarinatus
Nactus chevertiNactus galgajuga
Nactus vankampeniNactus eboracensisNactus acutus
Heteronotia speleaHeteronotia binoeiHeteronotia planiceps
Dixonius siamensisDixonius vietnamensis
Dixonius melanostictusGekko grossmanni
Gekko badeniiGekko petricolus
Gekko vittatusLuperosaurus iskandari
Ptychozoon lionotumPtychozoon kuhli
Ptychozoon rhacophorusGekko porosusGekko crombota
Gekko romblonGekko mindorensis
Gekko monarchusGekko athymus
Gekko swinhonisGekko auriverrucosus
Gekko hokouensisGekko japonicus
Gekko chinensisGekko gecko
Gekko smithiiLepidodactylus moestusLepidodactylus lugubris
Luperosaurus joloensisLuperosaurus cumingii
Luperosaurus macgregoriLepidodactylus orientalis
Pseudogekko compressicorpusPseudogekko smaragdinus
Lepidodactylus novaeguineae
99
100
97
100
99
98
97
91
91
96
6697
98
99
82
100
9698
57
100
100
9695 97
97
100
100
81
93
100100
9956
97
100
93
8988
76
100
100100
87
99
92
94
9898
99
98
77
96
79
88
10052
9798
92
100
100
75
95
79
99
100
99
100
86
9969
99
98
9692
10096
85
100
100
65 100
64
100
100
96
55
9898
9892
100
10097
70100
100
97
8459
91
100
100
10085
100
10083
98 100
100
100
100
100 100
100
100
100
9496
97
89
95
10099
80100
100
100
9598
54
100
100
7991 100
100
100
Gek
kon
idae
(i)
100
(i)
(ii)
Figure 4 Species-level squamate phylogeny continued (C).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 8 of 53http://www.biomedcentral.com/1471-2148/13/93

DCordylus cordylusCordylus tasmani
Cordylus oelofseniCordylus mclachlani
Cordylus nigerCordylus minorCordylus aridusCordylus imkeae
Cordylus macropholisCordylus rhodesianus
Cordylus jonesiiCordylus beraducciiCordylus ukingensisCordylus vittifer
Cordylus meculaeCordylus tropidosternum
Hemicordylus capensisHemicordylus nebulosusNinurta coeruleopunctatusPseudocordylus melanotusPseudocordylus spinosus
Pseudocordylus microlepidotusPseudocordylus langi
Chamaesaura anguinaChamaesaura aenea
Smaug giganteusSmaug warreni
Namazonurus peersiNamazonurus lawrenci
Namazonurus namaquensisNamazonurus pustulatus
Namazonurus campbelliKarusasaurus polyzonus
Karusasaurus jordaniOuroborus cataphractus
Platysaurus monotropisPlatysaurus minor
Platysaurus intermediusPlatysaurus capensis
Platysaurus broadleyiPlatysaurus mitchelli
Platysaurus pungweensisZonosaurus rufipes
Zonosaurus aeneusZonosaurus bemaraha
Zonosaurus subunicolorZonosaurus brygooiZonosaurus tsingy
Zonosaurus boettgeriZonosaurus laticaudatus
Zonosaurus anelanelanyZonosaurus karsteni
Zonosaurus quadrilineatusZonosaurus trilineatus
Zonosaurus ornatusZonosaurus madagascariensis
Zonosaurus haraldmeieriTracheloptychus petersi
Tracheloptychus madagascariensisGerrhosaurus flavigularis
Gerrhosaurus multilineatusGerrhosaurus nigrolineatus
Gerrhosaurus typicusGerrhosaurus skoogi
Gerrhosaurus validusTetradactylus tetradactylus
Tetradactylus africanusTetradactylus seps
Cordylosaurus subtessellatusGerrhosaurus major
Lepidophyma dontomasiLepidophyma radula
Lepidophyma loweiLepidophyma cuicatecaLepidophyma smithii
Lepidophyma lineriLepidophyma gaigeae
Lepidophyma sylvaticumLepidophyma micropholis
Lepidophyma occulorLepidophyma pajapanensis
Lepidophyma reticulatumLepidophyma flavimaculatumLepidophyma lipetziLepidophyma tuxtlae
Lepidophyma mayaeXantusia wigginsi
Xantusia bezyiXantusia vigilis
Xantusia arizonaeXantusia riversiana
Xantusia sancheziXantusia bolsonaeXantusia henshawi
Xantusia gracilisCricosaura typica
99
100
100
100
55
95
96
99
100
8586
100
84
92
83100
99
818283
88
95
86100
93
10099
100
94
100
100
99
100
92
91100
100
100
91
96
6682
8987
73
100100
100
100
100
100
79
91100
76
95
9869
100
100
100
87
95
9594
100100
94
100
100
100
100
100
8492
10066
97
100
100
Platysaurinae
Cordylinae
Cordylidae
Gerrhosauridae
Gerrhosaurinae
Zonosaurinae
Lepidophyminae
XantusiinaeCricosaurinae
Xantusiidae
Figure 5 Species-level squamate phylogeny continued (D).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 9 of 53http://www.biomedcentral.com/1471-2148/13/93

EAmphiglossus frontoparietalisAmphiglossus punctatus
Amphiglossus macrocercusAmphiglossus anosyensis
Amphiglossus splendidusVoeltzkowia rubrocaudataVoeltzkowia lineata
Voeltzkowia fierinensisAmphiglossus astrolabi
Amphiglossus reticulatusAmphiglossus tsaratananensis
Androngo trivittatusPygomeles braconnieri
Amphiglossus tanysomaAmphiglossus mandokava
Amphiglossus ornaticepsAmphiglossus melanurus
Madascincus intermediusMadascincus polleni
Madascincus stumpffiMadascincus nanus
Madascincus mouroundavaeMadascincus igneocaudatus
Pseudoacontias menamaintyMadascincus melanopleura
Paracontias rothschildiParacontias hildebrandti
Paracontias manifyParacontias brocchii
Paracontias holomelasScelotes kasneri
Scelotes montispectusScelotes gronovii
Scelotes sexlineatusScelotes bipes
Scelotes arenicolusScelotes mirus
Scelotes anguineusScelotes caffer
Proscelotes eggeliHakaria simonyi
Melanoseps loveridgeiMelanoseps ater
Melanoseps occidentalisFeylinia currori
Feylinia polylepisFeylinia grandisquamis
Typhlacontias punctatissimusTyphlacontias brevipes
Sepsina angolensisChalcides montanus
Chalcides polylepisChalcides manueli
Chalcides mionectonChalcides sphenopsiformis
Chalcides viridanusChalcides coeruleopunctatus
Chalcides sexlineatusChalcides lanzaiChalcides parallelus
Chalcides boulengeriChalcides bedriagaiChalcides colosii
Chalcides sepsoidesChalcides ocellatus
Chalcides striatusChalcides pseudostriatusChalcides chalcides
Chalcides guentheriChalcides minutus
Chalcides mauritanicusGongylomorphus bojerii
Janetaescincus braueriJanetaescincus veseyfitzgeraldi
Pamelaescincus gardineriScincus mitranusScincus scincus
Eumeces algeriensisScincopus fasciatus
Eumeces schneideriEurylepis taeniolatus
Plestiodon dugesiiPlestiodon brevirostris
Plestiodon copeiPlestiodon parvulus
Plestiodon ochoterenaePlestiodon lynxe
Plestiodon sumichrastiPlestiodon parviauriculatus
Plestiodon skiltonianusPlestiodon lagunensis
Plestiodon gilbertiPlestiodon longirostris
Plestiodon septentrionalisPlestiodon multivirgatus
Plestiodon fasciatusPlestiodon obsoletus
Plestiodon callicephalusPlestiodon tetragrammus
Plestiodon laticepsPlestiodon inexpectatus
Plestiodon anthracinusPlestiodon egregius
Plestiodon reynoldsiPlestiodon stimpsonii
Plestiodon marginatusPlestiodon elegans
Plestiodon latiscutatusPlestiodon japonicusPlestiodon barbouri
Plestiodon capitoPlestiodon tunganus
Plestiodon quadrilineatusPlestiodon chinensis
Plestiodon kishinouyeiPlestiodon tamdaoensis
Brachymeles pathfinderiBrachymeles gracilis
Brachymeles bicolorBrachymeles boulengeri
Brachymeles schadenbergiBrachymeles eleraeBrachymeles talinis
Brachymeles samarensisBrachymeles cebuensis
Brachymeles minimusBrachymeles bonitae
Brachymeles tridactylusBrachymeles miriamae
Brachymeles apusOphiomorus latastiiOphiomorus punctatissimus
Mesoscincus managuaeMesoscincus schwartzei
Acontias poecilusAcontias plumbeus
Acontias gracilicaudaAcontias breviceps
Acontias rieppeliAcontias percivali
Acontias meleagrisAcontias kgalagadi
Acontias litoralisAcontias lineatus
Acontias gariepensisTyphlosaurus lomiaeTyphlosaurus vermisTyphlosaurus caecusTyphlosaurus meyeri
Typhlosaurus braini
100
97
81
100
97
92
100
100
85
93
24
8070
9096
10095
93
86
100
90100
100
100
5297
64 100
87
89
100
6069
93
100
99
100 97
7594
8089
87
97
100
96
96100
100
98
100
57
100
88
100
9495
96
9899
74
9174
8799
55
10086
75
92
99
100
100100 100
77
85
100
70
88
100
1007376
98
9173
92
100
97
56100
747671
98 86
74
100
100
93100
86
97
98
100 100
10094
95
8388
85
85
73
94
99
91
99
98
100
9694
9898
92100
100
9599
100
89
94
Scincidae
Acontiinae
Scincinae
F-I
Figure 6 Species-level squamate phylogeny continued (E).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 10 of 53http://www.biomedcentral.com/1471-2148/13/93

Lygosominae
Lygosominaecont.F
Papuascincus stanleyanusLipinia noctuaLipinia pulchella
Insulasaurus victoriaInsulasaurus wrighti
Insulasaurus arborensPrasinohaema virens
Sphenomorphus buenloicusScincella assatus
Scincella cherrieiScincella gemmingeri
Scincella lateralisAsymblepharus sikimmensis
Scincella reevesiiSphenomorphus sabanusSphenomorphus cyanolaemus
Sphenomorphus variegatusSphenomorphus multisquamatus
Sphenomorphus indicusSphenomorphus maculatus
Larutia seribuatensisParvoscincus lawtoni
Parvoscincus kitangladensisParvoscincus luzonenseParvoscincus laterimaculatusParvoscincus beyeri
Parvoscincus sisoniParvoscincus tagapayo
Parvoscincus leucospilosParvoscincus decipiens
Parvoscincus steereiSphenomorphus acutusSphenomorphus diwataPinoyscincus llanosi
Pinoyscincus abdictusPinoyscincus coxi
Pinoyscincus jagoriPinoyscincus mindanen sis
Otosaurus cumingiSphenomorphus maindroni
Sphenomorphus craneiSphenomorphus leptofasciatus
Sphenomorphus fasciatusSphenomorphus solomonis
Sphenomorphus scutatusSphenomorphus concinnatus
Tytthoscincus atrigularisTytthoscincus hallieri
Tytthoscincus parvusTytthoscincus aesculeticola
Sphenomorphus muelleriSphenomorphus jobiensis
Sphenomorphus melanopogonIsopachys anguinoides
Lipinia vittigeraTropidophorus misaminiusTropidophorus partelloi
Tropidophorus brookeiTropidophorus beccarii
Tropidophorus baconiTropidophorus grayiTropidophorus microlepis
Tropidophorus cocincinensisTropidophorus thaiTropidophorus robinsoni
Tropidophorus sinicusTropidophorus baviensis
Tropidophorus hainanusTropidophorus murphyi
Tropidophorus noggeiTropidophorus latiscutatusTropidophorus matsuiiTropidophorus berdmorei
Sphenomorphus simusSphenomorphus praesignis
Ablepharus pannonicusAsymblepharus alaicus
Ateuchosaurus pellopleurus
100
98
89
97
77
94
93
76
99
91
99
98
99
94100
100
80
65
9693
5499
100
100100
100
99
90
96
100
89
100
97
996883
100
100
100
80
100
95
100
1008776
100
100
10092
98
86
100
10095
94
89
95
83
99
98
10098
60100
90
81
100
H-I GFigure 7 Species-level squamate phylogeny continued (F).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 11 of 53http://www.biomedcentral.com/1471-2148/13/93

Lerista kennedyensisLerista onsloviana
Lerista uniduoLerista connivens
Lerista variaLerista lineopunctulata
Lerista yunaLerista kendricki
Lerista nichollsiLerista gascoynensis
Lerista petersoniLerista humphriesi
Lerista praepeditaLerista bougainvillii
Lerista viduataLerista muelleri
Lerista haroldiLerista allochira
Lerista stictopleuraLerista planiventralis
Lerista lineataLerista christinae
Lerista distinguendaLerista elegans
Lerista emmottiLerista punctatovittata
Lerista dorsalisLerista terdigitata
Lerista tridactylaLerista elongata
Lerista speciosaLerista zietzi
Lerista flammicaudaLerista picturata
Lerista baynesiLerista edwardsae
Lerista arenicolaLerista microtis
Lerista macropisthopusLerista axillaris
Lerista gerrardiiLerista eupoda
Lerista desertorumLerista puncticaudaLerista neander
Lerista zonulataLerista ingrami
Lerista orientalisLerista taeniata
Lerista aericepsLerista xanthura
Lerista chordaeLerista fragilis
Lerista frostiLerista borealis
Lerista walkeriLerista simillima
Lerista vermicularisLerista robusta
Lerista greeriLerista labialis
Lerista bipesLerista ips
Lerista griffiniLerista apoda
Lerista kalumburuLerista wilkinsiLerista cinerea
Lerista amelesLerista carpentariae
Lerista karlschmidtiLerista stylis
Ctenotus pulchellusCtenotus gagudju
Ctenotus hilliCtenotus hebetior
Ctenotus essingtoniiCtenotus piankai
Ctenotus hanloniCtenotus angusticeps
Ctenotus grandisCtenotus atlas
Ctenotus maryaniCtenotus olympicus
Ctenotus regiusCtenotus septenarius
Ctenotus astarteCtenotus mimetes
Ctenotus tanamiensisCtenotus greeri
Ctenotus serventyiCtenotus quattuordecimlineatus
Ctenotus leonhardiiCtenotus leae
Ctenotus australisCtenotus uber
Ctenotus rutilansCtenotus taeniolatus
Ctenotus robustusCtenotus spaldingi
Ctenotus inornatusCtenotus saxatilis
Ctenotus fallensCtenotus rawlinsoni
Ctenotus calurusCtenotus schomburgkiiCtenotus youngsoni
Ctenotus strauchiiCtenotus pantherinus
Ctenotus nasutusCtenotus rubicundus
Ctenotus brooksiCtenotus labillardieri
Notoscincus ornatusEulamprus tympanum
Eulamprus heatwoleiEulamprus kosciuskoi
Eulamprus leuraensisEulamprus quoyii
Glaphyromorphus darwiniensisGlaphyromorphus cracens
Glaphyromorphus pumilusGlaphyromorphus fuscicaudis
Glaphyromorphus mjobergiGlaphyromorphus punctulatus
Hemiergis decresiensisHemiergis millewae
Hemiergis gracilipesHemiergis quadrilineatum
Hemiergis peroniiHemiergis initialis
Eremiascincus isolepisEremiascincus douglasiEremiascincus richardsoniiEremiascincus pardal is
Eremiascincus fasciolatusAnomalopus leuckartii
Anomalopus verreauxiAnomalopus mackayi
Anomalopus swansoniOphioscincus truncatus
Coeranoscincus reticulatusSaiphos equalis
Ophioscincus ophioscincusCoeranoscincus frontalis
Coggeria naufragusEulamprus murrayi
Eulamprus tryoniEulamprus luteilateralis
Eulamprus brachyosomaEulamprus sokosoma
Eulamprus martiniEulamprus tigrinus
Eulamprus tenuisEulamprus amplus
Gnypetoscincus queenslandiaeCalyptotis ruficauda
Calyptotis scutirostrumCalyptotis lepidorostrum
Nangura spinosaEulamprus frerei
100
95
55
100
85
82
87
90
93
93
89
98
84100
96
100100
1008177
56100
100
93
8996
90
92
100
90
100
61100
10099
100
100
100
98
90
100
99
10093
79
87
100
99
100100
100
96
8997
99
91100
9993
9890 100
88
10099
97
95
81
99
95
85
67
76
95
86
66
51
85
78
9165
90
7992
97
83
8873
63
100100
88
89
84
85
99
81
100
76
70
93
78
93 100
8287
95
99
91
100
82 60
100100
10097
62 88
100100
97
80
91
9595
77
8583
98
7896
88 100
10077
G
Lygosominaecont.
Figure 8 Species-level squamate phylogeny continued (G).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 12 of 53http://www.biomedcentral.com/1471-2148/13/93

Mabuya heathiMabuya caissara
Mabuya agilisMabuya guaporicola
Mabuya macrorhynchaMabuya agmosticha
Mabuya frenataMabuya bistriata
Mabuya falconensisMabuya unimarginata
Mabuya mabouyaMabuya meridensis
Mabuya dorsivittataMabuya cochabambae
Mabuya nigropunctataMabuya altamazonica
Mabuya sloaniiMabuya nigropalmata
Mabuya croizatiMabuya carvalhoiChioninia stangeri
Chioninia fogoensisChioninia spinalis
Chioninia cocteiChioninia delalandii
Chioninia vaillantiiEumecia anchietae
Trachylepis dumasiTrachylepis aureopunctata
Trachylepis vatoTrachylepis boettgeriTrachylepis madagascariensis
Trachylepis elegansTrachylepis gravenhorstiiTrachylepis acutilabris
Trachylepis homalocephalaTrachylepis sulcata
Trachylepis variegataTrachylepis hoeschi
Trachylepis striataTrachylepis spilogaster
Trachylepis occidentalisTrachylepis capensis
Trachylepis variaTrachylepis atlantica
Trachylepis margaritiferaTrachylepis quinquetaeniata
Trachylepis perrotetiiTrachylepis affinis
Trachylepis sechellensisTrachylepis wrightii
Trachylepis maculilabrisTrachylepis socotrana
Trachylepis brevicollisTrachylepis vittata
Trachylepis aurataDasia olivacea
Dasia griseaDasia vittata
Eutropis trivittataEutropis nagarjuniEutropis beddomii
Eutropis bibroniiEutropis clivicola
Eutropis multicarinataEutropis cumingi
Eutropis multifasciataEutropis macrophthalma
Eutropis rudisEutropis macularia
Eutropis longicaudataLankascincus fallax
Ristella rurkiiLiopholis montana
Liopholis guthegaLiopholis whitiiLiopholis margaretae
Liopholis modestaLiopholis pulchra
Liopholis kintoreiLiopholis multiscutata
Liopholis inornataLiopholis striata
Tiliqua scincoidesTiliqua gigas
Tiliqua nigroluteaTiliqua occipitalis
Tiliqua rugosaTiliqua adelaidensisCyclodomorphus branchialis
Cyclodomorphus casuarinaeCyclodomorphus michaeli
Egernia hosmeriEgernia stokesii
Lissolepis luctuosaEgernia napoleonis
Egernia richardiBellatorias frerei
Egernia kingiiEgernia depressa
Bellatorias majorEgernia saxatilis
Corucia zebrataTribolonotus pseudoponceleti
Tribolonotus ponceletiTribolonotus brongersmai
Tribolonotus schmidtiTribolonotus blanchardi
Tribolonotus gracilisTribolonotus novaeguineae
83
97
91
99
87
100
98
89
81
70
7780
10099
100
97
9695
100
78
9098
80
100
91
98
92
84
90
74
98100
9795
10095
95
54
7889
99100
100
100
100
99
83100
99
93
97
95
90
9796 100
80
73
100
96
88
94
93
100
96
91
99
5652
90
73
9890
85
88
97
8471
100
8195
66
86
74
73
90
77
100
8697
93 96
100
Lygosominaecont.
H
Figure 9 Species-level squamate phylogeny continued (H).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 13 of 53http://www.biomedcentral.com/1471-2148/13/93

Carlia vivaxCarlia dogare
Carlia rostralisCarlia amax
Carlia tetradactylaCarlia rubrigularis
Carlia rhomboidalisCarlia rufilatus
Carlia pectoralisCarlia munda
Carlia mysiCarlia longipes
Carlia fuscaCarlia storri
Carlia schmeltziiLygisaurus malleolus
Lygisaurus tanneriLygisaurus absconditaLygisaurus foliorum
Lygisaurus laevisLygisaurus aeratus
Lygisaurus novaeguineaeLygisaurus macfarlani
Lygisaurus parrhasiusLygisaurus sesbrauna
Carlia gracilisCarlia bicarinata
Carlia jarnoldaeCarlia johnstonei
Carlia triacanthaMenetia timlowi
Liburnascincus mundivensisLiburnascincus scirtetis
Liburnascincus coensisCautula zia
Niveoscincus metallicusNiveoscincus pretiosus
Niveoscincus ocellatusNiveoscincus greeni
Proablepharus reginaeLampropholis robertsi
Lampropholis coggeriLampropholis delicata
Lampropholis guichenotiSaproscincus spectabilis
Saproscincus roseiSaproscincus challengeri
Saproscincus mustelinusSaproscincus oriarus
Saproscincus hannahaeSaproscincus tetradactylus
Saproscincus czechuraiSaproscincus lewisi
Saproscincus basiliscusBartleia jigurru
Morethia adelaidensisMorethia butleri
Morethia ruficaudaBassiana trilineata
Emoia isolataEmoia pseudocyanura
Emoia cyanuraEmoia impar
Emoia physicaeEmoia jakati
Emoia atrocostataEmoia caeruleocauda
Emoia cyanogasterEmoia schmidti
Menetia alanaeMenetia greyii
Cryptoblepharus novocaledonicusCryptoblepharus boutonii
Cryptoblepharus nigropunctatusCaledoniscincus terma
Caledoniscincus aquiloniusCaledoniscincus chazeaui
Caledoniscincus atropunctatusCaledoniscincus haplorhinus
Caledoniscincus austrocaledonicusCaledoniscincus festivus
Caledoniscincus auratusCaledoniscincus renevieri
Caledoniscincus orestesSimiscincus aurantiacus
Graciliscincus shonaeTropidoscincus variabilis
Tropidoscincus boreus
Tropidoscincus aubrianusLioscincus tillieri
Lioscincus maruiaLioscincus novaecaledoniaeSigaloseps deplanchei
Sigaloseps ruficaudaPhoboscincus garnieri
Lacertoides pardalisLioscincus nigrofasciolatum
Kanakysaurus viviparusLioscincus steindachneri
Lioscincus vivaeCelatiscincus euryotis
Celatiscincus similisMarmorosphax tricolor
Marmorosphax montanaNannoscincus humectus
Nannoscincus hanchisteusNannoscincus greeriNannoscincus mariei
Nannoscincus gracilisNannoscincus slevini
Nannoscincus garrulusOligosoma whitakeri
Oligosoma oliveriOligosoma townsi
Oligosoma ornatumOligosoma macgregori
Oligosoma mocoOligosoma alani
Oligosoma fallaiOligosoma hardyi
Oligosoma levidensumOligosoma aeneum
Oligosoma microlepisOligosoma smithi
Oligosoma striatumOligosoma homalonotumOligosoma zelandicum
Oligosoma notosaurusOligosoma inconspicuumOligosoma maccanni
Oligosoma stenotisOligosoma grande
Oligosoma nigriplantareOligosoma longipes
Oligosoma lineoocellatumOligosoma chloronoton
Oligosoma otagenseOligosoma waimatenseOligosoma infrapunctatum
Oligosoma acrinasumOligosoma taumakaeOligosoma pikitanga
Oligosoma suteriOligosoma lichenigera
Pseudemoia entrecasteauxiiBassiana duperreyi
Pseudemoia pagenstecheriLeptosiaphos vigintiserierumLeptosiaphos kilimensis
Leptosiaphos amietiLeptosiaphos graueri
Leptosiaphos hackarsiLacertaspis lepesmei
Lacertaspis gemmiventrisLacertaspis chriswildi
Lacertaspis rohdeiLacertaspis reichenowi
Afroablepharus africanusAfroablepharus annobonensis
Afroablepharus wahlbergiPanaspis togoensis
Panaspis brevicepsLeiolopisma mauriti anaLeiolopisma telfairii
Emoia loyaltiensisEugongylus albofasciolatusEugongylus rufescens
Lygosoma bowringiiMochlus brevicaudisLygosoma punctata
Lygosoma sundevalliMochlus afer
Lygosoma lineolatumLepidothyris fernandi
Lygosoma albopunctataLygosoma koratense
Lygosoma quadrupesLamprolepis smaragdina
Emoia tonganaEmoia concolor
Sphenomorphus stellatusAblepharus kitaibelii
Ablepharus chernoviAblepharus budaki
90
100
100
84
52
79
100
72
87
88
98
90
88
68
80
85
7653
62
92
100
93
98100
73
91
75
98
8578
89 100
100 100
99
79
100
96
91
76100
92100
72
76
71100 97
100
96
98
96
73
93
91
100
7798 71 98
98 100
100
93
100
79
61
99
96
96
98
87
97
95
83
5387
9598
95
97
100
100 100
80
8585 92
100
80
90
9696
100100
98
8293
94
98
89
100
98
90
94
57
9893
899796
63
93
99
100
100100
86
99
8598 100
98
99
100
99
9891 100
95
9980
75
84
85
100
8195 99
95
9396
91
90
9999 85
100
100
100
100
6360
9593
7670
92 100
98
96
I
Lygosominaecont.
Figure 10 Species-level squamate phylogeny continued (I).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 14 of 53http://www.biomedcentral.com/1471-2148/13/93

Gymnophthalmus cryptusGymnophthalmus underwoodi
Gymnophthalmus speciosusGymnophthalmus vanzoiGymnophthalmus leucomystax
Gymnophthalmus pleeiPsilophthalmus paeminosus
Calyptommatus nicterusCalyptommatus leiolepis
Calyptommatus confusionibusCalyptommatus sinebrachiatus
Nothobachia ablepharaProcellosaurinus tetradactylusProcellosaurinus erythrocercus
Vanzosaura rubricaudaMicrablepharus maximiliani
Micrablepharus atticolusTretioscincus oriximinensis
Tretioscincus agilisAcratosaura mentalisStenolepis ridleyi
Colobosaura modestaIphisa elegans
Colobodactylus taunayiColobodactylus dalcyanus
Heterodactylus imbricatusRhachisaurus brachylepis
Bachia heteropaBachia intermediaBachia barbouriBachia bicolorBachia peruana
Bachia panopliaBachia trisanale
Bachia scolecoidesBachia huallagana
Bachia dorbignyiBachia bresslaui
Bachia flavescensPetracola ventrimaculatus
Proctoporus bolivianusProctoporus unsaacae
Proctoporus guentheriProctoporus pachyurus
Proctoporus subsolanusProctoporus sucullucu
Cercosaura eigenmanniCercosaura schreibersii
Cercosaura ocellataCercosaura argulus
Cercosaura oshaughnessyiCercosaura quadrilineata
Potamites juruazensisPotamites ecpleopus
Pholidobolus macbrydeiPholidobolus montium
Placosoma glabellumPlacosoma cordylinum
Neusticurus bicarinatusNeusticurus rudis
Riama simoterusRiama colomaromani
Riama unicolorLeposoma guianense
Leposoma southiLeposoma parietale
Leposoma osvaldoiLeposoma percarinatum
Arthrosaura kockiiArthrosaura reticulata
Anotosaura collarisColobosauroides cearensis
Leposoma annectansLeposoma scincoides
Leposoma pukLeposoma baturitensis
Leposoma nanodactylusEcpleopus gaudichaudii
Alopoglossus angulatusAlopoglossus copii
Alopoglossus atriventrisPtychoglossus brevifrontalis
Cnemidophorus lacertoidesCnemidophorus longicaudus
Dicrodon guttulatumAmeiva fuscataAmeiva erythrocephala
Ameiva griswoldiAmeiva pluvianotata
Ameiva coraxAmeiva plei
Ameiva maynardiAmeiva lineolata
Ameiva taeniuraAmeiva chrysolaema
Ameiva leberiAmeiva wetmorei
Ameiva exsulAmeiva polops
Ameiva dorsalisAmeiva auberi
Aspidoscelis ceralbensisAspidoscelis hyperythra
Aspidoscelis tigrisAspidoscelis marmorata
Aspidoscelis lineattissimaAspidoscelis guttata
Aspidoscelis deppeiAspidoscelis velox
Aspidoscelis laredoensisAspidoscelis gularis
Aspidoscelis costataAspidoscelis communis
Aspidoscelis burtiAspidoscelis sexlineata
Aspidoscelis inornataCnemidophorus arenivagus
Cnemidophorus lemniscatusCnemidophorus gramivagus
Cnemidophorus vanzoiAmeiva festivaAmeiva quadrilineataAmeiva undulata
Kentropyx viridistrigaKentropyx vanzoi
Kentropyx paulensisKentropyx pelviceps
Kentropyx altamazonicaKentropyx striata
Kentropyx calcarataCnemidophorus ocellifer
Ameiva bifrontataAmeiva ameivaTeius teyou
Dracaena guianensisCrocodilurus amazonicus
Tupinambis teguixinTupinambis longilineus
Tupinambis quadrilineatusTupinambis duseni
Tupinambis merianaeTupinambis rufescens
Callopistes maculatusCallopistes flavipunctatus
100
98
100
84
99100
100
50
100
10087 100
93
8893
86
72
9861
100
100
98
100
91 96
100
93
10077
10088
9795 100
7874
98
100
99
80
100
97
55
98
94 99
100
98
99
82
100
92
100
98100
94
100
91
100
97
99
65100
95
96
86
100
10098
97
100100 99
54
100
56
93
98
92
99
59
100
94
8597
55
98
93
100
97
9695 100
100
86
100
99
84
98
100
98
9993 99
8183
100
99
94
97 100
82
100
9196
71
99
100
9798
8766
91
100
97
J
Teiidae
Gymnophthalmidae
Tupinambinae
Teiinae
Alopoglossinae
Ecpleopinae
Cercosaurinae
Bachiinae
Rhachisaurinae
Gymnophthalminae
Figure 11 Species-level squamate phylogeny continued (J).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 15 of 53http://www.biomedcentral.com/1471-2148/13/93

Amphisbaena xeraAmphisbaena bakeriAmphisbaena caeca
Amphisbaena fenestrata
Amphisbaena manni
Amphisbaena schmidti
Amphisbaena carlgansi
Amphisbaena cubana
Amphisbaena barbouri
Amphisbaena anaemariae
Amphisbaena silvestrii
Amphisbaena leeseri
Amphisbaena angustifrons
Amphisbaena darwini
Amphisbaena munoai
Amphisbaena kingiiAmphisbaena cuiabana
Amphisbaena hastata
Amphisbaena infraorbitale
Amphisbaena microcephalum
Amphisbaena polystegum
Amphisbaena anomala
Amphisbaena bolivica
Amphisbaena camura
Amphisbaena alba
Amphisbaena vermicularis
Amphisbaena roberti
Amphisbaena kraoh
Amphisbaena saxosa
Amphisbaena ignatiana
Amphisbaena leali
Amphisbaena innocens
Amphisbaena hyporissor
Amphisbaena mertensii
Amphisbaena cunhai
Amphisbaena fuliginosa
Amphisbaena brasiliana
Monopeltis capensis
Geocalamus acutus
Chirindia swynnertoni
Cynisca leucura
Trogonophis wiegmanni
Diplometopon zarudnyi
Cadea blanoides
Blanus tingitanus
Blanus mettetali
Blanus cinereus
Blanus strauchi
Bipes canaliculatus
Bipes biporus
Bipes tridactylus
Rhineura floridana
99
100
84
100
91
86
89
95
90
92
87
100
100
97
100
100
90
78100
93
99
100
100
100
81
100
98
95100
100
91
100
55
96
67
95
89
100
100
100
10097
10073
Blanidae
Cadeidae
Trogonophiidae
Amphisbaenidae
K
Bipedidae
Rhineuridae
Figure 12 Species-level squamate phylogeny continued (K).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 16 of 53http://www.biomedcentral.com/1471-2148/13/93

Darevskia derjuginiDarevskia caucasica
Darevskia daghestanicaDarevskia mixta
Darevskia clarkorumDarevskia armeniaca
Darevskia bendimahiensisDarevskia sapphirina
Darevskia uzzelliDarevskia raddei
Darevskia rostombekoviDarevskia saxicola
Darevskia brauneriDarevskia lindholmi
Darevskia alpinaDarevskia chlorogaster
Darevskia praticolaDarevskia valentini
Darevskia portschinskiiDarevskia rudis
Darevskia parvulaIranolacerta zagrosicaIranolacerta brandtii
Algyroides fitzingeriAlgyroides marchi
Dinarolacerta montenegrinaDinarolacerta mosorensis
Algyroides nigropunctatusAlgyroides moreoticus
Anatololacerta danfordiAnatololacerta oertzeni
Anatololacerta anatolicaParvilacerta parva
Parvilacerta fraasiiIberolacerta aranicaIberolacerta bonnali
Iberolacerta aurelioiIberolacerta horvathi
Iberolacerta monticolaIberolacerta cyreni
Iberolacerta galaniApathya cappadocica
Archaeolacerta bedriagaeHellenolacerta graecaDalmatolacerta oxycephala
Podarcis carbonelliPodarcis bocagei
Podarcis hispanicusPodarcis vaucheri
Podarcis liolepisPodarcis peloponnesiacusPodarcis erhardii
Podarcis raffoneaePodarcis lilfordi
Podarcis pityusensisPodarcis siculus
Podarcis muralisPodarcis tiliguerta
Podarcis filfolensisPodarcis gaigeae
Podarcis milensisPodarcis tauricus
Podarcis melisellensisScelarcis perspicillata
Teira dugesiiLacerta pamphylicaLacerta trilineata
Lacerta mediaLacerta agilis
Lacerta schreiberiLacerta strigata
Lacerta bilineataLacerta viridis
Timon tangitanusTimon lepidusTimon pater
Timon princepsTakydromus stejnegeriTakydromus septentrionalisTakydromus toyamaiTakydromus wolteriTakydromus formosanus
Takydromus hsuehshanensisTakydromus amurensis
Takydromus sylvaticusTakydromus dorsalis
Takydromus intermediusTakydromus sauteri
Takydromus smaragdinusTakydromus tachydromoides
Takydromus sexlineatusTakydromus kuehnei
Zootoca viviparaPhoenicolacerta cyanisparsa
Phoenicolacerta laevisPhoenicolacerta kulzeri
Acanthodactylus schreiberiAcanthodactylus boskianus
Acanthodactylus opheodurusAcanthodactylus gongrorhynchatus
Acanthodactylus masiraeAcanthodactylus cantoris
Acanthodactylus schmidtiAcanthodactylus longipes
Acanthodactylus scutellatusAcanthodactylus aureus
Acanthodactylus maculatusAcanthodactylus pardalis
Acanthodactylus beershebensisAcanthodactylus busacki
Acanthodactylus erythrurusAcanthodactylus blanci
Acanthodactylus tristramiAcanthodactylus orientalisOphisops elegans
Ophisops occidentalisMesalina rubropunctata
Mesalina brevirostrisMesalina adramitana
Mesalina balfouriMesalina guttulata
Mesalina bahaeldiniMesalina olivieri
Mesalina simoniOmanosaura jayakariOmanosaura cyanura
Congolacerta vauereselliEremias grammica
Eremias pleskeiEremias multiocellata
Eremias przewalskiiEremias arguta
Eremias nigrolateralisEremias persica
Eremias montanusEremias velox
Eremias vermiculataEremias argus
Eremias brenchleyiAdolfus jacksoni
Adolfus alleniAdolfus africanus
Gastropholis prasinaGastropholis vittata
Holaspis guentheriHolaspis laevis
Pedioplanis gaerdesiPedioplanis inornata
Pedioplanis rubensPedioplanis undata
Pedioplanis husabensisPedioplanis namaquensis
Pedioplanis brevicepsPedioplanis burchelli
Pedioplanis laticepsNucras tessellata
Pedioplanis lineoocellataMeroles cuneirostrisMeroles micropholidotus
Meroles ctenodactylusMeroles anchietaeIchnotropis squamulosa
Meroles suborbitalisMeroles knoxii
Meroles reticulatusIchnotropis capensis
Tropidosaura gularisAustralolacerta australis
Heliobolus spekiiHeliobolus lugubris
Pseuderemias smithiiPhilochortus spinalis
Latastia longicaudataNucras lalandii
Poromera fordiiAtlantolacerta andreanskyiGallotia bravoana
Gallotia simonyiGallotia intermedia
Gallotia caesarisGallotia galloti
Gallotia atlanticaGallotia stehlini
Psammodromus blanciPsammodromus algirus
Psammodromus hispanicus
100
100
100
58
94
86
64
73
53
97
100
96
93
9870
9990
94
100
100
97
100
10087
10094
99
100
9498
100
96
99100
89
100
100100
93
9186
92
97
92
62
100
78
88
9792 100
93
79
95
99
92
85
85
7455
100
99
100
9899
98
80100
9498 69
95
10095
79
100100
9975
99
97
85100
100
100
10098
92
96
93
73
100
99
10085
7599
80
96
9593
9594
10076
86
99
100100
10089
6393
91
9798
96
100
95
9356
86
100
9976
97
100
7299 89
100
100
99
97
100
96100
7696
56
69
100
54
100
76100 97
80
84
73
9699
8890
100
10099
9568
100 97
9995
Lacertidae
Gallotiinae
Lacertinae
L
Figure 13 Species-level squamate phylogeny continued (L).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 17 of 53http://www.biomedcentral.com/1471-2148/13/93

Varanus baritjiVaranus acanthurus
Varanus storriVaranus primordius
Varanus kingorumVaranus bushiVaranus gilleni
Varanus caudolineatusVaranus eremius
Varanus brevicaudaVaranus semiremex
Varanus mitchelliVaranus timorensis
Varanus scalarisVaranus glauerti
Varanus tristisVaranus pilbarensis
Varanus glebopalmaVaranus komodoensis
Varanus variusVaranus salvadoriiVaranus gouldiiVaranus panoptes
Varanus rosenbergiVaranus giganteus
Varanus spenceriVaranus mertensi
Varanus bengalensisVaranus flavescens
Varanus dumeriliiVaranus rudicollis
Varanus salvatorVaranus marmoratus
Varanus finschiVaranus doreanusVaranus yuwonoiVaranus jobiensis
Varanus caerulivirensVaranus cerambonensisVaranus melinusVaranus indicus
Varanus rainerguentheriVaranus prasinusVaranus macraeiVaranus boehmei
Varanus beccariiVaranus keithhornei
Varanus olivaceusVaranus yemenensis
Varanus albigularisVaranus exanthematicusVaranus niloticus
Varanus griseusLanthanotus borneensis
Shinisaurus crocodilurusAbronia auritaAbronia campbelliAbronia anzuetoiAbronia matudai
Abronia fimbriataAbronia lythrochilaAbronia ornelasi
Abronia frostiAbronia graminea
Abronia oaxacaeAbronia chiszari
Mesaspis gadoviiMesaspis moreletiiBarisia rudicollis
Barisia herreraeBarisia imbricataBarisia levicollis
Abronia mixtecaGerrhonotus infernalisGerrhonotus liocephalusColoptychon rhombifer
Gerrhonotus parvusElgaria multicarinataElgaria panamintinaElgaria paucicarinataElgaria kingii
Elgaria coeruleaDopasia harti
Dopasia gracilisOphisaurus attenuatus
Anguis fragilisPseudopus apodus
Ophisaurus koellikeriOphisaurus ventralis
Celestus haetianusCelestus agasepsoides
Ophiodes striatusDiploglossus pleii
Diploglossus bilobatusCelestus enneagrammus
Anniella geronimensisAnniella pulchra
Heloderma horridumHeloderma suspectum
Xenosaurus grandisXenosaurus platyceps
100
94
100
100
100
84
100
100
88
97
99100
99
99
100100
100
99
8796
7686
100
100
100
99
100
9296
94
100
8572
86
69
95
86
100
767974
8678
88
10095
75
98
9597
99
100
95
100
90
100
100
97
89
79
9689
9697
95
84
7266
99
100
9594
88
92
100
998584
100
87
100
83
99
98
93 100100
100
100
100
M
XenosauridaeHelodermatidae
Anniellidae
Anguidae
Shinisauridae Lanthanotidae
Varanidae
Gerrhonotinae
Anguinae
Diploglossinae
Figure 14 Species-level squamate phylogeny continued (M).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 18 of 53http://www.biomedcentral.com/1471-2148/13/93

Trioceros sternfeldiTrioceros rudis
Trioceros elliotiTrioceros hoehneliiTrioceros schubotzi
Trioceros narraiocaTrioceros jacksonii
Trioceros bitaeniatusTrioceros fuelleborni
Trioceros goetzeiTrioceros tempeli
Trioceros werneriTrioceros deremensis
Trioceros melleriTrioceros johnstoni
Trioceros oweniTrioceros pfefferiTrioceros quadricornisTrioceros wiedersheimi
Trioceros cristatusTrioceros montiumTrioceros feae
Trioceros balebicornutusTrioceros harennae
Trioceros affinisChamaeleo arabicus
Chamaeleo calyptratusChamaeleo zeylanicus
Chamaeleo chamaeleonChamaeleo calcaricarensChamaeleo africanus
Chamaeleo monachusChamaeleo senegalensis
Chamaeleo roperiChamaeleo dilepis
Chamaeleo quilensisChamaeleo gracilis
Chamaeleo necasiChamaeleo laevigatus
Chamaeleo namaquensisCalumma globifer
Calumma parsoniiCalumma oshaughnessyi
Calumma capuroniRhampholeon chapmanorum
Rhampholeon platycepsRhampholeon nchisiensis
Rhampholeon beraducciiRhampholeon boulengeriRhampholeon acuminatus
Rhampholeon uluguruensisRhampholeon moyeri
Rhampholeon marshalliRhampholeon viridis
Rhampholeon spinosusRhampholeon temporalis
Rhampholeon spectrumNadzikambia mlanjensis
Calumma nasutumCalumma boettgeri
Calumma fallaxCalumma gallus
Calumma maltheCalumma guibei
Calumma hilleniusiCalumma tsaratananense
Calumma brevicorneCalumma crypticum
Calumma cucullatumRieppeleon brevicaudatus
Rieppeleon brachyurusRieppeleon kerstenii
Calumma gastrotaeniaCalumma furcifer
Bradypodion ventraleBradypodion karrooicumBradypodion taeniabronchum
Bradypodion gutturaleBradypodion atromontanum
Bradypodion occidentaleBradypodion setaroi
Bradypodion nemoraleBradypodion dracomontanumBradypodion transvaalense
Bradypodion thamnobatesBradypodion melanocephalum
Bradypodion cafferBradypodion damaranumBradypodion pumilum
Kinyongia fischeriKinyongia vosseleri
Kinyongia multituberculataKinyongia matschiei
Kinyongia boehmeiKinyongia tavetana
Kinyongia uthmoelleriKinyongia carpenteri
Kinyongia xenorhinaKinyongia excubitor
Kinyongia adolfifridericiKinyongia tenue
Kinyongia oxyrhinaFurcifer antimenaFurcifer labordi
Furcifer lateralisFurcifer belalandaensis
Furcifer verrucosusFurcifer oustaleti
Furcifer angeliFurcifer pardalis
Furcifer polleniFurcifer cephalolepis
Furcifer campaniFurcifer petteri
Furcifer willsiiFurcifer minor
Furcifer bifidusFurcifer balteatus
Brookesia lineataBrookesia betschiBrookesia thieliBrookesia vadoni
Brookesia ebenauiBrookesia antakaranaBrookesia ambreensis
Brookesia stumpffiBrookesia griveaudi
Brookesia valerieaeBrookesia superciliaris
Brookesia therezieniBrookesia bonsiBrookesia brygooi
Brookesia decaryiBrookesia perarmata
Brookesia peyrierasiBrookesia karchei
Brookesia tuberculataBrookesia minima
Brookesia exarmataBrookesia dentata
Brookesia nasusBrookesia lolontany
100
100
58
85
100
98
9297
7098
8587
9766
90
91
92
97
93
66
10096
50
10085
100
100
95
90
100
10098
100
95
99
99
100
100
10061
82
78
88
99
86100
7391
90100
81
73
9898
100
100
9193
8597
94
75
88
98
99
100
100
55
100
98
85987495
95
94
7287
100
96
95
82
92100
10059
100
100
100
6798
100
9999
93
94
99
9898
10094
93
91100
96
62
86
100
99
100
51
100
100
93
100
100
100 98
100
83 100
63
10079
ChamaeleonidaeBrookesiinae
Chamaeleoninae
N
Figure 15 Species-level squamate phylogeny continued (N).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 19 of 53http://www.biomedcentral.com/1471-2148/13/93

Pseudocalotes flavigulaPseudocalotes brevipes
Pseudocalotes kakhienensisJapalura splendida
Japalura flavicepsOtocryptis wiegmanniSitana ponticeriana
Acanthosaura lepidogasterAcanthosaura capraAcanthosaura armataAcanthosaura crucigera
Salea horsfieldiiCalotes htunwiniCalotes calotes
Calotes irawadiCalotes versicolor
Calotes liolepisCalotes nigrilabris
Calotes ceylonensisCalotes liocephalusCalotes emma
Calotes chincolliumCalotes mystaceus
Ceratophora erdeleniCeratophora stoddartii
Ceratophora karuCeratophora aspera
Cophotis ceylanicaCophotis dumbara
Lyriocephalus scutatusGonocephalus kuhliiGonocephalus chamaeleontinus
Gonocephalus grandisBronchocela cristatella
Coryphophylax subcristatusAphaniotis fusca
Gonocephalus robinsoniiJapalura polygonata
Phoxophrys nigrilabrisDraco ornatusDraco palawanensisDraco spilopterus
Draco cornutusDraco guentheri
Draco quadrasiDraco cyanopterusDraco reticulatusDraco boschmai
Draco timorensisDraco volans
Draco bimaculatusDraco rhytismaDraco bourouniensis
Draco beccariiDraco biaroDraco caerulhians
Draco spilonotusDraco obscurusDraco taeniopterus
Draco blanfordiiDraco indochinensisDraco haematopogon
Draco melanopogonDraco quinquefasciatus
Draco mindanensisDraco maximus
Draco cristatellusDraco fimbriatus
Draco maculatusDraco lineatus
Draco dussumieriPtyctolaemus collicristatus
Ptyctolaemus gularisJapalura tricarinata
Japalura variegataMantheyus phuwuanensis
Agama paragamaAgama planiceps
Agama finchiAgama agama
Agama doriaeAgama sankaranicaAgama mwanzae
Agama kaimosaeAgama lionotus
Agama rueppelliAgama caudospinosa
Agama aculeataAgama armata
Agama hispidaAgama atraAgama anchietae
Agama weidholziAgama gracilimembris
Agama insularisAgama boulengeri
Agama castroviejoiAgama impalearis
Agama bouetiAgama spinosa
Trapelus savigniiTrapelus flavimaculatus
Trapelus pallidusTrapelus ruderatus
Trapelus agilisTrapelus mutabilisTrapelus sanguinolentus
Bufoniceps laungwalaensisXenagama taylori
Acanthocercus atricollisPseudotrapelus sinaitus
Phrynocephalus guttatusPhrynocephalus albolineatus
Phrynocephalus melanurusPhrynocephalus przewalskiiPhrynocephalus versicolorPhrynocephalus raddei
Phrynocephalus mystaceusPhrynocephalus helioscopus
Phrynocephalus axillarisPhrynocephalus lidskiiPhrynocephalus vlangalii
Phrynocephalus putjataiPhrynocephalus theobaldiPhrynocephalus forsythiiPhrynocephalus interscapularis
Phrynocephalus scutellatusLaudakia erythrogasterLaudakia caucasia
Laudakia microlepisLaudakia stoliczkana
Laudakia himalayanaLaudakia lehmanni
Laudakia nuptaLaudakia tuberculataLaudakia sacra
Laudakia stellio
100
100
100
52
81
50
96
69
99100
95
100
100
100
100
99
91
100
90
99
55
72100
86
100
90100
97
100
97
8587
100
98
100
100
86
92
86
95
100
85
87
91
100
100
99
100
99
100
97
99
100
59
9661
100
6595
53
95
97
70100
100
100
100
91
98
100100
6868
99
100
9165
100
50100
100
76
8469
100100
96
66
10099
9989
100
93
100
95
84
10064
918994100
9690
100
100
907573
94
998488
100
85
99
Diporiphora lalliaeDiporiphora arnhemica
Diporiphora albilabrisDiporiphora bennettii
Diporiphora nobbiDiporiphora australis
Diporiphora amphiboluroidesDiporiphora valensDiporiphora pindanDiporiphora bilineataDiporiphora magnaDiporiphora reginae
Diporiphora winneckeiDiporiphora linga
Diporiphora superbaPogona vitticepsPogona minorPogona minimaPogona henrylawsoni
Pogona nullarborPogona barbata
Rankinia diemensisTympanocryptis lineataTympanocryptis pinguicollaTympanocryptis cephalusTympanocryptis tetraporophoraTympanocryptis intima
Tympanocryptis uniformisAmphibolurus norrisiAmphibolurus muricatus
Lophognathus gilbertiChlamydosaurus kingii
Lophognathus longirostrisLophognathus temporalis
Ctenophorus femoralisCtenophorus fordiCtenophorus maculatusCtenophorus pictusCtenophorus mckenzieiCtenophorus scutulatus
Ctenophorus isolepisCtenophorus ornatus
Ctenophorus caudicinctusCtenophorus fionni
Ctenophorus decresiiCtenophorus vadnappaCtenophorus tjantjalka
Ctenophorus rufescensCtenophorus reticulatusCtenophorus nuchalis
Ctenophorus cristatusCtenophorus salinarum
Ctenophorus gibbaCtenophorus clayi
Ctenophorus adelaidensisCtenophorus maculosus
Intellagama lesueuriiHypsilurus bruijniiHypsilurus nigrigularis
Hypsilurus papuensisHypsilurus modestus
Chelosania brunneaMoloch horridus
Hypsilurus dilophusHypsilurus boydii
Hypsilurus spinipesPhysignathus cocincinus
Hydrosaurus amboinensisLeiolepis guentherpetersiLeiolepis guttataLeiolepis reevesiiLeiolepis belliana
Uromastyx yemenensisUromastyx bentiUromastyx ocellata
Uromastyx ornataUromastyx leptieniUromastyx aegyptia
Uromastyx thomasiUromastyx disparUromastyx acanthinuraUromastyx geyri
Uromastyx macfadyeniUromastyx princepsUromastyx asmussi
Uromastyx loricataUromastyx hardwickii
100
96
100
100
100
71
100
100
71
99
95
95
56100
95
81
67
100
76
99
63
99
9994
100
100
10053
99
100
94
97
71
100
100
79
53
76
90100
94
92100
100
99
99
90
98
96
75
85
100
100 51
98
92
97
100 100
100
100
86
74
91
70
100100
99
100
100
94
94
95
Ag
amid
ae
Uromastycinae
LeiolepidinaeA
mph
ibol
urin
ae
Hydrosaurinae Aga
min
ae
Draconinae
O
(i)
(ii)
(i)
(ii)
Figure 16 Species-level squamate phylogeny continued (O).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 20 of 53http://www.biomedcentral.com/1471-2148/13/93

Ctenosaura oaxacanaCtenosaura quinquecarinataCtenosaura flavidorsalisCtenosaura palearis
Ctenosaura bakeriCtenosaura oedirhinaCtenosaura melanosterna
Ctenosaura pectinataCtenosaura acanthuraCtenosaura hemilophaCtenosaura similis
Conolophus pallidusConolophus subcristatusAmblyrhynchus cristatus
Cyclura cychluraCyclura nubila
Cyclura rileyiCyclura colleiCyclura carinataCyclura ricordiCyclura cornuta
Cyclura pinguisSauromalus variusSauromalus hispidusSauromalus klauberiSauromalus aterIguana delicatissima
Iguana iguanaBrachylophus vitiensisBrachylophus fasciatus
Dipsosaurus dorsalisTropidurus torquatusTropidurus hispidus
Tropidurus oreadicusTropidurus insulanus
Tropidurus erythrocephalusTropidurus mucujensis
Tropidurus etheridgeiTropidurus cocorobensis
Tropidurus itambereTropidurus psammonastes
Tropidurus montanusTropidurus hygomi
Eurolophosaurus nanuzaeEurolophosaurus amathites
Eurolophosaurus divaricatusTropidurus spinulosus
Tropidurus callathelysPlica plica
Plica lumariaPlica umbra
Strobilurus torquatusTropidurus bogerti
Uracentron flavicepsUranoscodon superciliosus
Microlophus duncanensisMicrolophus albemarlensisMicrolophus pacificusMicrolophus grayii
Microlophus delanonisMicrolophus stolzmanni
Microlophus habeliiMicrolophus bivittatus
Microlophus occipitalisMicrolophus koepckeorum
Microlophus yaneziMicrolophus theresioidesMicrolophus atacamensis
Microlophus quadrivittatusMicrolophus peruvianus
Microlophus tigrisMicrolophus heterolepis
Microlophus theresiaeMicrolophus thoracicus
Stenocercus stigmosusStenocercus melanopygus
Stenocercus latebrosusStenocercus orientalis
Stenocercus ornatissimusStenocercus chrysopygus
Stenocercus cupreusStenocercus boettgeriStenocercus empetrusStenocercus eunetopsisStenocercus imitator
Stenocercus variusStenocercus torquatus
Stenocercus crassicaudatusStenocercus marmoratus
Stenocercus humeralisStenocercus angel
Stenocercus guentheriStenocercus festae
Stenocercus chotaStenocercus rhodomelasStenocercus anguliferStenocercus iridescens
Stenocercus puyangoStenocercus percultusStenocercus ornatus
Stenocercus caducusStenocercus limitaris
Stenocercus doellojuradoiStenocercus azureusStenocercus roseiventris
Stenocercus ochoaiStenocercus formosus
Stenocercus apurimacusStenocercus scapularis
100
72
100
100
56
100
99
100
9972
74
98
629699
100
100
9557
8586
94
97
92
100
61
99100
100
100
100
10073
97
10098
89
83
98
100
100100
100
98
78
99
74
100
100
100100
6495
100100
100
99100
100
98
10096
100
78
84
100
73
88
79
10085
88
78100
8994
71100
88
96
8299
96100
89100
100
69
64100
86
94
94
Iguanidae
Tropiduridae
PQ-R
Figure 17 Species-level squamate phylogeny continued (P).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 21 of 53http://www.biomedcentral.com/1471-2148/13/93

Sceloporus adleriSceloporus stejnegeriSceloporus formosusSceloporus subpictusSceloporus cryptus
Sceloporus lundelliSceloporus malachiticusSceloporus smaragdinus
Sceloporus taeniocnemisSceloporus spinosusSceloporus edwardtaylori
Sceloporus horridusSceloporus cautusSceloporus undulatus
Sceloporus consobrinusSceloporus woodiSceloporus virgatus
Sceloporus occidentalisSceloporus olivaceus
Sceloporus clarkiiSceloporus cyanogenysSceloporus serriferSceloporus ornatus
Sceloporus minorSceloporus dugesii
Sceloporus poinsettiiSceloporus macdougalli
Sceloporus mucronatusSceloporus bulleri
Sceloporus torquatusSceloporus jarrovii
Sceloporus insignisSceloporus megalepidurus
Sceloporus goldmaniSceloporus samcolemani
Sceloporus chaneyiSceloporus slevini
Sceloporus scalarisSceloporus bicanthalis
Sceloporus aeneusSceloporus heterolepis
Sceloporus palaciosiSceloporus grammicus
Sceloporus melanorhinusSceloporus hunsakeriSceloporus orcuttiSceloporus lickiSceloporus lineatulus
Sceloporus zosteromusSceloporus magister
Sceloporus arenicolusSceloporus graciosusSceloporus vandenburgianus
Sceloporus ochoterenaeSceloporus jalapae
Sceloporus maculosusSceloporus gadoviae
Sceloporus nelsoniSceloporus pyrocephalus
Sceloporus merriamiSceloporus siniferus
Sceloporus squamosusSceloporus carinatus
Sceloporus utiformisSceloporus angustus
Sceloporus grandaevusSceloporus cozumelae
Sceloporus smithiSceloporus variabilis
Sceloporus teapensisSceloporus chrysostictus
Sceloporus couchiiSceloporus parvus
Urosaurus auriculatusUrosaurus clarionensisUrosaurus ornatus
Urosaurus graciosusUrosaurus nigricaudusUrosaurus lahtelaiUrosaurus bicarinatus
Urosaurus gadoviUta stejnegeri
Uta stansburianaUta palmeriUta squamataPetrosaurus thalassinusPetrosaurus repens
Petrosaurus mearnsiPhrynosoma hernandesi
Phrynosoma douglassiiPhrynosoma ditmarsiPhrynosoma orbiculare
Phrynosoma modestumPhrynosoma taurusPhrynosoma braconnieri
Phrynosoma asioPhrynosoma wigginsiPhrynosoma cerroense
Phrynosoma blainvilliiPhrynosoma coronatum
Phrynosoma platyrhinosPhrynosoma mcalliiPhrynosoma solare
Phrynosoma cornutumHolbrookia maculataHolbrookia propinquaHolbrookia lacerata
Cophosaurus texanusCallisaurus draconoides
Uma notataUma inornataUma scoparia
Uma paraphygasUma exsul
Crotaphytus collarisCrotaphytus nebriusCrotaphytus bicinctores
Crotaphytus grismeriCrotaphytus insularis
Crotaphytus vestigiumCrotaphytus reticulatus
Crotaphytus antiquusGambelia copeiiGambelia sila
Gambelia wislizeniiLeiocephalus barahonensisLeiocephalus schreibersii
Leiocephalus personatusLeiocephalus psammodromus
Leiocephalus ravicepsLeiocephalus carinatus
54
81
87
100
100
86
90
75
100
96
89
100
99
96
82
99
100
100
61100
97
100
98100
100
74
10065
100
91
89
100
989898100
100
96
92
97
89
100
9898
100
99
99
100
97
100
10099
9999
10099
82
100
100
95
100
100100
100
100
100
9197
96
100
10099
100100
99100
10076
100
100100
100
100
95
10097
10099
8892
96
100
96
100
100
99
99100
100
100100
100
100
100
83
93100
100
82
99
100
100
92
93
94
Lei
oce
ph
alid
ae Crotaphytidae
Ph
ryn
oso
mat
idae
Liolaemus lavillaiLiolaemus ornatusLiolaemus calchaquiLiolaemus albicepsLiolaemus irregularis
Liolaemus crepuscularisLiolaemus abaucanLiolaemus espinozai
Liolaemus quilmesLiolaemus koslowskyi
Liolaemus laurentiLiolaemus darwiniiLiolaemus grosseorum
Liolaemus olongastaLiolaemus chacoensis
Liolaemus uspallatensisLiolaemus hermannuneziLiolaemus xanthoviridisLiolaemus fitzingerii
Liolaemus chehuachekenkLiolaemus morenoiLiolaemus canqueli
Liolaemus melanopsLiolaemus donosobarrosiLiolaemus cuyanus
Liolaemus telsenLiolaemus inacayali
Liolaemus boulengeriLiolaemus rothi
Liolaemus wiegmanniiLiolaemus azarai
Liolaemus scapularisLiolaemus multimaculatusLiolaemus riojanusLiolaemus salinicola
Liolaemus lutzaeLiolaemus occipitalis
Liolaemus pseudoanomalusLiolaemus andinus
Liolaemus multicolorLiolaemus molinaiLiolaemus dorbignyiLiolaemus huacahuasicusLiolaemus fabianiLiolaemus audituvelatus
Liolaemus ruibaliLiolaemus vallecurensisLiolaemus famatinae
Liolaemus orientalisLiolaemus stolzmanni
Liolaemus archeforusLiolaemus tristisLiolaemus zullyaeLiolaemus scolaroi
Liolaemus sarmientoiLiolaemus gallardoi
Liolaemus kingiiLiolaemus escarchadosiLiolaemus tari
Liolaemus bagualiLiolaemus somuncurae
Liolaemus uptoniLiolaemus magellanicus
Liolaemus kolenghLiolaemus silvanaeLiolaemus lineomaculatus
Liolaemus hatcheriLiolaemus saxatilis
Liolaemus gracilisLiolaemus robertmertensiLiolaemus ramirezae
Liolaemus yanalcuLiolaemus walkeriLiolaemus puna
Liolaemus chaltinLiolaemus bitaeniatus
Liolaemus pagaburoiLiolaemus bibronii
Liolaemus hernaniLiolaemus pictusLiolaemus cyanogaster
Liolaemus chiliensisLiolaemus schroederi
Liolaemus gravenhorstiiLiolaemus belliiLiolaemus coeruleusLiolaemus elongatus
Liolaemus kriegiLiolaemus leopardinus
Liolaemus ceiiLiolaemus buergeriLiolaemus petrophilusLiolaemus umbriferLiolaemus capillitasLiolaemus heliodermis
Liolaemus dicktracyiLiolaemus thermarum
Liolaemus austromendocinusLiolaemus paulinaeLiolaemus plateiLiolaemus nigromaculatus
Liolaemus pseudolemniscatusLiolaemus zapallarensisLiolaemus atacamensis
Liolaemus fuscusLiolaemus nigroviridisLiolaemus nitidus
Liolaemus monticolaLiolaemus lemniscatus
Liolaemus tenuisPhymaturus antofagastensisPhymaturus mallimacciiPhymaturus punaePhymaturus palluma
Phymaturus dorsimaculatusPhymaturus patagonicusPhymaturus somuncurensis
Phymaturus indistinctusCtenoblepharys adspersa
Leiosaurus catamarcensisLeiosaurus paronae
Leiosaurus belliiPristidactylus torquatus
Diplolaemus darwiniiPristidactylus scapulatus
Urostrophus vautieriAnisolepis longicauda
Urostrophus gallardoiEnyalius leechii
Enyalius bilineatusOplurus fierinensis
Oplurus grandidieriOplurus saxicola
Oplurus quadrimaculatusOplurus cuvieri
Oplurus cyclurusChalarodon madagascariensis
Enyalioides praestabilisEnyalioides microlepisEnyalioides palpebralis
Enyalioides oshaughnessyiEnyalioides heterolepis
Enyalioides laticepsMorunasaurus annularis
Hoplocercus spinosusPolychrus marmoratus
Polychrus femoralisPolychrus acutirostris
Polychrus gutturosus
63
95
96
95
100
100
100
99
78
100
66
996786
100
94100
94100
9599100
10098649579
93
99
100
85
97
85
9998100
65100
99
100
82
72
8487
100
9882
78 99
100
100
100
100
999878
83
10095
99
9470
100
99
100
99
86
100
97
7484
100
91
5653
10068
100
9899
9897
100
59100
6698
100
100
9883
99
95
916766 97
100
99
68 93
100
100
100
968988
10098
99
100100
7877
70
98
97
99
100
7396 88
9099
99100
88
9696
96
100
100
92
69
Po
lych
roti
dae
Ho
plo
cerc
idae Opluridae
Lei
osa
uri
dae
Lio
laem
idae
Leiosaurinae
Enyaliinae
(i)
(ii)
(i)(ii)
QR
Figure 18 Species-level squamate phylogeny continued (Q).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 22 of 53http://www.biomedcentral.com/1471-2148/13/93

Anolis smaragdinusAnolis allisoniAnolis maynardi
Anolis brunneusAnolis longiceps
Anolis porcatusAnolis carolinensisAnolis isolepisAnolis oporinus
Anolis argillaceusAnolis centralis
Anolis pumilusAnolis loysiana
Anolis guazumaAnolis garridoi
Anolis angusticepsAnolis paternusAnolis alayoniAnolis sheplaniAnolis placidus
Anolis alutaceusAnolis inexpectatus
Anolis vanidicusAnolis alfaroi
Anolis macilentusAnolis cupeyalensisAnolis cyanopleurusAnolis rejectus
Anolis clivicolaAnolis lucius
Anolis whitemaniAnolis cybotes
Anolis armouriAnolis shreveiAnolis haetianusAnolis longitibialis
Anolis strahmiAnolis marcanoi
Anolis barahonaeAnolis baleatus
Anolis ricordiAnolis eugenegrahami
Anolis christopheiAnolis cuvieri
Anolis guamuhayaAnolis chamaeleonides
Anolis porcusAnolis barbatus
Anolis argenteolusAnolis alumina
Anolis semilineatusAnolis olssoni
Anolis barbouriAnolis insolitus
Anolis fowleriAnolis etheridgei
Anolis equestrisAnolis luteogularis
Anolis baracoaeAnolis nobleiAnolis smallwoodi
Anolis alinigerAnolis singularisAnolis chlorocyanusAnolis coelestinus
Anolis hendersoniAnolis dolichocephalus
Anolis bahorucoensisAnolis darlingtoni
Anolis monticolaAnolis occultus
Anolis vermiculatusAnolis bartschi
Anolis maculigulaAnolis casildae
Anolis frenatusAnolis princeps
Anolis chocorumAnolis fraseri
Anolis danieliAnolis insignisAnolis microtus
Anolis agassiziAnolis neblininusAnolis calimae
Anolis anatolorosAnolis jacare
Anolis tigrinusAnolis transversalisAnolis punctatus
Anolis inderenaeAnolis vanzoliniiAnolis heterodermusAnolis nicefori
Anolis euskalerriariAnolis gemmosusAnolis aequatorialisAnolis ventrimaculatusAnolis chlorisAnolis festae
Anolis peraccaeAnolis huilaeAnolis fitchi
Anolis extremusAnolis roquet
Anolis aeneusAnolis richardii
Anolis trinitatisAnolis griseus
Anolis bonairensisAnolis luciae
Anolis boettgeriBasiliscus basiliscus
Basiliscus plumifronsBasiliscus vittatus
Basiliscus galeritusCorytophanes cristatus
Corytophanes percarinatusLaemanctus longipes
99
100
100
93
60
80
86
98
95
99
10074
97
100
100
89
97
100
100
85
100 100
100
100
6576
100
99
97
100
100
9699
95
99
99
95
86
9090
10095
100
8666
9786
9799
9898
99
59
100
5989
90
94
10099
757982
56100
100
100
100
90
83
71
100
87
100100
80
98
100
98
100
99100
100
100100
9965
89
9599
96
96
67
64
10062
96100
100
100
9892
100
Anolis trachydermaAnolis poecilopusAnolis tropidogaster
Anolis lionotusAnolis oxylophus
Anolis zeusAnolis limifrons
Anolis lemurinusAnolis bicaorum
Anolis carpenteriAnolis ocelloscapularisAnolis tropidonotus
Anolis capitoAnolis polylepis
Anolis cupreusAnolis fuscoauratusAnolis kemptoniAnolis altae
Anolis humilisAnolis pachypus
Anolis sericeusAnolis isthmicus
Anolis intermediusAnolis laeviventris
Anolis ortoniiAnolis quercorum
Anolis aquaticusAnolis woodiAnolis biporcatus
Anolis bitectusAnolis uniformisAnolis polyrhachis
Anolis sminthusAnolis crassulusAnolis utilensisAnolis purpurgularis
Anolis loveridgeiAnolis chrysolepis
Anolis bombicepsAnolis nitens
Anolis meridionalisAnolis lineatus
Anolis oncaAnolis annectens
Anolis auratusAnolis grahamiAnolis conspersus
Anolis garmaniAnolis opalinusAnolis valencienni
Anolis reconditusAnolis lineatopus
Anolis bremeriAnolis quadriocellifer
Anolis sagreiAnolis ophiolepis
Anolis mestreiAnolis homolechisAnolis jubar
Anolis confususAnolis guafe
Anolis ahliAnolis allogus
Anolis rubribarbarisAnolis imias
Anolis cristatellusAnolis desechensis
Anolis ernestwilliamsiAnolis scriptus
Anolis cookiAnolis monensis
Anolis krugiAnolis pulchellus
Anolis gundlachiAnolis poncensis
Anolis evermanniAnolis stratulusAnolis acutus
Anolis caudalisAnolis marron
Anolis brevirostrisAnolis websteriAnolis distichus
Anolis marmoratusAnolis sabanusAnolis nubilus
Anolis lividusAnolis ferreus
Anolis terraealtaeAnolis oculatus
Anolis gingivinusAnolis bimaculatus
Anolis leachiiAnolis pogus
Anolis wattsi
75
100
99
92
100
93
90
100
56
94
97
9692
86
91
100
100
99
98
69
100 86
86
93100
100
90
72
95100
94
96
100
95
99
5284
5999
100
58
100
90
100
100
90
100
10099
8999
100
7294
98
10098
100
95
100
98
100
96100
100
100
91
100
88
100
100
9694
100
100
99
99
1009557
77
56
90
100
Corytophanidae
Dactyloidae
(i)
(ii)
(i)
(ii)
R
Figure 19 Species-level squamate phylogeny continued (R).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 23 of 53http://www.biomedcentral.com/1471-2148/13/93

Austrotyphlops pilbarensisAustrotyphlops hamatus
Austrotyphlops endoterusAustrotyphlops australisAustrotyphlops waitiiAustrotyphlops splendidusAustrotyphlops pinguisRamphotyphlops bicolor
Austrotyphlops ligatusAustrotyphlops ganei
Austrotyphlops troglodytesAustrotyphlops kimberleyensis
Austrotyphlops ammodytesAustrotyphlops unguirostris
Austrotyphlops leptosomusAustrotyphlops longissimus
Austrotyphlops grypusAustrotyphlops bituberculatus
Austrotyphlops guentheriAustrotyphlops howi
Austrotyphlops diversusRamphotyphlops polygrammicusAcutotyphlops kunuaensis
Acutotyphlops subocularisRamphotyphlops acuticaudus
Ramphotyphlops lineatusTyphlopidae sp. (Sri Lanka)
Ramphotyphlops braminusTyphlops pammeces
Ramphotyphlops albicepsTyphlops ruber
Typhlops luzonensisTyphlops arenarius
Typhlops vermicularisAfrotyphlops punctatus
Afrotyphlops congestusAfrotyphlops lineolatus
Afrotyphlops bibroniiAfrotyphlops fornasinii
Megatyphlops schlegeliiAfrotyphlops angolensis
Typhlops elegansLetheobia obtusa
Letheobia newtoniLetheobia feae
Rhinotyphlops lalandeiTyphlops catapontusTyphlops richardiTyphlops platycephalusTyphlops hypomethesTyphlops granti
Typhlops dominicanusTyphlops monastusTyphlops notorachiusTyphlops anousiusTyphlops anchaurusTyphlops contorhinusTyphlops aratorTyphlops caymanensis
Typhlops agoralionisTyphlops sylleptor
Typhlops jamaicensisTyphlops capitulatus
Typhlops rostellatusTyphlops sulcatus
Typhlops titanopsTyphlops schwartzi
Typhlops lumbricalisTyphlops eperopeus
Typhlops syntherusTyphlops hectusTyphlops pusillus
Typhlops biminiensisTyphlops reticulatus
Typhlops brongersmianusXenotyphlops grandidieri
Gerrhopilus mirusGerrhopilus hedraeusLeptotyphlops scutifrons
Leptotyphlops distantiLeptotyphlops conjunctusLeptotyphlops sylvicolus
Leptotyphlops nigricansLeptotyphlops nigroterminus
Namibiana occidentalisMyriopholis adleri
Myriopholis macrorhynchaMyriopholis blanfordi
Myriopholis bouetiMyriopholis rouxestevae
Myriopholis algeriensisMyriopholis longicauda
Mitophis leptipileptusMitophis pyrites
Mitophis asbolepisTetracheilostoma breuili
Tetracheilostoma carlaeRena humilisRena dulcis
Trilepida macrolepisEpictia columbiEpictia goudotii
Epictia albifronsSiagonodon septemstriatus
Tricheilostoma bicolorTyphlophis squamosus
Liotyphlops albirostris
100
90
100
83
100
87
76
99
72
91
100
98
76
99
988174
91
96
90
92
93
87
96
59
75
100 52
74
87
100
65
84100
97
100
87
100
77
99
879972
89
100
100
100
90
92
92
86
84999575
100
100
74
10077
7792
99
71878456
95
96
91
100
100
100100
10051
100
99
100
56
7969 100
100
97
97
100 100
98
100
100
100
93
100100
100 Anomalepididae
Leptotyphlopidae
GerrhopilidaeXenotyphlopidae
Typ
hlo
pid
ae
ST-AA
Figure 20 Species-level squamate phylogeny continued (S).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 24 of 53http://www.biomedcentral.com/1471-2148/13/93

Antaresia childreniAntaresia stimsoniAntaresia perthensisAntaresia maculosaMorelia bredliMorelia spilota
Morelia carinataMorelia viridisBothrochilus boaLeiopython albertisii
Liasis fuscusLiasis mackloti
Apodora papuanaLiasis olivaceus
Aspidites melanocephalusAspidites ramsayi
Morelia amethistinaMorelia boeleniMorelia oenpelliensisBroghammerus timoriensisBroghammerus reticulatus
Python molurusPython sebaePython curtusPython regius
Python brongersmaiLoxocemus bicolor
Xenopeltis unicolorRhinophis blythiiRhinophis homolepisPseudotyphlops philippinusRhinophis oxyrhynchus
Rhinophis philippinusRhinophis dorsimaculatus
Rhinophis drummondhayiUropeltis phillipsiUropeltis melanogaster
Rhinophis travancoricusUropeltis ceylanicus
Uropeltis liuraBrachyophidium rhodogaster
Melanophidium punctatumCylindrophis ruffus
Cylindrophis maculatusAnomochilus leonardi
Epicrates striatusEpicrates exsulEpicrates chrysogasterEpicrates subflavus
Epicrates fordiEpicrates monensisEpicrates inornatus
Epicrates anguliferEunectes notaeusEunectes murinus
Epicrates cenchriaCorallus annulatus
Corallus caninusCorallus hortulanusBoa constrictorEryx tataricusEryx miliaris
Eryx elegansEryx jaculusEryx johnii
Eryx conicusEryx colubrinus
Eryx jayakariCandoia bibroni
Candoia carinataCandoia aspera
Lichanura trivirgataCharina bottae
Ungaliophis continentalisExiliboa placata
Calabaria reinhardtiiAcrantophis dumeriliAcrantophis madagascariensisSanzinia madagascariensisCasarea dussumieriXenophidion schaeferiTropidophis melanurusTropidophis feicki
Tropidophis haetianusTropidophis greenwayi
Tropidophis wrightiTropidophis pardalis
Trachyboa boulengeriTrachyboa gularis
Anilius scytale
100
97
69
88
89
98
100
100
100
96
71
96
85
5183100
84100
100
75
98
100
100
7697
98
99
8696
99
100
9886
9073
80
69
77
848580
82
100
65
100
99
87
83
98
100
96
90
95100
92
9485
100
99
100
95
9693100
87
53
100
81
99100
100
100
100
98
100
100
5398
100
100
100 Aniliidae
Tropidophiidae
XenophidiidaeBolyeriidae
Boidae
AnomochilidaeCylindrophiidae
Uropeltidae
XenopeltidaeLoxocemidae
Pythonidae
CalabariidaeSanziniinae
Ungaliophiinae
Candoiinae
Erycinae
Boinae
TU-AA
Figure 21 Species-level squamate phylogeny continued (T).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 25 of 53http://www.biomedcentral.com/1471-2148/13/93

Bitis rubida
Bitis cornuta
Bitis atropos
Bitis xeropaga
Bitis caudalis
Bitis peringueyi
Bitis gabonica
Bitis nasicornisBitis arietans
Bitis worthingtoni
Atheris desaixi
Atheris nitschei
Atheris squamigera
Atheris hispida
Atheris chlorechis
Atheris barbouri
Atheris ceratophora
Echis pyramidum
Echis leucogaster
Echis coloratus
Echis omanensis
Echis jogeri
Echis ocellatus
Echis carinatus
Causus defilippii
Causus rhombeatus
Causus resimus
Cerastes gasperettii
Cerastes cerastes
Cerastes vipera
Proatheris superciliaris
Vipera lotievi
Vipera renardi
Vipera dinniki
Vipera ursinii
Vipera eriwanensis
Vipera kaznakovi
Vipera barani
Vipera berus
Vipera nikolskii
Vipera seoanei
Vipera aspis
Vipera latastei
Vipera ammodytes
Macrovipera mauritanica
Macrovipera deserti
Daboia russelii
Daboia palaestinae
Montivipera wagneri
Montivipera albizona
Montivipera bornmuelleri
Montivipera raddei
Montivipera xanthina
Macrovipera schweizeri
Macrovipera lebetina
Pseudocerastes fieldi
Pseudocerastes persicus
Eristicophis macmahoni
Pareas nuchalis
Pareas carinatus
Pareas hamptoni
Pareas macularius
Pareas margaritophorus
Pareas formosensis
Pareas boulengeri
Pareas monticola
Aplopeltura boa
Asthenodipsas vertebralis
Achalinus rufescens
Achalinus meiguensis
Xenodermus javanicus
Stoliczkia borneensis
Acrochordus arafurae
Acrochordus granulatus
Acrochordus javanicus
100
100
95
100
100
97
94
100
96
98
99
98100
100
100
63
100
51
77
53
100
93
86
89
100
62
91
100
100
100
100
100
100
100
88
95
100
77
99
97
99
855980
93
100
100
96
59
100
100
99
9483
100
100
99
99
96
95
100100
91
100
95
100
98
97
10095
Crotalus adamanteusCrotalus tigris
Crotalus mitchelliiCrotalus viridis
Crotalus oreganusCrotalus scutulatusCrotalus tortugensis
Crotalus atroxCrotalus catalinensis
Crotalus ruberCrotalus totonacusCrotalus molossus
Crotalus basiliscusCrotalus simusCrotalus durissus
Crotalus horridusCrotalus willardi
Crotalus ravusCrotalus cerastesCrotalus polystictus
Crotalus enyoCrotalus aquilus
Crotalus lepidusCrotalus pusillus
Crotalus triseriatusCrotalus tancitarensis
Crotalus transversusCrotalus intermedius
Crotalus priceiSistrurus miliarius
Sistrurus catenatusAgkistrodon bilineatus
Agkistrodon tayloriAgkistrodon piscivorus
Agkistrodon contortrixBothriechis rowleyi
Bothriechis auriferBothriechis bicolor
Bothriechis thalassinusBothriechis marchi
Bothriechis lateralisBothriechis nigroviridis
Bothriechis schlegeliiLachesis mutaLachesis stenophrys
Mixcoatlus melanurusMixcoatlus barbouri
Ophryacus undulatusBothrops leucurus
Bothrops moojeniBothrops atrox
Bothrops colombiensisBothrops marajoensis
Bothrops asperBothrops lanceolatus
Bothrops caribbaeusBothrops punctatusBothrops jararacussu
Bothrops braziliBothriopsis bilineata
Bothriopsis taeniataBothriopsis chloromelasBothriopsis pulchra
Bothropoides alcatrazBothropoides insularis
Bothropoides jararacaBothropoides neuwiedi
Bothropoides diporusBothropoides erythromelas
Rhinocerophis itapetiningaeRhinocerophis alternatus
Rhinocerophis fonsecaiRhinocerophis cotiara
Rhinocerophis ammodytoidesBothrops pictus
Bothrocophias hyoproraBothrocophias microphthalmus
Bothrocophias campbelliPorthidium lansbergii
Porthidium porrasiPorthidium nasutum
Porthidium yucatanicumPorthidium dunni
Porthidium ophryomegasCerrophidion petlalcalensis
Cerrophidion tzotzilorumCerrophidion godmani
Atropoides picadoiAtropoides olmec
Atropoides nummiferAtropoides occiduus
Gloydius saxatilisGloydius intermedius
Gloydius shedaoensisGloydius halys
Gloydius strauchiGloydius blomhoffiiGloydius ussuriensis
Gloydius brevicaudusGloydius tsushimaensis
Trimeresurus gracilisOvophis okinavensis
Protobothrops tokarensisProtobothrops flavoviridis
Protobothrops mucrosquamatusProtobothrops elegans
Protobothrops jerdoniiProtobothrops xiangchengensis
Protobothrops cornutusProtobothrops mangshanensis
Protobothrops sieversorumProtobothrops kaulbacki
Ovophis tonkinensisOvophis zayuensisOvophis monticola
Trimeresurus purpureomaculatusTrimeresurus erythrurus
Trimeresurus cantoriTrimeresurus albolabris
Trimeresurus andersoniiTrimeresurus septentrionalis
Trimeresurus insularisTrimeresurus fasciatus
Trimeresurus macropsTrimeresurus venustus
Trimeresurus kanburiensisTrimeresurus vogeli
Trimeresurus stejnegeriTrimeresurus gumprechti
Trimeresurus yunnanensisTrimeresurus medoensis
Trimeresurus tibetanusTrimeresurus popeiorum
Trimeresurus sumatranusTrimeresurus schultzei
Trimeresurus flavomaculatusTrimeresurus malcolmiTrimeresurus hageni
Trimeresurus trigonocephalusTrimeresurus gramineus
Trimeresurus malabaricusTrimeresurus borneensis
Trimeresurus puniceusHypnale zara
Hypnale hypnaleHypnale nepa
Calloselasma rhodostomaDeinagkistrodon acutus
Tropidolaemus wagleriGarthius chaseni
Azemiops feae
100
100
97
97
100
91
96
100
74
97
99
100
7789
90
98
58
100100
82
100100
91
100
63
87
99
56
100
95
52
10098 100
97
100
9999
85
5398
100 100
84
98
51
100
100
100
100
90
91
99
100
10090
9972
87
97
100
100
9895
100100
90
100
94
96
9499
100
99
99
90
78
100100
8690
96
9499
100100
85
100
96100
9995
100100
99
100
98
99
90100
63 100
100
92100
97
100
100
98
93
64
10093
77100
99
85
96
100
94
8172 96
89
99
9989
97
98
100 96
99
100100
95
91
50
Vip
erid
ae
Acrochordidae
Xen
od
erm
atid
ae
Vip
erin
ae
Azemiopinae
Cro
talin
ae
UV-AA
(i)
(ii)
(i)
(ii)
Par
eati
dae
Figure 22 Species-level squamate phylogeny continued (U).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 26 of 53http://www.biomedcentral.com/1471-2148/13/93

Liophidium chabaudiLiophidium mayottensis
Liophidium torquatumLiophidium therezieni
Liophidium vaillantiLiophidium rhodogaster
Liopholidophis sexlineatusLiopholidophis dolicocercus
Liopholidophis dimorphusHeteroliodon occipitalis
Pseudoxyrhopus ambreensisThamnosophis lateralis
Thamnosophis stumpffiThamnosophis epistibes
Thamnosophis martaeThamnosophis infrasignatus
Dromicodryas bernieriDromicodryas quadrilineatus
Lycodryas granulicepsLycodryas pseudogranuliceps
Lycodryas inopinaeLycodryas sanctijohannis
Lycodryas citrinusLycodryas inornatus
Madagascarophis meridionalisMadagascarophis colubrinus
Ithycyphus oursiIthycyphus miniatus
Micropisthodon ochraceusLangaha madagascariensis
Leioheterodon modestusLeioheterodon madagascariensis
Leioheterodon geayiParastenophis betsileanus
Alluaudina bellyiCompsophis infralineatus
Compsophis laphystiusCompsophis albiventris
Compsophis boulengeriDitypophis vivaxDuberria lutrix
Duberria variegataAmplorhinus multimaculatus
Lycodonomorphus whytiiLycodonomorphus laevissimus
Lycodonomorphus rufulusLycodonomorphus inornatus
Lamprophis fiskiiLamprophis aurora
Lamprophis fuscusLamprophis guttatus
Boaedon olivaceusBoaedon fuliginosus
Boaedon lineatusBoaedon virgatus
Bothrophthalmus lineatusBothrophthalmus brunneus
Bothrolycus aterPseudoboodon lemniscatus
Gonionotophis capensisGonionotophis poensis
Gonionotophis brussauxiGonionotophis nyassae
Gonionotophis stenophthalmusInyoka swazicus
Hormonotus modestusLycophidion ornatum
Lycophidion capenseLycophidion laterale
Lycophidion nigromaculatumBuhoma depressiceps
Buhoma procteraePsammodynastes pictus
Psammodynastes pulverulentusPseudaspis cana
Pythonodipsas carinataAparallactus guentheri
Aparallactus capensisAparallactus werneriAparallactus modestus
Polemon acanthiasPolemon collaris
Polemon notatusAmblyodipsas dimidiata
Xenocalamus transvaalensisAmblyodipsas polylepis
Macrelaps microlepidotusAtractaspis corpulenta
Atractaspis micropholisAtractaspis bibronii
Atractaspis boulengeriAtractaspis microlepidota
Atractaspis irregularisHomoroselaps lacteus
Psammophis phillipsiPsammophis mossambicusPsammophis leopardinus
Psammophis sibilansPsammophis rukwae
Psammophis orientalisPsammophis sudanensis
Psammophis subtaeniatusPsammophis lineatusPsammophis biseriatus
Psammophis tanganicusPsammophis praeornatus
Psammophis punctulatusPsammophis schokari
Psammophis angolensisPsammophis notostictusPsammophis leightoniPsammophis jallae
Psammophis trigrammusPsammophis condanarus
Psammophis lineolatusPsammophis crucifer
Psammophylax tritaeniatusPsammophylax variabilis
Psammophylax rhombeatusPsammophylax acutus
Hemirhagerrhis hildebrandtiiHemirhagerrhis kelleri
Hemirhagerrhis viperinaDipsina multimaculata
Mimophis mahfalensisRhagerhis moilensis
Malpolon monspessulanusRhamphiophis rubropunctatus
Rhamphiophis oxyrhynchusProsymna meleagris
Prosymna greigertiProsymna janii
Prosymna visseriProsymna ruspolii
Oxyrhabdium leporinumMicrelaps bicoloratus
Cerberus rynchopsCerberus microlepis
Cerberus australisHomalopsis buccataEnhydris punctata
Fordonia leucobaliaGerarda prevostiana
Cantoria violaceaBitia hydroides
Enhydris bocourtiErpeton tentaculatum
Pseudoferania polylepisMyron richardsonii
Enhydris innominataEnhydris longicauda
Enhydris jagoriiEnhydris enhydris
Enhydris matannensisEnhydris plumbea
Enhydris chinensis
100
58
98
95
96
90
100
95
76
99
10089 99
73
99
94
98100
95
98
100 9398
95
100
90
97
94100
98
54 100
100
100
100
100100
82
98100
94
99
99100
100
92
98
95
95
100 98
99
100100
97100
93
100
100
97
100
6992
95
64
92
100
100
100
97
54100
92100
100 93
96
100 94
98
9698
58 99
95
100
90
100
100
80
69
9673
10080
71100
100
97
60
100
76
1008550
70
100
97
6597
10095
100
54
9996
99
10086
100
100
85
88
8897
100 100
96
7098
7189
100
97
100100
100
53100
Homalopsidae
Prosymninae
Psammophiinae
Atractaspidinae
Aparallactinae
Pseudaspidinae
Lamprophiinae
Pseudoxyrhophiinae
Lamprophiidae
VX-AA W
Figure 23 Species-level squamate phylogeny continued (V).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 27 of 53http://www.biomedcentral.com/1471-2148/13/93

Hydrophis ornataHydrophis peronii
Hydrophis kingiiHydrophis macdowelli
Hydrophis schistosaHydrophis majorHydrophis czeblukovi
Hydrophis caerulescensHydrophis brooki
Hydrophis atricepsHydrophis stokesii
Hydrophis platuraHydrophis curtus
Hydrophis parvicepsHydrophis melanocephala
Hydrophis cyanocinctaHydrophis pacificaHydrophis semperi
Hydrophis spiralisHydrophis lapemoides
Hydrophis elegansHydrelaps darwiniensis
Ephalophis greyaeParahydrophis mertoni
Aipysurus laevisAipysurus fuscusAipysurus apraefrontalisAipysurus duboisii
Aipysurus eydouxiiEmydocephalus annulatus
Hemiaspis dameliiHemiaspis signata
Tropidechis carinatusNotechis scutatus
Echiopsis atricepsHoplocephalus bitorquatus
Austrelaps superbusAustrelaps labialis
Drysdalia coronoidesDrysdalia mastersii
Echiopsis curtaPseudonaja textilisPseudonaja modesta
Oxyuranus scutellatusOxyuranus microlepidotus
Denisonia devisiSimoselaps calonotus
Vermicella intermediaSuta spectabilis
Suta sutaSuta monachus
Suta fasciataRhinoplocephalus bicolor
Cryptophis nigrescensElapognathus coronata
Cacophis squamulosusPseudechis guttatus
Pseudechis papuanusPseudechis colletti
Pseudechis australisPseudechis butleri
Pseudechis porphyriacusAcanthophis antarcticusAcanthophis praelongus
Aspidomorphus muelleriAspidomorphus lineaticollis
Aspidomorphus schlegeliSimoselaps bertholdi
Simoselaps anomalusSimoselaps semifasciatus
Furina diademaFurina ornata
Toxicocalamus loriaeDemansia psammophis
Demansia papuensisDemansia vestigiata
Toxicocalamus preussiMicropechis ikaheka
Laticauda colubrinaLaticauda guineai
Laticauda saintgironsiLaticauda laticaudata
Bungarus candidusBungarus multicinctus
Bungarus nigerBungarus caeruleus
Bungarus ceylonicusBungarus sindanus
Bungarus fasciatusBungarus bungaroidesBungarus flavicepsElapsoidea sundevalliiElapsoidea semiannulata
Elapsoidea nigraNaja ashei
Naja nigricollisNaja mossambica
Naja katiensisNaja pallida
Naja nubiaeNaja haje
Naja annuliferaNaja nivea
Naja annulataNaja melanoleuca
Naja multifasciataNaja sumatranaNaja siamensis
Naja mandalayensisNaja naja
Naja atraNaja kaouthia
Hemachatus haemachatusAspidelaps scutatus
Walterinnesia aegyptiaDendroaspis polylepis
Dendroaspis angusticepsOphiophagus hannah
Hemibungarus calligasterMicrurus brasiliensisMicrurus frontalisMicrurus spixii
Micrurus ibibobocaMicrurus altirostris
Micrurus pyrrhocryptusMicrurus baliocoryphus
Micrurus decoratusMicrurus lemniscatusMicrurus hemprichii
Micrurus surinamensisMicrurus dissoleucus
Micrurus mipartitusMicrurus narduccii
Micrurus diastemaMicrurus fulvius
Micrurus psychesMicrurus albicinctus
Micrurus corallinusSinomicrurus macclellandi
Sinomicrurus kelloggiSinomicrurus japonicus
Micruroides euryxanthusMaticora bivirgata
Calliophis melanurus
100
98
95
90
100
92
85
67
89
75
99
91
100
98
72
100
95
67
83
938874
100
8377
9854
94
10097100 75
94
96
100
9490 100
97
100
78
97
9899
98
98
100
100
71
100 97
100
94
88
98100
90
8895
68
100 100100
100
100
100
99100
82 100
100100
91
10098
100
98
100100
99 100
9797
10099
87
100
91
90
83
92
9996
9676
78
100
88
69
58
100
9987
63
84
84 100
61
85
100
91
9498
88
8959
97
97
98
10090
99
10095
Elapidae
W
Figure 24 Species-level squamate phylogeny continued (W).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 28 of 53http://www.biomedcentral.com/1471-2148/13/93

Eirenis punctatolineatusEirenis barani
Eirenis collarisEirenis eiselti
Eirenis coronelloidesEirenis rothiiEirenis lineomaculatus
Eirenis thospitisEirenis medus
Eirenis levantinusEirenis decemlineatus
Eirenis persicusEirenis aurolineatus
Eirenis modestusHierophis spinalisHierophis gemonensis
Hierophis viridiflavusDolichophis caspius
Dolichophis schmidtiDolichophis jugularis
Platyceps florulentusPlatyceps collaris
Platyceps najadumPlatyceps ventromaculatus
Platyceps kareliniPlatyceps rogersi
Platyceps rhodorachisSpalerosophis diadema
Spalerosophis microlepisHemorrhois ravergieri
Hemorrhois nummiferHemorrhois algirus
Hemorrhois hippocrepisHemerophis socotrae
Macroprotodon abubakeriMacroprotodon cucullatus
Bamanophis dorriColuber zebrinus
Lytorhynchus diademaHapsidophrys principis
Hapsidophrys smaragdinaHapsidophrys lineatus
Philothamnus hoplogasterPhilothamnus natalensisPhilothamnus thomensis
Philothamnus girardiPhilothamnus angolensis
Philothamnus semivariegatusPhilothamnus nitidus
Philothamnus heterodermusPhilothamnus carinatus
Thelotornis capensisDispholidus typus
Thrasops jacksoniiCoelognathus flavolineatus
Coelognathus subradiatusCoelognathus erythrurus
Coelognathus helenaCoelognathus radiatus
Oligodon planicepsOligodon torquatus
Oligodon theobaldiOligodon cruentatus
Oligodon splendidusOligodon maculatus
Oligodon cinereusOligodon formosanus
Oligodon ocellatusOligodon chinensisOligodon taeniatusOligodon barroniOligodon cyclurus
Oligodon octolineatusOligodon taeniolatusOligodon sublineatus
Oligodon arnensisScaphiophis albopunctatus
Dasypeltis fasciataDasypeltis gansi
Dasypeltis sahelensisDasypeltis scabra
Dasypeltis atraDasypeltis confusa
Toxicodryas pulverulentaBoiga multomaculata
Boiga beddomeiBoiga ceylonensisBoiga trigonata
Boiga barnesiiBoiga cynodon
Boiga forsteniBoiga dendrophila
Boiga irregularisTelescopus fallax
Dipsadoboa unicolorCrotaphopeltis tornieri
Boiga kraepeliniDendrelaphis tristis
Dendrelaphis schokariDendrelaphis caudolineatus
Dendrelaphis caudolineolatusDendrelaphis bifrenalis
Chrysopelea paradisiChrysopelea ornata
Chrysopelea taprobanicaAhaetulla nasuta
Ahaetulla pulverulentaAhaetulla fronticincta
Grayia smithiiGrayia ornataGrayia tholloni
Sibynophis chinensisSibynophis collaris
Sibynophis triangularisSibynophis subpunctatus
Sibynophis bistrigatusScaphiodontophis annulatus
Pseudoxenodon karlschmidtiPseudoxenodon macrops
Pseudoxenodon bambusicolaPlagiopholis styani
Calamaria yunnanensisCalamaria pavimentata
Pseudorabdion oxycephalum
100
68
74
71
100
57
100
68
100
94
60
94
100
92
83
100
83
97
91100
8659
86
82
100
100
100
100
99
100
100
9852
61
8695
100
100
100
9286 100
60
99
69
8768 99
99
9490
88
52
9098
100
10099
86
100
99
98
98 100
97
93
56
100
89
9990
85
99
100
83
100
79
9870
9277
90
100
74
85
73
8996
9668
7199
100
10097
91
69
100100
100
100
9668
100100
100
97
10097
85
99 100
Co
lub
rid
ae
Calamariinae
Pseudoxenodontinae
Sibynophiinae
Grayiinae
Col
ubrin
ae
X YZ-AA Colubrinae cont.
Figure 25 Species-level squamate phylogeny continued (X).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 29 of 53http://www.biomedcentral.com/1471-2148/13/93

Lampropeltis californiaeLampropeltis splendidaLampropeltis holbrooki
Lampropeltis getulaLampropeltis nigraLampropeltis alterna
Lampropeltis extenuatumLampropeltis triangulumLampropeltis ruthveniLampropeltis mexicanaLampropeltis elapsoidesLampropeltis zonataLampropeltis pyromelana
Lampropeltis webbiLampropeltis calligaster
Cemophora coccineaPseudelaphe flavirufa
Arizona elegansRhinocheilus lecontei
Bogertophis rosaliaeBogertophis subocularis
Pantherophis bairdiPantherophis obsoletusPantherophis alleghaniensis
Pantherophis spiloidesPantherophis slowinskii
Pantherophis emoryiPantherophis guttatus
Pantherophis vulpinusPituophis cateniferPituophis ruthveniPituophis melanoleucus
Pituophis lineaticollisPituophis vertebralis
Pituophis deppeiSenticolis triaspis
Oocatochus rufodorsatusCoronella austriaca
Coronella girondicaElaphe sauromates
Elaphe quatuorlineataElaphe quadrivirgata
Elaphe schrenckiiElaphe climacophora
Elaphe dioneElaphe bimaculata
Elaphe carinataElaphe davidi
Zamenis situlaRhinechis scalaris
Zamenis lineatusZamenis longissimus
Zamenis persicusZamenis hohenackeri
Orthriophis cantorisOrthriophis moellendorffi
Orthriophis hodgsoniOrthriophis taeniurus
Oreocryptophis porphyraceusEuprepiophis mandarinus
Euprepiophis conspicillataArchelaphe bella
Lycodon aulicusLycodon zawi
Lycodon osmanhilliLycodon capucinus
Lycodon carinatusDryocalamus nympha
Lycodon semicarinatumLycodon rufozonatum
Lycodon paucifasciatusLycodon fasciatus
Lycodon laoensisLycodon ruhstrati
Rhadinophis frenatusRhynchophis boulengeri
Rhadinophis prasinusGonyosoma jansenii
Gonyosoma oxycephalumSympholis lippiens
Pseudoficimia frontalisGyalopion canum
Ficimia streckeriConopsis nasusConopsis biserialis
Sonora semiannulataChilomeniscus stramineus
Chionactis occipitalisStenorrhina freminvillei
Rhinobothryum lentiginosumMastigodryas bifossatus
Mastigodryas boddaertiMastigodryas melanolomus
Drymoluber braziliDrymoluber dichrous
Chironius multiventrisChironius laurenti
Chironius exoletusChironius monticola
Chironius bicarinatusChironius flavolineatus
Chironius grandisquamisChironius laevicollisChironius scurrulus
Chironius fuscusDrymarchon corais
Pseustes sulphureusSpilotes pullatus
Phyllorhynchus decurtatusTrimorphodon biscutatus
Drymobius rhombiferDendrophidion percarinatum
Dendrophidion dendrophisLeptophis ahaetulla
Chironius carinatusChironius quadricarinatus
Coluber flagellumColuber constrictor
Coluber taeniatusTantilla melanocephala
Salvadora mexicanaOpheodrys aestivusOpheodrys vernalisOxybelis aeneus
Oxybelis fulgidusPtyas mucosa
Ptyas korrosCyclophiops major
64
83
100
99
90
75
84
98
100
87
10075
92
82
10096
7496
6098
98
100
7279
94
98
100
100
96
100 98100
100
99
100
91
9499
9883
100
61
83 100
55
93100
99
64100
100
100
99
8898
100
53
83
98 94 76
86
9293
60
98100
100
100
93
93
100
87
94
100
99
71
89
10092
87
100
98
87
86
8183
5499
74
9182
9792
9966
92
93
88
96 10082
100
81
Colubrinae cont.
Y
Figure 26 Species-level squamate phylogeny continued (Y).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 30 of 53http://www.biomedcentral.com/1471-2148/13/93

Thamnophis butleriThamnophis radixThamnophis brachystomaThamnophis elegansThamnophis gigasThamnophis atratusThamnophis couchii
Thamnophis ordinoidesThamnophis hammondii
Thamnophis marcianusThamnophis equesThamnophis cyrtopsis
Thamnophis chrysocephalusThamnophis fulvus
Thamnophis mendaxThamnophis sumichrasti
Thamnophis scaligerThamnophis godmaniThamnophis exsulThamnophis melanogaster
Thamnophis validaAdelophis foxi
Thamnophis rufipunctatusThamnophis proximusThamnophis sauritus
Thamnophis sirtalisNerodia sipedon
Nerodia fasciataNerodia harteriNerodia erythrogaster
Nerodia rhombiferNerodia taxispilota
Nerodia floridanaNerodia cyclopion
Tropidoclonion lineatumRegina grahami
Regina septemvittataRegina alleni
Regina rigidaSeminatrix pygaea
Storeria dekayiStoreria occipitomaculata
Virginia striatulaClonophis kirtlandii
Natrix tessellataNatrix natrix
Natrix mauraSinonatrix annularis
Sinonatrix percarinataSinonatrix aequifasciata
Opisthotropis cheniOpisthotropis latouchii
Opisthotropis lateralisOpisthotropis guangxiensis
Aspidura guentheriAspidura ceylonensis
Aspidura trachyproctaAspidura drummondhayi
Atretium yunnanensisXenochrophis asperrimus
Xenochrophis punctulatusAtretium schistosum
Xenochrophis piscatorXenochrophis flavipunctatusXenochrophis vittatus
Amphiesma stolatumRhabdophis nuchalis
Rhabdophis tigrinusRhabdophis subminiatus
Balanophis ceylonensisMacropisthodon rudis
Lycognathophis seychellensisAfronatrix anoscopus
Natriciteres olivaceaAmphiesma craspedogasterAmphiesma sauteri
Trachischium monticola
100
98
100
100
100
100
96
86
99
100
92
63
73
94100
89
9969
99
100
93
100
85100 98
72
69100
96
100
99
100
100
100
76
50
97
80100
72
98
100
88
100
82
100
86100
100
99
100
99
68
69
98
100
88
88
100
6296
98
99
100
Natricinae
Z
Figure 27 Species-level squamate phylogeny continued (Z).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 31 of 53http://www.biomedcentral.com/1471-2148/13/93

Sibynomorphus neuwiedi
Sibynomorphus ventrimaculatus
Dipsas albifrons
Dipsas articulata
Sibynomorphus turgidus
Sibynomorphus mikanii
Dipsas pratti
Dipsas variegata
Dipsas neivai
Dipsas catesbyi
Dipsas indica
Tropidodipsas sartorii
Sibon nebulatus
Ninia atrata
Atractus zebrinus
Atractus trihedrurus
Atractus reticulatus
Atractus badius
Atractus flammigerus
Atractus elaps
Atractus albuquerquei
Atractus schach
Atractus wagleri
Atractus zidoki
Geophis godmani
Geophis carinosus
Cryophis hallbergi
Tretanorhinus nigroluteus
Hydromorphus concolor
Adelphicos quadrivirgatum
Leptodeira rubricata
Leptodeira maculata
Leptodeira bakeri
Leptodeira annulata
Leptodeira septentrionalis
Leptodeira punctata
Leptodeira splendida
Leptodeira frenata
Leptodeira uribei
Imantodes cenchoa
Imantodes gemmistratus
Imantodes lentiferus
Imantodes inornatus
Leptodeira nigrofasciata
Tretanorhinus variabilis
Hypsiglena chlorophaea
Hypsiglena torquata
Hypsiglena ochrorhyncha
Hypsiglena affinis
Hypsiglena jani
Hypsiglena slevini
Trimetopon gracile
Pseudoleptodeira latifasciata
Rhadinaea fulvivittis
Rhadinaea flavilata
Coniophanes fissidens
Tantalophis discolor
Amastridium veliferum
Nothopsis rugosus
Farancia erytrogramma
Farancia abacura
Carphophis amoenus
Diadophis punctatus
Heterodon nasicus
Heterodon simus
Heterodon platirhinos
Contia tenuis
Thermophis baileyi
Thermophis zhaoermii
100
100
93
63
89
90
91
83
70
100
82
70
75
79
88
79
96
84
100
90
86
96
73
75
73
97
89
68
68
94
100
64
67
100
91
97
88
100
98
100
71
10086
99
100
99
84
85
80
96
86
68
58
87
8098
72
100
99
100
Pseudoboa coronataPseudoboa nigra
Pseudoboa neuwiediiRhachidelus brazili
Boiruna maculataDrepanoides anomalus
Mussurana bicolorClelia clelia
Phimophis gueriniParaphimophis rusticus
Rodriguesophis iglesiasiOxyrhopus guibei
Oxyrhopus melanogenysOxyrhopus rhombifer
Oxyrhopus clathratusOxyrhopus trigeminus
Oxyrhopus formosusOxyrhopus petolarius
Siphlophis longicaudatusSiphlophis pulcher
Siphlophis compressusSiphlophis cervinus
Hydrodynastes gigasHydrodynastes bicinctus
Caaeteboia amaraliXenopholis scalarisXenopholis undulatus
Tropidodryas striaticepsTropidodryas serra
Thamnodynastes rutilusThamnodynastes strigatus
Thamnodynastes hypoconiaThamnodynastes laneiThamnodynastes pallidusTomodon dorsatus
Ptychophis flavovirgatusTachymenis peruviana
Pseudotomodon trigonatusCalamodontophis paucidens
Gomesophis brasiliensisHelicops angulatus
Helicops gomesiHelicops carinicaudus
Helicops hagmanniHelicops infrataeniatus
Pseudoeryx plicatilisHydrops triangularis
Manolepis putnamiApostolepis cearensisApostolepis sanctaeritae
Apostolepis flavotorquataApostolepis albicollaris
Apostolepis dimidiataApostolepis assimilis
Elapomorphus quinquelineatusPhalotris mertensi
Phalotris lativittatusPhalotris nasutusPhalotris lemniscatusTaeniophallus affinisEchinanthera undulataEchinanthera melanostigma
Taeniophallus nicagusTaeniophallus brevirostris
Sordellina punctataPhilodryas patagoniensis
Philodryas agassiziiPhilodryas aestiva
Philodryas psammophideaPhilodryas mattogrossensis
Philodryas georgeboulengeriPhilodryas argentea
Philodryas olfersiiPhilodryas viridissima
Philodryas nattereriPhilodryas baroni
Erythrolamprus aesculapiiErythrolamprus mimus
Eryrthrolamprus typhlusEryrthrolamprus pygmaea
Eryrthrolamprus brevicepsEryrthrolamprus reginae
Eryrthrolamprus miliarisEryrthrolamprus juliae
Eryrthrolamprus epinephelusEryrthrolamprus poecilogyrusEryrthrolamprus ceii
Eryrthrolamprus almadensisEryrthrolamprus jaegeri
Eryrthrolamprus atraventerXenodon pulcherXenodon matogrossensisXenodon semicinctusXenodon guentheri
Xenodon nattereriXenodon histricus
Xenodon dorbignyiXenodon neuwiedii
Xenodon werneriXenodon merremi
Xenodon severusLygophis meridionalis
Lygophis flavifrenatusLygophis paucidens
Lygophis lineatusLygophis elegantissimus
Lygophis anomalusUromacer frenatusUromacer oxyrhynchusUromacer catesbyi
Schwartzophis funereumSchwartzophis polylepis
Schwartzophis callilaemumAntillophis parvifrons
Hypsirhynchus feroxDarlingtonia haetiana
Ialtris dorsalisAlsophis rufiventrisAlsophis antiguaeAlsophis rijgersmaei
Alsophis antillensisBorikenophis portoricensisHaitiophis anomalusCaraiba andreae
Cubophis vudiiCubophis cantherigerus
Magliophis exiguumArrhyton dolichura
Arrhyton tanyplectumArrhyton procerum
Arrhyton landoiArrhyton vittatum
Arrhyton supernumArrhyton taeniatum
Pseudalsophis elegansPseudalsophis biserialis
Psomophis jobertiPsomophis genimaculatus
Psomophis obtususCrisantophis nevermanni
Conophis lineatus
91
90
90
91
68
100
95
87
96
7492
100
87
90
96
95
6763
9567
98
100
91
87
56100
100
100
91
100
8786
9490
93 99
90
97
55
10098
91 99
79
100
5695
9794
6999
95
100100
95
6667
75
100
100
68
68
6294
96
100
88
51
81
89
78
100
94
99
87
9688
87100
97
94
100
92
73
100
7599
7161 98
62
98
9852
9976
7370
100
100
99
75
99
80
97
83
98100
100
99
99
100
97100
100
93
100
100
73
10098
100100 94
95
AA
Dip
sadi
nae
(i)
(ii)
(ii)
(i)
Figure 28 Species-level squamate phylogeny continued (AA).
Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 32 of 53http://www.biomedcentral.com/1471-2148/13/93

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 33 of 53http://www.biomedcentral.com/1471-2148/13/93
need for considering these clades as families, since thefamily Scincidae is clearly monophyletic, based on our re-sults and others (see above). Thus, their new taxonomychanges the long-standing definition of Scincidae un-necessarily (see [113]). Furthermore, these changes weredone without defining the full content (beyond a typegenus) of any of these families other than Scincidae (theformer Scincinae + Feylininae) and Acontiidae (the formerAcontiinae).Most importantly, the new taxonomy proposed by these
authors [112] is at odds with the phylogeny estimatedhere, with respect to the familial and subfamilial classifica-tion of >1000 skink species (Figures 6, 7, 8, 9, 10). For in-stance, Sphenomorphus stellatus is found in a stronglysupported clade containing Lygosoma (presumablyLygosomidae; Figure 10), which is separate from the otherclade (presumably Sphenomorphidae) containing theother sampled Sphenomorphus species (Figure 7; note thatthese Sphenomorphus species are divided among severalsubclades within this latter clade). An additional problemis that Egernia, Lygosoma, and Sphenomorphus are thetype genera of Egerniidae, Lygosomidae, and Sphenomor-phidae, but are paraphyletic as currently defined (Figures 7,8, 9, 10), leading to further uncertainty in the content anddefinition of these putative families.Furthermore, Mabuyidae apparently refers to the
clade (Figure 9) containing Chioninia, Dasia, Mabuya,Trachylepis, with each of these genera placed in its ownsubfamily (Chioniniinae, Dasiinae, Mabuyinae, andTrachylepidinae). However, several other genera arestrongly placed in this group, such as Eumecia, Eutropis,Lankaskincus, and Ristella (Figure 9). These other ge-nera cannot be readily fit into these subfamilial groups(i.e. they are not the sister group of any genera in thosesubfamilies), and Trachylepis is paraphyletic with respectto Eumecia, Chioninia, and Mabuya (Figure 9). Also, wefind that Emoia is divided between clades containingLygosoma (Lygosomidae) and Eugongylus (Eugongylidae),and many of these relationships have strong support(Figure 10). Finally, Ateuchosaurus is apparently notaccounted for in their classification, and here is weaklyplaced as the sister-group to a clade comprising theirSphenomorphidae, Egerniidae, Mabuyidae, and Lygoso-midae (i.e. Lygosominae as recognized here; Figures 7,8, 9, 10).These authors [112] argued that a more heavily
subdivided classification for skinks may be desirable forfacilitating future taxonomic revisions and species de-scriptions. However, this classification seems likely toonly exacerbate existing taxonomic problems (e.g. pla-cing congeneric species in different families without re-vising the genus-level taxonomy). Here, we retain theprevious definition of Mabuya (restricted to the NewWorld clade; sensu [114]), and we support the traditional
definitions of Scincidae, Acontiinae, Scincinae (but in-cluding Feylininae), and Lygosominae (Figures 6, 7, 8, 9,10; note that we leave Ateuchosaurus as incertae sedis).The other taxonomic issues in Scincidae identified hereand elsewhere should be resolved in future studies. Ourphylogeny provides a framework in which these analysescan take place (i.e. identifying major subclades withinskinks), which we think may be more useful than a classi-fication lacking clear taxon definitions.Of the 133 scincid genera [1], we can assign the 113
sampled in our tree to one of the three subfamilies in ourclassification (Acontiinae, Lygosominae, and Scincinae;Appendix I). We place 19 of the remaining genera intoone of the three subfamilies based on previous classifica-tions (e.g. [110]), with Ateuchosaurus as incertae sedis inScincidae. Below, we review the non-monophyletic generain our tree. Many of these problems have been reportedby previous authors [51,111,115-118], and for brevity wedo not distinguish between cases reported in previousstudies, and potentially new instances found here.Within Acontiinae, we find that the two genera are
both monophyletic (Figure 6). Within Scincinae, manygenera are now strongly monophyletic (thanks in part tothe dismantling of Eumeces; [49,50,119]), but some prob-lems remain (Figure 6). The genera Scincus and Scincopusare strongly supported as being nested inside of theremaining Eumeces. Among Malagasy scincines (see[49,120]), Pseudacontias is nested inside Madascincus,and the genera Androngo, Pygomeles, and Voeltzkowia areall nested in Amphiglossus (Figure 6).We also find numerous taxonomic problems within
lygosomines (Figures 7, 8, 9, 10). Species of Sphenomor-phus are widely dispersed among other lygosomine gen-era. The genus Tropidophorus is paraphyletic with respectto a clade containing many other genera (Figure 7). Thesampled species of Asymblepharus are only distantly re-lated to each other, including one species (A. sikimmensis)nested inside of Scincella (Figure 7). The genus Lipinia ispolyphyletic, with one species (L. vittigera) strongly placedas the sister taxon to Isopachys, and with two other species(L. pulchella and L. noctua) placed in a well-supportedclade that also includes Papuascincus (Figure 7).Among Australian skinks, the genus Eulamprus
is polyphyletic with respect to Nangura, Calyptotis,Gnypetoscincus, Coggeria, Coeranoscincus, Ophioscincus,Saiphos, Anomalopus, Eremiascincus, Hemiergis, Gla-phyromorphus, Notoscincus, Ctenotus, and Lerista, andmost of the relevant nodes are strongly supported(Figure 8). The genera Coeranoscincus and Ophios-cincus are polyphyletic with respect to each other andto Saiphos and Coggeria (Figure 8). The genus Glaphy-romorphus is paraphyletic with respect to a clade ofEulamprus (Figure 8). The genus Egernia is paraphy-letic with respect to Bellatorias (which is paraphyletic

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 34 of 53http://www.biomedcentral.com/1471-2148/13/93
with respect to Egernia and Lissolepis) and Lissolepis,although many of the relevant nodes are not stronglysupported (Figure 9). The genera Cyclodomorphus andTiliqua are paraphyletic with respect to each other(Figure 9).Among other lygosomines, Trachylepis is non-
monophyletic [121], with two species (T. aurata and T.vittata) that fall outside the strongly supported cladecontaining the other species (Figure 9). The latter clade isweakly supported as the sister group to a clade containingChioninia, Eumecia, and Mabuya. In Mabuya, a few spe-cies (M. altamazonica, M. bistriata, and M. nigropuncata)have unorthodox placements within a monophyleticMabuya, potentially due to uncertain taxonomic assign-ment of specimens by previous authors [51,122]. Thegenus Lygosoma is paraphyletic with respect to Lepido-thyris and Mochlus, and many of the relevant nodes arestrongly supported (Figure 10). Among New Caledonianskinks, the genus Lioscincus is polyphyletic with respectto Marmorosphax, Celatiscincus, and Tropidoscincus,and both Lioscincus and Tropidoscincus are paraphyleticwith respect to, Kanakysaurus, Lacertoides, Phoboscincus,Sigaloseps, Tropidoscincus, Graciliscincus, Simiscincus, andCaledoniscincus, with strong support for most relevantnodes (Figure 10). The genera Emoia and Bassiana aremassively polyphyletic and divided across multiplelygosomine clades (Figure 10). The genus Lygisaurus ap-pears to be nested inside of Carlia, although many of therelevant branches are only weakly supported (Figure 10).
LacertoideaWithin Lacertoidea (Figure 1), we corroborate recentmolecular analyses (e.g. [16,17,19,20]) and morphology-based phylogenies and classifications (e.g. [13,15]) insupporting the clade including the New World familiesGymnophthalmidae and Teiidae (Figure 11). Within aweakly supported Teiidae (Figure 11), the subfamiliesTupinambinae and Teiinae are each strongly supportedas monophyletic, as in previous studies [61]. In Tupi-nambinae, Callopistes is the sister group to a cladecontaining Tupinambis, Dracaena, and Crocodilurus.The clade Dracaena + Crocodilurus is nested withinTupinambis, and the associated clades have strong sup-port (Figure 11). We find that the teiine genera Ameivaand Cnemidophorus are non-monophyletic (Figure 11),interdigitating with each other and the monophyleticgenera Aspidoscelis, Dicrodon (monotypic), and Kentropyx,as in previous phylogenies [62].A recent study [123] proposed a re-classification of the
family Teiidae based on analysis of 137 morphologicalcharacters for 101 terminal species (with ~150 species inthe family [1]). Those authors erected several new gen-era and subfamilies in an attempt to deal with the appar-ent non-monophyly of currently recognized taxa in their
tree. However, in our tree, some of these new taxa conflictstrongly with the phylogeny or are rendered unnecessary.First, they recognize Callopistinae as a distinct subfamilyfor Callopistes, arguing that failure to do so wouldproduce a taxonomy inconsistent with teiid phylogeny.However, we find strong support for Callopistes in itstraditional placement as part of Tupinambinae (Figure 11),and this change is thus not needed based on our results.The genus Ameiva is paraphyletic under traditional defini-tions [62]. In our tree, their conception of Ameiva is alsonon-monophyletic, with species found in three distinctclades (Figure 11). We also find non-monophyly of manyof their species groups within Ameiva, including theameiva, bifrontata, dorsalis, and erythrocephala groups(Figure 11). Their genera Aurivela (Cnemidophorus longi-caudus) and Contomastyx (Cnemidophorus lacertoides)are strongly supported as sister taxa in our tree, and arenested within Ameiva in their erythrocephala speciesgroup, along with Dicrodon (Figure 11).On the positive side, many of the genera they recognize
are monophyletic and are not nested in other genera in ourtree, including their Ameivula (Cnemidophorus ocellifer),Aspidoscelis (unchanged from previous definitions),Cnemidophorus (excluding C. ocellifer, C. lacertoides, andC. longicaudus), Holcosus (Ameiva undulata, A. festiva, andA. quadrilineatus), Kentropyx (unchanged from previousdefinitions), Salvator (Tupinambis rufescens, T. duseni, andT. merianae), and Teius (unchanged from previous defini-tions). We did not sample Ameiva edracantha (theirMedopheos).Given our results, major taxonomic rearrangements
within Teiidae seem problematic at present, especiallywith the extensive paraphyly of many traditional and re-defined teiid genera, the lack of strong resolution of manyof these relationships based on molecular and morpho-logical data, and incomplete taxon sampling in all studiesso far. Thus, we provisionally retain the traditional tax-onomy of Teiidae, pending additional data and analyses.However, we note that Ameiva, Cnemidophorus, andTupinambis are clearly non-monophyletic based on bothour results and those of recent authors [123], and will re-quire taxonomic changes in the future. We anticipate thatmany of these newly proposed genera [123] will be usefulin such revisions.We find strong support (SHL = 98) for monophyly of
Gymnophthalmidae (Figure 11). Within Gymnophthal-midae, we find strong support for the monophyly of thepreviously recognized subfamilies [63,64,124], with the ex-ception of Cercosaurinae (Figure 11). Previous researchersconsidered the genus Bachia a distinct tribe (Bachiini)within Cercosaurinae, based on a poorly supported sister-group relationship with the tribe Cercosaurini [63,64].Here, we find a moderately well supported relationship(SHL = 84) between Bachia and Gymnophthalminae +

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 35 of 53http://www.biomedcentral.com/1471-2148/13/93
Rhachisaurinae, and we find that this clade is only dis-tantly related to other Cercosaurinae. Therefore, we re-strict Cercosaurinae to the tribe Cercosaurini, and elevatethe tribe Bachiini [64] to the subfamily level. The subfa-mily Bachiinae contains only the genus Bachia (Figure 11),identical in content to the previously recognized tribe[64]. Within Cercosaurinae, we find that the genusPetracola is nested within Proctoporus (Figure 11). InEcpleopinae, Leposoma is divided into two clades, sepa-rated by Anotosaura, Colobosauroides, and Arthrosaura(Figure 11), and many of the relevant nodes are verystrongly supported. These issues should be addressed infuture studies.Our results show strong support for a clade uniting
Lacertidae and Amphisbaenia (Figure 1), as in many previ-ous studies [16-20,23]. We also find strong support formonophyly of amphisbaenians (SHL = 99), in contrastto some molecular analyses [19,20]. Relationshipsamong amphisbaenian families are generally stronglysupported and similar to those in earlier molecularstudies (Figure 12), including the placement of the NewWorld family Rhineuridae as sister group to all otheramphisbaenians [70,71,125]. The family Cadeidae is placedas the sister-group to Amphisbaenidae + Trogonophiidae(Figures 1 and Figure 12) with weak support, but has beenplaced with Blanidae in previous studies, with strong sup-port but less- extensive taxon sampling [125,126].We find strong support for monophyly of the Old World
family Lacertidae (Figure 13). Within Lacertidae, branchsupport for the monophyly of most genera and for the sub-families Gallotiinae and Lacertinae is very high (Figure 13).However, we find that relationships among many generaare poorly supported, as in previous studies [65,67,68]. Ourresults (Figure 13) also indicate that several lacertid generaare non-monophyletic with strong support for the associ-ated nodes, including Algyroides (paraphyletic with respectto Dinarolacerta), Ichnotropis (paraphyletic with respect toMeroles), Meroles (paraphyletic with respect to Ichno-tropis), Nucras (polyphyletic with respect to several genera,including Pedioplanis, Poromera, Latastia, Philocortus,Pseuderemias, and Heliobolus), and Pedioplanis (paraphy-letic with respect to Nucras).
Higher-level phylogeny of ToxicoferaWe find strong support (SHL = 96) for monophyly ofToxicofera (Anguimorpha, Iguania, and Serpentes;Figure 1), and moderate support for a sister-group re-lationship between Iguania and Anguimorpha (SHL =79). Relationships among Anguimorpha, Iguania, andSerpentes were weakly supported in some Bayesianand likelihood analyses [16-19], but strongly sup-ported in others [20]. We further corroborate previ-ous studies in also placing Anguimorpha with Iguania[16-20]. In contrast, some other studies have placed
anguimorphs with snakes as the sister group toiguanians [127,128].
AnguimorphaOur hypothesis for family-level anguimorphan relation-ships (Figures 1, 14) is generally similar to that of otherrecent studies [17,19,20,129], and is strongly supported.Our results differ from some analyses based only onmorphology, which place Anguidae near the base ofAnguimorpha [130]. Here, Shinisauridae is stronglysupported as the sister taxon to a well-supportedclade of Varanidae + Lanthanotidae (Figures 1, 14).Varanid relationships are similar to previous estimates(e.g. [131]). Xenosauridae is here strongly supportedas the sister-group to a strongly supported cladecontaining Helodermatidae and the strongly supportedAnniellidae + Anguidae clade (Figures 1, 14). However,previous molecular analyses have placed Helodermatidaeas the sister to Xenosauridae + (Anniellidae + Anguidae),typically with strong support [16,17,19,20].Within Anguidae (Figure 14), our phylogeny indicates
non-monophyly of genera within every subfamily, includ-ing Diploglossinae (Diploglossus and Celestus are stronglysupported as paraphyletic with respect to each other andto Ophiodes), Anguinae (Ophisaurus is strongly supportedas paraphyletic with respect to Anguis, Dopasia, andPseudopus), and Gerrhonotinae (Abronia and Mesaspisare non-monophyletic, and Coloptychon is nested insideGerrhonotus). Some of these problems were not reportedpreviously (e.g. Coloptychon, Abronia, and Mesaspis), dueto incomplete taxon sampling in previous studies[129,132,133], but relationships within Gerrhonotinae areunder detailed investigation by other researchers, so theseissues are likely to be resolved in the near future.
IguaniaWe find strong support (SHL = 100) for the monophylyof Iguania (Figure 1). This clade is strongly supported bynuclear data [16,17,19,20], but an apparent episode ofconvergent molecular evolution in several mitochondrialgenes has seemingly misled some analyses of mtDNA,leading to weak support for Iguania [134], or even separ-ation of the acrodonts and pleurodonts [17,135] in previ-ous studies. Within Iguania (Figure 1), we find strongsupport (SHL = 100) for a sister-group relationship be-tween Chamaeleonidae and Agamidae (Acrodonta), andfor a clade of mostly New World families (Pleurodonta;SHL = 100).We find strong support for the monophyly of Cha-
maeleonidae and the subfamily Chamaeleoninae, andweak support for the paraphyly of Brookesiinae (Figures 1,15). The sampled species of the Brookesia nasus group ap-pear as the sister group to all other chamaeleonids (thelatter clade weakly supported) as found by some previous

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 36 of 53http://www.biomedcentral.com/1471-2148/13/93
authors [136], though other studies have recovereda monophyletic Brookesia [137,138]. Within Chamae-leoninae (Figure 15), we find strong support for the mono-phyly of most genera and species-level relationships.However, we find strong support for the non-monophylyof Calumma, with some species strongly placed withChamaeleo, others strongly placed with Rieppeleon, and athird set weakly placed with Nadzikambia + Rhampoleon.While non-monophly of Calumma has also been found inprevious studies [138], a recent study strongly supportsmonophyly of Brookesia and weakly supports monophylyof Calumma [139].Monophyly of Agamidae is strongly supported
(Figure 16; SHL = 100), contrary to some previous esti-mates [15,31]. Most relationships among agamid subfam-ilies and genera are strongly supported (Figure 16), andlargely congruent with earlier studies [17,29,34,140].There are some differences with earlier studies. For ex-ample, previous studies based on 29–44 loci [20,29,34]placed Hydrosaurinae as sister to Amphibolurinae +(Agaminae + Draconinae) with strong support, whereaswe place Hydrosaurinae as the sister-group to Amphibo-lurinae with weak support. Other authors [140] placedLeiolepiedinae with Uromastycinae, but we (and mostother studies) place Uromastycinae as the sister group toall other agamids.Our phylogeny indicates several taxonomic problems
within amphibolurine agamids (Figure 16). The generaMoloch and Chelosania render Hypsilurus paraphyletic,although the support for the relevant clades is weak.The species Lophognathus gilberti is placed in a stronglysupported clade with Chlamydosaurus and Amphibolurus,a clade that is not closely related to the otherLophognathus. Many of these taxonomic problems werealso noted by previous authors [141].Within agamine agamids (Figure 16), most relation-
ships are well supported and monophyly of all sam-pled genera is strongly supported. In contrast, withindraconine agamids (Figure 16), many intergeneric rela-tionships are weakly supported, and some genera arenon-monophyletic (Figure 16; see also [142]), includingGonocephalus (G. robinsonii is only distantly related toother Gonocephalus) and Japalura (with species distri-buted among three distantly related clades, includingone allied with Ptyctolaemus, another with G. robinsonii,and a third with Pseudocalotes).Recent authors suggested dividing Laudakia into
three genera (Stellagama, Paralaudakia, and Laudakia)based on a non-phylogenetic analysis of morphology[143]. Here, Laudakia (as previously defined) isstrongly supported as monophyletic (Figure 16), andthis change is not necessitated by the phylogeny. Simi-larly, based on genetic and morphological data, recentauthors [144] suggested resurrecting the genus Saara for
the basal clade of Uromastyx (U. asmussi, U. hardwickii,and U. loricata). However, Uromastyx (as previously de-fined) is strongly supported as monophyletic in our results(Figure 16) and in those of the recent revision [144], andthis change is not needed. We therefore retain Laudakiaand Uromastyx as previously defined, to preserve taxo-nomic stability in these groups [113]. We note that recentstudies have also begun to revise species limits in othergroups such as Trapelus [145], and taxa such as T.pallidus (Figure 16) may represent populations withinother species.Within Pleurodonta we generally confirm the mono-
phyly and composition of the clades that were ranked asfamilies (or subfamilies) within the group (e.g. Phry-nosomatidae, Opluridae, Leiosauridae, Leiocephalidae, andCorytophanidae; Figures 1, 17, 18, 19) based on previousmolecular studies [31,33,34] and earlier morphologicalanalyses [146,147].One important exception is the previously recognized
Polychrotidae. Our results confirm that Anolis andPolychrus are not sister taxa (Figures 1, 18, 19), as alsofound in some previous molecular studies [31,33,34], butnot others [20,29]. Our results provide strong supportfor non-monophyly of Polychrotidae, placing Polychruswith Hoplocercidae (SHL = 99) and Anolis withCorytophanidae (SHL = 99; the latter also found by[34]). Recent analyses placing Anolis with Polychrusshowed only weak support for this relationship [20,29],despite many loci (30–44). We support continued rec-ognition of Dactyloidae for Anolis and Polychrotidaefor Polychrus [34], based on a limited number of locibut extensive taxon sampling. We note that these fa-milies are still monophlyetic, even if they prove to be sistertaxa.Interestingly, our results for relationships among pleu-
rodont families differ from most previous studies, and aresurprisingly well-supported in some cases by SHL values(but see below). In previous studies, many relationshipsamong pleurodont families were poorly supported byBayesian posterior probabilities and by parsimony andlikelihood bootstrap values, though typically samplingfewer taxa or characters [17,31,33,148-151]. Studies in-cluding 29 nuclear loci found strong concatenated Bayes-ian support for many relationships but weak support fromML bootstrap analyses for many of the same relationships[34]. The latter pattern (typically weak ML support) wasalso found in an analysis including those same 29 loci andmitochondrial data for >150 species [29]. We also find amixture of strongly and weakly supported clades, but withmany relationships that are incongruent with these previ-ous studies. First, we find that Tropiduridae is weakly sup-ported as the sister group to all other pleurodonts (alsofound by [29]), followed successively (Figures 1, 17, 18, 19)by Iguanidae, Leiocephalidae, Crotaphytidae + Phrynoso-

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 37 of 53http://www.biomedcentral.com/1471-2148/13/93
matidae, Polychrotidae + Hoplocercidae, and Coryto-phanidae + Dactyloidae.The relatively strong support for the clades
Crotaphytidae + Phrynosomatidae (SHL = 87), Poly-chrotidae + Hoplocercidae (SHL = 99), and Coryto-phanidae + Dactyloidae (SHL = 99) is largelyunprecedented in previous studies (although Coryto-phanidae + Dactyloidae is strongly supported in someBayesian analyses [34]). As in many previous analyses,deeper relationships among the families remain weaklysupported. We also find a strongly supported cladecontaining Liolaemidae, Opluridae, and Leiosauridae(SHL = 95), with Opluridae + Leiosauridae also stronglysupported (SHL = 99). Both clades have also been foundin previous studies [149,151], including studies based on29 or more nuclear loci [20,29,34].We note that previous studies have shown strong sup-
port for some pleurodont relationships (e.g. basal place-ment of phrynosomatids; see [34]), only to be stronglyoverturned with additional data [20,29]. Therefore, the re-lationships found here should be taken with some caution(even if strongly supported), with the possible exceptionof the recurring clade of Liolaemidae + (Opluridae +Leiosauridae).All pleurodont families are strongly supported as mono-
phyletic (SHL > 85). Within the pleurodont families, ourresults generally support the current generic-level ta-xonomy (Figures 17, 18, 19). However, there are some ex-ceptions. Within Tropiduridae (Figure 17), Tropidurus isparaphyletic with respect to Eurolophosaurus, Strobilurus,Uracentron, and Plica. Within Opluridae (Figure 18), themonotypic genus Chalarodon renders Oplurus para-phyletic. Two leiosaurid genera are also problematic(Figure 18). In Enyaliinae, Anisolepis is paraphyletic withrespect to Urostrophus, and this clade is nested withinEnyalius. In Leiosaurinae, Pristidactylus is rendered para-phyletic by Leiosaurus and Diplolaemus (Figure 18).Within Dactyloidae, a recent study re-introduced a
more subdivided classification of anoles [152], an issuethat has been debated extensively in the past [153-156].Our results support the monophyly of all the genera rec-ognized by recent authors [152], including Anolis,Audantia, Chamaelinorops, Dactyloa, Deiroptyx, Norops,and Xiphosurus (see Figure 19). However, since Anolis ismonophyletic as traditionally defined, we retain that def-inition here (including the seven listed genera) for con-tinuity with the recent literature [113,157].
SerpentesRelationships among the major serpent groups (Figure 1)are generally similar to other recent studies [20,35,36,38,41,44,47,158-160]. We find that the blindsnakes, Sco-lecophidia (Figures 1, 20) are not monophyletic, as inprevious studies [19,20,36,44,159,160]. Similar to some
previous studies [44,159], our data weakly place Ano-malepididae as the sister taxon to all snakes, and thescolecophidian families Gerrhopilidae, Leptotyphlopidae,Typhlopidae, and Xenotyphlopidae as the sister-group toall other snakes excluding Anomalepididae (Figures 1, 20).Previous studies have also placed Anomalepididaeas the sister-group to all non-scolecophidian snakes[19,20,35,36,158,160], in some cases with strong sup-port [20]. Although it might appear that recent ana-lyses of scolecophidian relationships [38] supportmonophyly of Scolecophidia (e.g. Figure 1 of [38]), thetree including non-snake outgroups from that studyshows weak support for placing anomalepidids withalethinophidians, as in other studies [19,20,36,160].We follow recent authors [38] in recognizing Xeno-
typhlopidae (strongly placed as the sister taxon ofTyphlopidae) and Gerrhopilidae (strongly placed as thesister group of Xenotyphlopidae + Typhlopidae) as dis-tinct families (Figure 20). Leptotyphlopidae is stronglysupported as the sister group of a clade compri-sing Gerrhopilidae, Xenotyphlopidae, and Typhlopidae(Figure 20). As in previous studies [38,44], we find strongsupport for non-monophyly of several typhlopid genera(Afrotyphlops, Austrotyphlops, Ramphotyphlops, Letheo-bia, and Typhlops; Figure 20). There are also undescribedtaxa (e.g. Typhlopidae sp. from Sri Lanka; [44]) of uncer-tain placement within this group (Figure 20). The syste-matics of typhlopoid snakes will thus require extensiverevision in the future, with additional taxon and charactersampling.Within Alethinophidia (SHL = 100), Aniliidae is
strongly supported (SHL = 98) as the sister taxon ofTropidophiidae (together comprising Anilioidea), and allother alethinophidians form a strongly supported sistergroup to this clade (SHL = 97; Figures 1, 21). The enig-matic family Xenophidiidae is weakly placed as the sister-group to all alethinophidians exclusive of Anilioidea(Figures 1, 21). The family Bolyeriidae is weakly placedas the sister-group to pythons, boas, and relatives(Booidea), which are strongly supported (SHL = 88). Re-lationships in this group are generally consistent withother recent molecular studies [20,35-37,47,159].Relationships among other alethinophidians are a mix-
ture of strongly and weakly supported nodes (Figures 1,21). We find strong support (SHL = 100) for a cladecontaining Anomochilidae + Cylindrophiidae + Uro-peltidae. This clade of three families is strongly supported(SHL = 89) as the sister taxon to Xenopeltidae +(Loxocemidae + Pythonidae). Together, these six familiesform a strongly supported clade (SHL = 89; Figures 1, 21)that is weakly supported as the sister group to the stronglysupported clade of Boidae + Calabariidae. Within theclade of Anomochilidae, Cylindrophiidae, and Uropeltidae(Figure 21), we weakly place Anomochilus as the sister

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 38 of 53http://www.biomedcentral.com/1471-2148/13/93
group to Cylindrophiidae [44], in contrast to previousstudies which placed Anomochilus within Cylindrophis[161]. However, support for monophyly of Cylindrophisexcluding Anomochilus is weak (Figure 21). As in previousstudies [44,162], we find several taxonomic problemswithin Uropeltidae (Figure 21). Specifically, Rhinophis andUropeltis are paraphyletic with respect to each other andto Pseudotyphlops. The problematic taxa are primarily SriLankan [44], and forthcoming analyses will address theseissues.Within Pythonidae (Figure 21), the genus Python is
the sister group to all other genera. Some species thatwere traditionally referred to as Python (P. reticulatusand P. timoriensis) are instead sister to an Australasianclade consisting of Antaresia, Apodora, Aspidites,Bothrochilus, Leiopython, Liasis, and Morelia (Figure 21).These taxa (P. reticulatus and P. timoriensis) have beenreferred to as Broghammerus, a name originating froman act of "taxonomic vandalism" (i.e. an apparentlyintentional attempt to disrupt stable taxonomy) in anon-peer reviewed organ without data or analyses[163,164]. However, this name was, perhaps inadvert-ently, subsequently used by researchers in peer-reviewedwork [165] and has entered into somewhat widespreadusage [1]. This name should be ignored and replacedwith a suitable substitute. Within the Australasian clade(Figure 21), Morelia is paraphyletic with respect to allother genera, and Liasis is non-monophyletic with re-spect to Apodora, although many of the relevant rela-tionships are weakly supported.Within Boidae (Figure 21), our results and those of
other recent studies [20,36,47,48,150,166] have con-verged on estimated relationships that are generallysimilar to each other but which differ from traditionaltaxonomy [167]. However, the classification has yet tobe modified to reflect this, and we rectify this situationhere. We find that Calabariidae is nested within Boidae[150], but this is poorly supported, and contrary to mostprevious studies [47,48]. While Calabaria has been clas-sified as an erycine boid in the past, this placement isstrongly rejected here and in other studies [47,48]. If thecurrent placement of Calabaria is supported in thefuture, it would require recognition as the subfamilyCalabariinae in Boidae.The Malagasy boine genera Acrantophis and Sanzinia
are placed as the sister taxa to a weakly-supported cladecontaining Calabariidae and a strongly supported clade(SHL = 99) comprising the currently recognized sub-families Erycinae, Ungaliophiinae, and other boines(Figure 21). Regardless of the position of Calabariidae,this placement of Malagasy boines renders Boinae para-phyletic. We therefore resurrect the subfamily San-ziniinae [168] for Acrantophis and Sanzinia. Thissubfamily could be recognized as a distinct family if
future studies also support placement of this clade asdistinct from other Boidae + Calabariidae.The genera Lichanura and Charina are currently clas-
sified as erycines [1], but are strongly supported as thesister group to Ungaliophiinae, as in previous studies[20,36,47,166]. We expand Ungaliophiinae to includethese two genera (Figure 21), rather than erect a newsubfamily for these taxa. The subfamily Ungaliophiinaeis placed as the sister group to a well-supported clade(SHL = 87) containing the rest of the traditionally recog-nized Erycinae and Boinae. We restrict Erycinae to theOld World genus Eryx.The genus Candoia (Boinae) from Oceania and New
Guinea [1], is placed as the sister taxon to a moderatelysupported clade (SHL = 83) consisting of Erycinae (Eryx)and the remaining genera of Boinae (Boa, Corallus,Epicrates, and Eunectes). To solve the non-monophyly ofBoinae with respect to Erycinae (due to Candoia), weplace Candoia in a new subfamily (Candoiinae, subfam.nov.; see Appendix I). Boinae then comprises the fourNeotropical genera that have traditionally been classifiedin this group (Boa, Corallus, Epicrates, and Eunectes).We acknowledge that non-monophyly of Boinae couldbe resolved in other ways (e.g. expanding it to includeErycinae). However, our taxonomy maintains the trad-itionally used subfamilies Boinae, Erycinae, and Ungalio-phiinae, modifies them to reflect the phylogeny, andrecognizes the phylogenetically distinct boine clades asseparate subfamilies (Candoiinae, Sanziniinae). WithinBoinae, Eunectes renders Epicrates paraphyletic, but thisis not strongly supported (see also [48]).Our results for advanced snakes (Caenophidia) are gen-
erally similar to those of other recent studies [41,42,169],and will only be briefly described. However, in contrast tomost recent studies [20,36,41,42,81,159,160], Acrochor-didae is here strongly placed (SHL = 95) as the sistergroup to Xenodermatidae. This clade is then the sistergroup to the remaining Colubroidea, which form astrongly supported clade (SHL = 100; Figures 1, 22). Thisrelationship has been found in some previous studies[169,170], and was hypothesized by early authors [171].Further evidence will be required to resolve this conclu-sively. Analyses based on concatenation of 20–44 loci donot support this grouping [20,36], though preliminaryspecies-tree analyses of >400 loci do (Pyron et al., inprep.). Relationships in Pareatidae are similar to recentstudies [172], and the group is strongly placed as the sistertaxon to colubroids excluding xenodermatids (SHL = 100;Figures 1 , 22), as in most recent analyses (e.g. [41,43,44]).The family Viperidae is the sister group to all colubroids
excluding xenodermatids and pareatids (Figure 1), as inother recent studies. The family Viperidae is strongly sup-ported (Figure 22), as is the subfamily Viperinae, and thesister-group relationship between Azemiopinae and Cro-

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 39 of 53http://www.biomedcentral.com/1471-2148/13/93
talinae (SHL = 100). Our results generally supportthe existing generic-level taxonomy within Viperinae(Figure 22). However, we recover a strongly supportedclade within Viperinae consisting of Daboia russelii, D.palaestinae, Macrovipera mauritanica, and M. deserti(Figure 22), as in previous studies [173]. We corroborateprevious suggestions that these taxa be included in Daboia[174], though this has not been widely adopted [1]. Theother Macrovipera species (including the type species) re-main in that genus (Figure 22).Within Crotalinae (Figure 22), a number of genera ap-
pear to be non-monophyletic. The species Trimeresurusgracilis is strongly supported as the sister taxon to Ovophisokinavensis and distantly related to other Trimeresurus,whereas the other Ovophis are strongly placed as the sistergroup to Protobothrops. A well-supported clade (SHL = 90)containing Atropoides picadoi, Cerrophidion, and Por-thidium renders Atropoides paraphyletic (see also [175]).The species Bothrops pictus, considered incertae sedis inprevious studies [176], is here strongly supported as thesister taxon to a clade containing Rhinocerophis, Bothro-poides, Bothriopsis, and Bothrops (Figure 22). Most of theserelationships are strongly supported.Viperidae is strongly placed (SHL = 95) as the sister
taxon to a well-supported clade (SHL = 100) containingColubridae, Elapidae, Homalopsidae, and Lamprophiidae(Figure 1). Monophyly of Homalopsidae is also stronglysupported (Figure 23). Within Homalopsidae, non-monophyly of the genus Enhydris is strongly supported(Figure 23), and it should likely be split into multiplegenera. Homalopsidae is weakly supported (SHL = 58) asthe sister group of Elapidae + Lamprophiidae (Figure 1).This same relationship was also weakly supported by pre-vious analyses [41,44], but other studies have foundstrong support for placing Homalopsidae as the sistergroup of a strongly supported clade including Elapidae,Lamprophiidae, and Colubridae [20,36], including datafrom >400 loci (Pyron et al., in prep.).Support for the monophyly of Lamprophiidae is
strong (but excluding Micrelaps; see below), and mostof its subfamilies are well-supported [40,41,177,178]including Atractaspidinae, Aparallactinae, Lamprophiinae,Prosymninae (weakly placed as the sister-group toOxyrhabdium), Pseudaspidinae, Psammophiinae, andPseudoxyrhophiinae (Figure 23). In Lamprophiidae,most genera are monophyletic based on our sam-pling (Figure 23). However, within Aparallactinae,Xenocalamus is strongly placed within Amblyodipsas,and in Atractaspidinae, Homoroselaps is weaklyplaced in Atractaspis. In Lamprophiinae, Lamprophisis paraphyletic with respect to Lycodonomorphus butsupport for the relevant clades is weak.The enigmatic genera Buhoma from Africa and Psam-
modynastes from Asia were both previously considered
incertae sedis within Lamprophiidae [41]. Here they areweakly placed as sister taxa, and more importantly, theyform a strongly supported clade with the African genusPseudaspis (Pseudaspidinae; SHL = 95; Figure 23). There-fore, we expand Pseudaspidinae to include these twogenera.The genus Micrelaps (putatively an aparallactine; [1])
is weakly placed as the sister taxon to Lamprophiidae +Elapidae. Along with Oxyrhabdium (see above) andMontaspis [40], this genus is treated as incertae sedis inour classification (Appendix I). If future studies stronglysupport these relationships, they may require a new fam-ily for Micrelaps and possibly a new subfamily forOxyrhabdium, though placement of these taxa has beenhighly variable in previous studies [40,41,44,81].Monophyly of Elapidae is strongly supported (Figure 24),
and Calliophis melanurus is strongly supported as the sis-ter group to all other elapids (see also [44]). WithinElapidae (Figure 24), relationships are generally concord-ant with previous taxonomy, with some exceptions. Thegenera Toxicocalamus, Simoselaps, and Echiopsis are alldivided across multiple clades, with strong support formany of the relevant branches. A recent study [46] hasprovided a generic re-classification of the sea snakes(Hydrophis group) to resolve the extensive paraphyly ofgenera found in previous studies (e.g. [179,180]). Our re-sults support this classification.Monophyly of Colubridae and most of its subfamilies
(sensu [41,44]) are strongly supported (Figures 1, 25, 26,27, 28). However, relationships among many of thesesubfamilies are weakly supported (Figure 1), as in mostprevious studies [41,43,45,181]. The subfamilies Calama-riinae and Pseudoxenodontinae are strongly supportedas sister taxa, and weakly placed as the sister-group tothe rest of Colubridae (Figure 25). There is a weaklysupported clade (Figure 1) comprising Natricinae +(Dipsadinae + Thermophis), but the clade uniting the NewWorld Dipsadinae with the Asian genus Thermophis isstrongly supported (SHL = 100; Figure 28). Here, we placeThermophis (recently in Pseudoxenodontinae [41]) inDipsadinae (following [182]), making it the first and onlyAsian member of this otherwise exclusively New Worldsubfamily. However, despite the strong support for itsplacement here, placement of this taxon has been variablein previous analyses [41,182,183], and we acknowledgethat future analyses may support recognition of a distinctsubfamily (Thermophiinae).The clade of Natricinae and Dipsadinae is weakly sup-
ported as the sister group (Figures 1, 25, 26, 27, 28) to aclade containing Sibynophiinae [181] + (Colubrinae +Grayiinae). The subfamily Colubrinae is weakly sup-ported; we find that the colubrine genera Ahaetulla,Chrysopelea, and Dendrelaphis form a strongly sup-ported clade that is weakly placed as the sister group to

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 40 of 53http://www.biomedcentral.com/1471-2148/13/93
the rest of Colubrinae, which form a strongly supportedclade (Figure 25). This clade was also placed withGrayiinae or Sibynophiinae in many preliminary ana-lyses, rendering Colubrinae paraphyletic. This group ofthree genera has been strongly supported in the past,and only weakly placed with Colubrinae [41,44]. It ispossible that future analyses will reveal that the clade ofAhaetulla, Chrysopelea, and Dendrelaphis is placed else-where in Colubridae with strong support, and thus meritrecognition as a distinct subfamily (Ahaetuliinae). A not-able feature of this clade is the presence of gliding flightin most species of Chrysopelea, less-developed non-flight jumping with similar locomotor origins inDendrelaphis, and homologous glide-related traits inAhaetulla [184].Numerous colubroid genera are not included in our
tree and are not clearly placed in subfamilies based onprevious morphological evidence. In our classification,these genera are also considered incertae sedis withinColubridae, including Blythia, Cyclocorus, Elapoidis,Gongylosoma, Helophis, Myersophis, Oreocalamus, Poeci-lopholis, Rhabdops, and Tetralepis, as in previous classi-fications [1].Our phylogeny reveals numerous taxonomic problems
within Colubrinae (Figures 25, 26). The genus Boiga isparaphyletic with respect to Crotaphopeltis, Dipsadoboa,Telescopus, Toxicodryas, and Dasypeltis, with strongsupport (Figure 25). The genus Philothamnus is para-phyletic with respect to Hapsidophrys (Figure 25). Thegenus Coluber is split between Old World and NewWorld clades (Figures 25, 26). The species Hierophisspinalis is sister to Eirenis to the exclusion of the otherHierophis species (Figure 25). The genus Dryocalamus isnested within Lycodon (Figure 26). The species Chironiuscarinatus and C. quadricarinatus are weakly placed in aclade of Neotropical colubrines only distantly relatedto the other Chironius species (Figure 26). The ge-nus Drymobius renders Dendrophidion paraphyletic(Figure 26). The monotypic genus Rhynchophis rendersthe two species of Rhadinophis paraphyletic (Figure 26).The genus Coronella is rendered paraphyletic (Figure 26)by Oocatochus with weak support (see also [185]). Finally,the genus Rhinechis is nested within Zamenis (Figure 26).We find numerous non-monophyletic genera within
Natricinae (Figure 27), as in previous studies [41,44,78,186].These non-monophyletic genera include the Asian generaAmphiesma, Atretium, and Xenocrophis. Among NewWorld genera, we find Regina to be non-monophyleticwith respect to most other genera, as in previous phylo-genetic studies (e.g. [41,186]). Also, as in previous studies(e.g. [41,187]), we find that Adelophis [188] is nested deepwithin Thamnophis [189].Finally, within a weakly supported Dipsadinae
(Figure 28), we find non-monophyly of numerous genera,
as in many earlier studies (e.g. [41-43,190]). These prob-lems of non-monophyly include Leptodeira (with respectto Imantodes), Geophis (with respect to Atractus),Atractus (with respect to Geophis), Sibynomorphus (withrespect to Dipsas), Dipsas (with respect to Sibynomor-phus), Taeniophallus (with respect to Echinanthera), andEchinanthera (with respect to Taeniophallus). Recent revi-sions have begun to tackle these problems [42,43,190], butadditional taxon and character sampling will be crucial toresolve relationships and taxonomy.
DiscussionIn this study, we provide a phylogenetic estimate for4161 species of squamates based on molecular data fromup to 12 genes per species, combining much of the re-levant data used in previous molecular phylogeneticanalyses. This tree provides a framework for future evo-lutionary studies, spanning from the species level to rela-tionships among families, utilizing a common set ofbranch lengths. These estimated branch lengths are crit-ically important for most phylogenetic comparativemethods. To further facilitate use of this phylogeny incomparative studies, we provide the Newick version ofthis tree (with estimated branch lengths) in DataDryadrepository 10.5061/dryad.82h0m and Additional file 1:Data File S1. Our results also suggest that the branchlengths in this tree should not generally be compromisedby missing data for some genes in some taxa.Our results also reveal many problems in squamate
classification at nearly all phylogenetic levels. We makeseveral changes to higher-level taxonomy based on thisphylogeny, including changes to the traditionally re-cognized subfamilies of boid snakes (i.e. resurrectingSanziniinae for the boine genera Acrantophis andSanzinia, erecting Candoiinae for the boine genusCandoia, and moving Lichanura and Charina fromErycinae to Ungaliophiinae), lamprophiid snakes (expan-sion of Pseudaspidinae to include the formerly incertaesedis genera Buhoma and Psammodynastes), colubridsnakes (expansion of Dipsadinae to include the Asianpseudoxenodontine genus Thermophis), and gymno-phthalmid lizards (recognition of Bachiinae for the tribeBachiini, containing Bachia) and scincid lizards (synony-mizing Feylininae with Scincinae to yield a total of threescincid subfamilies: Acontiinae, Lygosominae, andScincinae). In Appendix I, we list the generic content ofall families and subfamilies. We also find dozens of prob-lems at the genus level, many of which have been identi-fied previously, and which we defer the resolution of tofuture studies. Our results also highlight potential prob-lems in recent proposals to modify the classification ofscincid [112] and teiid lizards [123].In addition to synthesizing existing molecular data for
squamate phylogeny, our analyses also reveal several

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 41 of 53http://www.biomedcentral.com/1471-2148/13/93
apparently novel findings (Figures 1, 2, 3, 4, 5, 6, 7, 8, 9,10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24,25, 26, 27, 28). Given space constraints, we cannot detailevery deviation from previous phylogenetic hypotheses(especially pre-molecular studies). Nevertheless, wefocus on three sets of examples. First, we find some rela-tively novel, strongly-supported relationships at the fam-ily level. These include the placement of Helodermatidae(as sister to Xenosauridae, Anguidae, and Anniellidae)and the placement of Xenodermatidae as the sister taxonto Acrochordidae (rendering Colubroidea paraphyletic),in contrast to most recent analyses of anguimorphs andsnakes (see above). We also find some novel, stronglysupported relationships among pleurodont families, butwe acknowledge that these may be overturned in futurestudies.The second example is the higher-level relationships
within Scincidae, the largest family of lizards [1]. Noprevious studies examining higher-level relationshipswithin the group included more than ~50 species[50,51]. In this study, we sample 683 skink species(Figures 6, 7, 8, 9, 10), and our phylogeny provides aunique resolution of higher-level skink relationships.Some previous researchers [51] placed acontiines as thesister group to all other skinks, but suggested thatscincines and lygosomines were paraphyletic with respectto each other (with feyliniines placed with scincines). Incontrast, others [50] suggested that acontiines, feyliniines,and lygosomines were all nested inside scincines (but witheach of those three subfamilies as monophyletic), althoughmany clades were only weakly supported. Here (Figures 1,6, 7, 8, 9, 10), we find that acontiines are the sister groupto a strongly supported clade consisting of a monophyleticScincinae and a monophyletic Lygosominae (exceptingthe weakly supported placement of Ateuchosaurus andplacement of Feyliniinae in Scincinae).Third, our phylogeny reveals numerous genera that ap-
pear to be non-monophyletic, with many of these caseshaving strong support for the associated nodes. Our exam-ples include genera in many families, including dibamids,diplodactylids, gekkonids, phyllodactylids, gerrhosaurids,scincids, teiids, gymnophthalmids, lacertids, anguids, cha-maeleonids, agamids, tropidurids, oplurids, leiosaurids,typhlopids, pythonids, uropeltids, boids, viperids, lam-prophiids, elapids, and colubrids (see Results). Althoughmany problems noted here were found in previous studies,some seem to be new, such as placement of Crocodilurusand Draceana within Tupinambis (in Teiidae; Figure 11)and Coloptychon within Gerrhonotus (in Anguidae;Figure 14).Our study also offers an important test of higher-level
squamate relationships using a very different samplingstrategy than that used in most previous analyses. Squa-mates have traditionally been divided into two clades based
on morphology, Iguania and Scleroglossa (e.g. [13,21]).Despite considerable disagreement among morphology-based hypotheses, this basic division is supported bynearly all phylogenetic analyses based on morphologicaldata [13-15,19,22,95,191]. In contrast, our results andthose of most previous molecular analyses strongly sup-port placement of iguanians with anguimorphs and snakes[16-20,23,24]. The causes of this conflict remain unclear,but may be related to morphological traits associated withdifferent feeding strategies of iguanian and (traditional)scleroglossan squamates [3,17].Additionally, analyses of morphology often place
dibamids, amphisbaenians, snakes, and (in some cases)some scincids and anguids in a single clade [13-15]. Ouranalyses do not support such a clade (Figure 1), nor doother analyses of molecular data alone [17-20], or ana-lyses of combined molecular and morphological data[19]. Instead, these morphological results seem to beexplained by convergence associated with burrowing(e.g. [19]). Overall, molecular datasets have shown over-whelmingly strong support for placement of dibamidsand gekkonids at the base of the tree, amphisbaenianswith lacertoids, and iguanians with snakes and angui-morphs [17-20,23]. These results have now been corro-borated with up to 22 genes (15794 bp) for 45 taxa [19],25 genes (19020 bp) for 64 taxa [16], and 44 genes(33717 bp) for 161 taxa [20]. We now support this basictopology with 4161 species sampled for up to 12 geneseach (up to 12896 bp).Nevertheless, despite the overall strong support for most
of the tree (i.e. 70% of all nodes have SHL > 85), certainclades remain poorly supported (e.g. relationships amongmany pleurodont iguanian families; Figures 1, 17, 18, 19).A potential criticism of the supermatrix approach usedhere is that this weak support may occur due to missingdata. However, previous studies of 8 datasets have shownexplicitly that there is typically little relationship betweenbranch support for terminal taxa and the amount of miss-ing data [85]. Instead, these patterns of weak support aremore likely to reflect short underlying branch lengths[20,36,41], and may be difficult to resolve even with morecomplete taxonomic and genomic sampling. Indeed, asnoted above, many of the weakly supported nodes in ourphylogeny are also weakly supported in analyses with littlemissing data (<20%) and large numbers of genes (e.g. 44genes as in [20]), such as the relationships of manypleurodont lizard families and colubroid snake familiesand subfamilies.We acknowledge that the differences between our re-
sults and previous studies (noted above) do not necessarilymean that our results are right and those of previous stud-ies are wrong. In some cases, we provide strong supportfor novel relationships when previous, conflicting studiesshowed only weak support (as in scincids, see above). In

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 42 of 53http://www.biomedcentral.com/1471-2148/13/93
other cases, our results disagree with other studies forclades that were strongly supported (e.g. placement ofxenodermatids). In the best-case scenario, these conflictsmay be resolved because our results are correct, possiblyreflecting the beneficial effects of adding taxa and theassociated subdivision of long branches [25,26,28,87,192-194]. Furthermore, in many cases, we are includingmore genes than used in previous studies of particularclades, increasing sampling of characters and loci. Thisshould generally reduce spurious results caused by sam-pling few characters and by incongruence between geneand species trees.However, other explanations for incongruence between
our results and previous studies are also possible.Adding taxa can potentially lead to incorrect results insome cases (e.g. [195]), such as when a long terminalbranch is added that further subdivides a short internalbranch. In other cases, conflicts with our results mightreflect the impact of our sampling fewer nuclear genesand a correspondingly increased influence of mitochon-drial data. Mitochondrial genes have relatively fast evo-lutionary rates (potentially exacerbating the impacts oflong branches), and their phylogenetic resolution for aparticular node may also reflect introgression or incom-plete lineage sorting rather than the species phylogeny(review in [196]). Many taxa in the matrix are repre-sented only by mitochondrial data, and highly variablemitochondrial genes might also overcome the influenceof less variable nuclear genes in combined analyses(although this scenario does not seem to be common[196]). Such cases might explain some strongly sup-ported conflicts between our results and those based onmultiple nuclear loci. Another possibility is that somecases may represent failure to find the optimal tree(although we assume that these cases will likely showonly weak support). We acknowledge that there aremany reasons why our results may differ from previousstudies, and the ultimate test of these novel findings willbe corroboration in future studies that include moretaxa and characters.This analysis also corroborates several recent studies
suggesting that the supermatrix approach is a powerfulstrategy for large-scale phylogenetic inference [41,72,73,75,76,197]. For example, even though each specieshad 81% missing data on average, we found that mostspecies were placed in the families and genera expectedbased on previous taxonomy, often with very strong sup-port. Furthermore, we found that incompleteness of ter-minal taxa is not related to branch lengths (at least notterminal branch lengths), suggesting that missing dataare not significantly biasing branch-length estimates (seealso [84,86,87]). Also, the ML models we used have beenshown to be robust to missing data in large, sparsesupermatrices [84].
Even though we did find some subfamilies and generato be non-monophyletic, similar relationships were oftenfound in previous studies based on data matrices withonly limited missing data (e.g. non-monophyly of boidsnake subfamilies [47], lacertid and scincid lizard genera[67,111], and scolecophidian, dipsadine, and natricinesnake genera [38,43,186]). We suggest that further reso-lution of the squamate tree will be greatly facilitated ifresearchers deliberately sample mitochondrial genes andnuclear genes that include the set of genes used hereand in recent phylogenomic studies (e.g. [20]), to in-crease overlap between genes and taxa, and decreasemissing data.With over 5000 species remaining to be included and
only 12 genes sampled, our study is far from the lastword on squamate phylogeny. We note that new datacan easily be added to this matrix, in terms of both newtaxa and new genes. Increased sampling of other nucleargenes is likely to be advantageous as well. Next-generationsequencing strategies and phylogenomic methods shouldhelp resolve difficult nodes [16,20,36,198-200], as shouldapplication of species-tree methods [201,202]. Species-treeanalyses of 44 nuclear loci support many of these sameclades across squamates [20], and data from >400 nuclearloci reinforces many of the relationships found hereamong the colubroid snake subfamilies (Pyron et al., inprep.). In addition, it would be useful to incorporate fossiltaxa in future studies [15,19,22,86], utilizing the large mor-phological datasets that are now available [14,15]. Despitethese areas for future studies, the present tree providesa framework for researchers analyzing patterns of squa-mate evolution at both lower and higher taxonomiclevels (e.g. [10,11,203,204]), and for building a morecomplete picture of squamate phylogeny.
ConclusionsIn this study, we provide a phylogenetic estimate for4161 squamate species, based on a supermatrix ap-proach. Our results provide important confirmation forprevious studies based on more limited taxon sampling,and reveal new relationships at the level of families, gen-era, and species. We also provide a phylogenetic frame-work for future comparative studies, with a large-scaletree including a common set of estimated branchlengths. Finally, we provide a revised classification forsquamates based on this tree, including changes in thehigher-level taxonomy of gymnophthalmid and scincidlizards and boid, colubrid, and lamprophiid snakes.
MethodsInitial classificationOur initial squamate classification is based on the June2009 version of the Reptile Database [1] (http://www.reptile-database.org/), accessed in September of 2009

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 43 of 53http://www.biomedcentral.com/1471-2148/13/93
when this research was begun. Minor modifications tothis scheme were made, primarily to update changes incolubroid snake taxonomy [41-44,205]. This initial taxo-nomic database consists of 8650 species (169 amp-hisbaenians, 5270 lizards, 3209 snakes, and 2 tuataras),against which the classification of species in the molecularsequence database was fixed. While modifications and up-dates (i.e. new species, revisions) have been made to squa-mate taxonomy subsequently, these are minor and shouldhave no impact on our phylogenetic results. This databaserepresents ~92% of the current estimated diversity ofsquamates (~9400 species as of December 2012).Throughout the paper, we refer to the updated version
of squamate taxonomy from the December 2012 updateof the Reptile Database [1], incorporating major, well-accepted changes from recent studies (summarized in[1]). However, for large, taxonomic groups that have re-cently been broken up for reasons other than resolvingparaphyly or matters of priority (e.g. in dactyloid andscincid lizards; see Results), we generally retain the older,more inclusive name in the interest of clarity, while pro-viding references to the recent revision. We attempt toalter existing classifications as little as possible (see also[113]). Therefore, we generally only make changes whenthere is strong support for non-monophyly of currentlyrecognized taxa and our proposed changes yield stronglysupported monophyletic groups. Similarly, we only erectnew taxa if they are strongly supported. Finally, althoughnumerous genera are identified as being non-monophyleticin our tree, we refrain from changing genus-level tax-onomy, given that our taxon sampling within many generais limited.
Molecular dataPreliminary literature searches were conducted to iden-tify candidate genes for which a substantial number ofsquamate species were sequenced and available onGenBank (with the sampled species spread across mul-tiple families), and which were demonstrably useful inprevious phylogenetic studies of squamates (see Intro-duction for references). Twelve genes were identified asmeeting these criteria: seven nuclear genes (brain-derivedneurotrophic factor [BDNF], oocyte maturation factor[c-mos], neurotrophin-3 [NT3], phosducin [PDC], Gprotein-coupled receptor 35 [R35], and recombination-activating genes 1 [RAG-1] and 2 [RAG-2]); and fivemitochondrial genes (12S/16S long and short subunitRNAs, cytochrome b [cyt-b], and nicotinamide adeninedehydrogenase subunits 2 [ND2] and 4 [ND4]). Thissampling of genes does not include all available markers.For example, we omitted several nuclear and mitochon-drial genes because they were available only for a limitedsubset of taxa. We also excluded tRNAs associated with
the protein-coding sequences, given their short lengthsand difficulty in alignment across the large time scalesconsidered here.To ensure maximal taxonomic coverage from the avail-
able data, searches were conducted on GenBank by family(stopping in October 2012), and the longest sequence forevery species was gathered. Sequences totaling less than250 bp for any species were not included. Only species inthe taxonomic database were included in the sequencematrix, which resulted in the exclusion of numerousnamed taxa of ambiguous status, a few taxa described veryrecently, and many sequences labeled 'sp.' Some recentlydescribed phylogeographic lineages were also omitted.Species and GenBank accession numbers are available inAdditional file 2: Table S1.With respect to the December 2012 update of the
Reptile Database [1], we sampled 52 of 183 amphisbae-nian species (28%) from 11 of 19 (58%) genera; 2847 of5799 lizard species (49%) from 448 of 499 genera (90%);and 1262 of 3434 snake species (39%) in 396 of 500 gen-era (80%). This yielded a total of 4161 species in 855genera in the final matrix, 44% of the 9416 known, ex-tant squamate species in 84% of 1018 genera [1]. Thespecies-level classification of squamates is in constantflux, and the numbers of species and genera changedeven as this paper was under review. The extant speciesof tuatara (Sphenodon punctatus) was included as a non-squamate outgroup taxon (see below). We acknowledgethat our sampling of outgroup taxa is not extensive.However, placement of Sphenodon as the sister group tosquamates is well-established by molecular analyses withextensive taxon sampling (e.g. [16,128,206]) and mor-phological data (e.g. [13]).Alignment for protein-coding sequences was relatively
straightforward. We converted them to amino acids, andthen used the translation alignment algorithm in theprogram Geneious v4.8.4 (GeneMatters Corp.), with thedefault cost matrix (Blosum62) and gap penalties(open=12, extension=3). Alignments were relativelyunambiguous after being trimmed for quality and max-imum coverage (i.e. ambiguous end regions were re-moved, and most sequences began and ended at thesame point).For the ribosomal RNA sequences (12S and 16S se-
quences), alignment was more challenging. Preliminaryglobal alignments using the algorithms MUSCLE [207]and CLUSTAL [208] under a variety of gap-cost parame-ters yielded low-quality results (i.e. alignments with largenumbers of gaps and little overlap of potentially hom-ologous characters). We subsequently employed a two-step strategy for these data. We first grouped sequencesby higher taxa (i.e. Amphisbaenia, Anguimorpha, Gek-kota, Iguania, Scincomorpha, and Serpentes, thoughthese are not all monophyletic as previously defined), for

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 44 of 53http://www.biomedcentral.com/1471-2148/13/93
which alignments were relatively straightforward underthe default MUSCLE parameters.These were then combined using the profile alignment
feature of MUSCLE, and the global alignment was sub-sequently updated using the "refine alignment" option.Minor adjustments were then made by eye, and ambigu-ously aligned end-regions were trimmed for maximumcoverage and quality. We did not include partitions forstems and loops for the ribosomal sequences, althoughthis has been shown to improve model fit in previoussquamate studies (e.g. [82]). Although it is possible toassign individual nucleotide positions to these partitions,this would have been challenging given the large numberof sequences, and the potential for stems and loops toshift across the many species and large time scalesinvolved.Each species was represented by a single terminal
taxon in the matrix. In many cases, sequences from mul-tiple individuals of the same species were combined, toallow us to combine data from different genes for thesame species. We acknowledge the possibility that insome cases this approach may cause us to combinegenes from different species in the same terminal taxon(e.g. due to changing taxonomy or incorrect identifica-tions). Additionally, many sequences are not fromvouchered specimens, and it is possible that misidenti-fied species are present on GenBank and in our matrix.However, most of our data came from lower-level phylo-genetic studies, in which the identification of species byprevious authors should be highly accurate. In addition,any such mistakes should be among closely related spe-cies, and lead to minimal phylogenetic distortion, as thegrossest errors are easily identified.Some species were removed after preliminary analyses,
due either to obvious sequencing errors (e.g. highBLAST homology with unrelated families or non-squamates, excessive ambiguities) or a lack of overlap ingenes sampled with other members of the same genus(leading to seemingly artificial paraphyly). We also ex-cluded species with identical sequences between taxaacross all genes, arbitrarily choosing the first taxon in al-phabetical order to remain in the matrix. Additionally,we also removed a few apparent "rogue taxa" [75,77].These were identified by their poor support and suspectplacement (e.g. in a clearly incorrect family), and weretypically represented in the matrix by short fragments ofsingle genes (e.g. an ND4 fragment from the enigmaticcolubroid snake Oreocalamus hanitchsi).The final combined matrix contained sequence data
for: 2335 species for 12S (including 56% of all 4162 taxa,1395 bp), 2377 for 16S (57%, 1970 bp), 730 for BDNF(18%, 714 bp), 1671 for c-mos (40%, 903 bp), 1985 forcyt-b (48%, 1000 bp), 437 for NT3 (10%, 675 bp), 1860for ND2 (45%, 960 bp), 1556 for ND4 (37%, 696 bp), 393
for PDC (9%, 395 bp), 401 for R35 (10%, 768 bp), 1379for RAG-1 (33%, 2700 bp), and 471 for RAG2 (11%, 720bp). The total alignment consists of 12896 bp for 4162taxa (4161 squamates and 1 outgroup). The mean lengthis 2497 bp of sequence data present per species from3.75 genes (19% of the total matrix length of 12896 bp,or 81% missing data), and ranges from 270–11153 bp(2–86% complete). The matrix and phylogeny (seebelow) are available in DataDryad repository 10.5061/dryad.82h0m.Clearly, many taxa had large amounts of missing data
(some >95%), and on average each species had 81%missing cells. However, several lines of evidence suggestthat these missing data are not generally problematic.First, a large body of empirical and theoretical studieshas shown that highly incomplete taxa can be accuratelyplaced in model-based phylogenetic analyses (and withhigh levels of branch support), especially if a large num-ber of characters have been sampled (recent reviews in[84,85]). Second, several recent empirical studies haveshown that the supermatrix approach (with extensivemissing data in some taxa) yields generally well-supported large-scale trees that are generally congruentwith previous taxonomies and phylogenetic estimates(e.g. [41,48,72,73,75,76,197]). Third, recent studies havealso shown that there is generally little relationship be-tween the amount of missing data in individual taxa andthe support for their placement on the tree [41,73,85].Finally, we note that some highly incomplete taxa wereunstable in their placement (“rogue taxa;" [75]), butthese were removed prior to the analysis of the finalmatrix (see above).Our sampling design should be especially robust to
the impacts of missing data for several reasons. Mostimportantly, most terminal taxon (species) had substan-tial data present (mean of 2497 bp per species) regard-less of the number of missing data cells. Simulations (seereviews in [84,85]) suggest that the amount of datapresent is a key parameter in determining the accuracywith which incomplete taxa are placed in phylogenies,not the amount of data absent. Additionally, severalgenes (e.g. 12S/16S, cyt-b, and c-mos) were shared bymany (>40%) of taxa. Thus, there was typically extensiveoverlap among the genes present for each taxon (as alsoindicated by the mean bp per species being much greaterthan the length of most genes). Limited overlap in genesampling among taxa could be highly problematic, irre-spective of the amount of missing data per se, but thisdoes not appear to be a problem in our dataset. Finally,several nuclear genes (e.g. BDNF, c-mos, R35, and RAG-1)were congruently sampled in previous studies to representmost (>80%) squamate families and subfamilies (e.g. [20]),providing a scaffold of well-sampled taxa spanning allmajor clades, as recommended by recent authors [84].

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 45 of 53http://www.biomedcentral.com/1471-2148/13/93
Phylogenetic analysesWe performed phylogenetic analyses of the 12-geneconcatenated matrix using Maximum Likelihood (ML).We assessed node support using the non-parametricShimodaira-Hasegawa-Like (SHL) implementation of theapproximate likelihood-ratio test (aLRT; [94]). This in-volved a two-stage strategy. We first performed initialML tree- inference using the program RAxML-Lightv1.0.7 [209], a modification of the original RAxML algo-rithm [210]. This program uses the GTRCAT strategyfor all genes and partitions, a high-speed approximationof the GTR+Γ model (general time-reversible withgamma-distribution of rate heterogeneity among sites).The GTR model is the only substitution model imple-mented in RAxML [210], and all other substitution modelsare simply special cases of the GTR model [211]. Previousanalyses suggest that GTR is generally the best-fittingmodel for these genes and that they should be partitionedby gene and codon position [16,17,19,20,36,81].To generate an initial ML estimate for final optimization
and support estimation, we performed 11 ML searchesfrom 11 parsimony starting trees generated under the de-fault parsimony model in RAxMLv7.2.8. This number islikely to be sufficient when datasets contain many charac-ters that have strong phylogenetic signal (A. Stamatakis,pers. comm.). Additionally, the dataset was analyzed withthese settings (GTRCAT search from a randomized parsi-mony starting tree) numerous times (>20) as the finalmatrix was assembled and tested, representing hundredsof independent searches from random starting points. Allof the estimated trees from these various analyses showedhigh overall congruence with the final topology. The con-cordance between the preliminary and final results sug-gests that the tree was not strongly impacted by searchesstuck on local optima, and that it should be a good ap-proximation of the ML tree.We then performed a final topology optimization and
assessed support. We passed our best ML estimate ofthe phylogeny (based on GTRCAT) from RAxML-Lightto RAxMLv7.2.8, which does an additional search (usingthe GTRGAMMA model) to produce a nearest-neighborinterchange (NNI)-optimized estimate of the ML tree.This optimization is needed to calculate the SHL versionof the aLRT for estimating support values [94]. The SHL-aLRT strategy approximates a test of the null hypothesisthat the branch length subtending each node equals 0(i.e. that the node can be resolved, rather than estimatedas a polytomy) with a test of the more general null hy-pothesis that "the branch is incorrect" relative to the fournext suboptimal arrangements of that node relative to theNNI-optimal arrangement [94]. Based on initial analyses,generating sufficient ML bootstrap replicates for a tree ofthis size proved computationally intractable, so we rely onSHL values alone to assess support.
The SHL approach has at least two major advantagesover non-parametric bootstrapping for large ML trees:(i) values are apparently robust to many potential modelviolations and have the same properties as bootstrapproportions for all but the shortest branches [41,94,212],and (ii) values for short branches may be more accuratethan bootstrap proportions, as support is evaluatedbased on whole-alignment likelihoods, rather than thefrequency of re-sampled characters [94,213]. Additio-nally, the SHL approach is orders of magnitude fasterthan traditional bootstrapping [94,212,213], and it ap-pears to be similarly robust to matrices with extensivemissing data [41]. As in previous studies, we take a con-servative view, considering SHL values of 85 or greater(i.e. a 15% chance that a branch is "incorrect") as strongsupport [41,212,213].These analyses were performed on a 360-core SGI ICE
supercomputing system ("ANDY") at the High Perform-ance Computing Center at the City University of NewYork (CUNY). The final analysis was completed in 8.8days of computer time using 188 nodes of the CUNYsupercomputing cluster.Finally, we assessed the potential impact of missing data
on our branch-length estimates. We performed linear re-gression (in R) of the proportional completeness of eachterminal taxa (non-missing data in bp / maximum amountof non-missing data, 12896 bp) against the length of itsterminal branch. This test addresses whether incompletetaxa have branch-length estimates that are consistentlybiased in one direction (shorter vs. longer) relative tomore complete terminals. However, it does not directlytest whether branch length estimates are correct or not,nor how branch length estimates are impacted by re-placing non-missing data with missing data (see [87] forresults suggesting that such replacements have little effectin real data sets).
Appendix INote that we only provide here an account for the onesubfamily newly erected in this study. We do not provideaccounts for subfamilies with changes in content (Boinae,Erycinae, Dipsadinae, Pseudaspidinae, Scincinae, Unga-liophiinae), that have been resurrrected (Sanziniinae), orthat represent elevation of lower-ranked taxa (tribe Bachi-ini here recognized as Bachiinae).
Candoiinae subfam. nov. (family Boidae)Type: genus and species Candoia carinata [214].Content: one genus, 4 species; C. aspera, C. bibroni, C.
carinata, C. paulsoni.Definition: this subfamily consists of the most recent
common ancestor of the extant species of Candoia, andall its descendants. These species are morphologically dis-tinguished in part from other boid snakes by a flattened

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 46 of 53http://www.biomedcentral.com/1471-2148/13/93
rostrum leading to an angular snout [215] and a wide pre-maxillary floor [167].Distribution: these snakes are primarily restricted to
the South Pacific islands of New Guinea and Melanesia,and the eastern Indonesian archipelago [150].Remarks: the three species from this subfamily that are
sampled in our tree are strongly supported as monophyletic(SHL = 100), and are well supported (SHL = 87) as the sis-ter taxon to a moderately supported clade consisting ofErycinae + Boinae (SHL = 83).
Proposed Generic Composition of Higher TaxaBelow, we list the familial and subfamilial assignment ofall squamate genera from the December, 2012 update ofthe Reptile Database [1], updated to reflect some recentchanges and the proposed subfamily level changes listedabove. As this classification includes numerous taxa notsampled in our tree, we deal with them conservatively.For traditionally recognized families and subfamilies thatwe found to be monophyletic, we include all taxa trad-itionally assigned to them. Taxa are denoted incertaesedis if they are of ambiguous familial or subfamilial as-signment due to uncertain placement in our tree, or dueto absence from our tree and lack of assignment by pre-vious authors. This classification includes 67 families and56 subfamilies, and accounts for >9400 squamate species in1018 genera [1]. Higher taxa are listed (more-or-less)phylogenetically (starting closest to the root; Figure 1), fa-milies are listed alphabetically within higher taxa, and sub-families and genera are listed alphabetically within families.
SquamataDibamidae (Anelytropsis, Dibamus)
GekkotaCarphodactylidae (Carphodactylus, Nephrurus, Orraya,Phyllurus, Saltuarius, Underwoodisaurus, Uvidicolus);Diplodactylidae (Amalosia, Bavayia, Correlophus,Crenadactylus, Dactylocnemis, Dierogekko, Diplodactylus,Eurydactylodes, Hesperoedura, Hoplodactylus, Lucasium,Mniarogekko, Mokopirirakau, Naultinus, Nebulifera,Oedodera, Oedura, Paniegekko, Pseudothecadactylus,Rhacodactylus, Rhynchoedura, Strophurus, Toropuku,Tukutuku, Woodworthia); Eublepharidae (Aelurosca-labotes, Coleonyx, Eublepharis, Goniurosaurus, Hemithe-conyx, Holodactylus); Gekkonidae (Afroedura, Afrogecko,Agamura, Ailuronyx, Alsophylax, Asiocolotes, Blaeso-dactylus, Bunopus, Calodactylodes, Chondrodactylus,Christinus, Cnemaspis, Colopus, Crossobamon, Cryptactites,Cyrtodactylus, Cyrtopodion, Dixonius, Ebenavia, Elasmo-dactylus, Geckolepis, Gehyra, Gekko, Goggia, Hemidactylus,Hemiphyllodactylus, Heteronotia, Homopholis, Lepidoda-ctylus, Luperosaurus, Lygodactylus, Matoatoa, Mediodac-tylus, Nactus, Narudasia, Pachydactylus, Paragehyra,
Paroedura, Perochirus, Phelsuma, Pseudoceramodactylus,Pseudogekko, Ptenopus, Ptychozoon, Rhinogecko, Rhoptro-pella, Rhoptropus, Stenodactylus, Tropiocolotes, Urocotyledon,Uroplatus); Phyllodactylidae (Asaccus, Gymnodactylus,Haemodracon, Homonota, Phyllodactylus, Phyllopezus,Ptyodactylus, Tarentola, Thecadactylus); Pygopodidae(Aprasia, Delma, Lialis, Ophidiocephalus, Paradelma, Ple-tholax, Pygopus); Sphaerodactylidae (Aristelliger, Chato-gekko, Coleodactylus, Euleptes, Gonatodes, Lepidoblepharis,Pristurus, Pseudogonatodes, Quedenfeldtia, Saurodactylus,Sphaerodactylus, Teratoscincus)
ScincoideaCordylidae, Cordylinae (Chamaesaura, Cordylus, Hemi-cordylus, Karusasaurus, Namazonurus, Ninurta, Ouro-borus, Pseudocordylus, Smaug), Platysaurinae (Platysaurus);Gerrhosauridae, Gerrhosaurinae (Cordylosaurus, Gerrho-saurus, Tetradactylus), Zonosaurinae (Tracheloptychus,Zonosaurus); Scincidae, Acontiinae (Acontias, Typhlosau-rus), Lygosominae (Ablepharus, Afroablepharus, Anomalo-pus, Asymblepharus, Ateuchosaurus, Bartleia, Bassiana,Bellatorias, Caledoniscincus, Calyptotis, Carlia, Cautula,Celatiscincus, Chioninia, Coeranoscincus, Coggeria, Cophos-cincopus, Corucia, Cryptoblepharus, Ctenotus, Cyclodomor-phus, Dasia, Egernia, Emoia, Eremiascincus, Eroticoscincus,Eugongylus, Eulamprus, Eumecia, Eutropis, Fojia, Geo-myersia, Geoscincus, Glaphyromorphus, Gnypetoscincus,Graciliscincus, Haackgreerius, Hemiergis, Hemisphaeriodon,Insulasaurus, Isopachys, Kaestlea, Kanakysaurus, Lacertaspis,Lacertoides, Lamprolepis, Lampropholis, Lankascincus,Larutia, Leiolopisma, Lepidothyris, Leptoseps, Leptosiaphos,Lerista, Liburnascincus, Liopholis, Lioscincus, Lipinia,Lissolepis, Lobulia, Lygisaurus, Lygosoma, Mabuya, Marmo-rosphax, Menetia, Mochlus, Morethia, Nangura, Nanno-scincus, Niveoscincus, Notoscincus, Oligosoma, Ophioscincus,Otosaurus, Panaspis, Papuascincus, Parvoscincus, Phobosci-ncus, Pinoyscincus, Prasinohaema, Proablepharus, Pseude-moia, Ristella, Saiphos, Saproscincus, Scincella, Sigaloseps,Simiscincus, Sphenomorphus, Tachygyia, Tiliqua, Trachy-lepis, Tribolonotus, Tropidophorus, Tropidoscincus, Tytthos-cincus, Vietnascincus), Scincinae (Amphiglossus, Androngo,Barkudia, Brachymeles, Chabanaudia, Chalcides, Chalci-doseps, Eumeces, Eurylepis, Feylinia, Gongylomorphus,Hakaria, Janetaescincus, Jarujinia, Madascincus, Melano-seps, Mesoscincus, Nessia, Ophiomorus, Pamelaescincus,Paracontias, Plestiodon, Proscelotes, Pseudoacontias, Pygo-meles, Scelotes, Scincopus, Scincus, Scolecoseps, Sepsina,Sepsophis, Sirenoscincus, Typhlacontias, Voeltzkowia);Xantusiidae, Cricosaurinae (Cricosaura), Lepidophyminae(Lepidophyma), Xantusiinae (Xantusia)
Lacertoidea (including Amphisbaenia)Amphisbaenidae (Amphisbaena, Ancylocranium, Baikia,Chirindia, Cynisca, Dalophia, Geocalamus, Loveridgea,

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 47 of 53http://www.biomedcentral.com/1471-2148/13/93
Mesobaena, Monopeltis, Zygaspis); Bipedidae (Bipes);Blanidae (Blanus); Cadeidae (Cadea); Gymnoph-thalmidae, Alopoglossinae (Alopoglossus, Ptychoglossus),Bachiinae (Bachia), Cercosaurinae (Anadia, Cercosaura,Echinosaura, Euspondylus, Macropholidus, Neusticurus,Opipeuter, Petracola, Pholidobolus, Placosoma, Pota-mites, Proctoporus, Riama, Riolama, Teuchocercus), Ec-pleopinae (Adercosaurus, Amapasaurus, Anotosaura,Arthrosaura, Colobosauroides, Dryadosaura, Ecpleopus,Kaieteurosaurus, Leposoma, Marinussaurus, Pantepuisaurus),Gymnophthalminae (Acratosaura, Alexandresaurus, Calyp-tommatus, Caparaonia, Colobodactylus, Colobosaura,Gymnophthalmus, Heterodactylus, Iphisa, Micrablepharus,Nothobachia, Procellosaurinus, Psilophthalmus, Scripto-saura, Stenolepis, Tretioscincus, Vanzosaura), Rhachisau-rinae (Rhachisaurus); Lacertidae, Gallotiinae (Gallotia,Psammodromus), Lacertinae (Acanthodactylus, Adolfus,Algyroides, Anatololacerta, Apathya, Archaeolacerta,Atlantolacerta, Australolacerta, Congolacerta, Dalmato-lacerta, Darevskia, Dinarolacerta, Eremias, Gastropholis,Heliobolus, Hellenolacerta, Holaspis, Iberolacerta, Ichno-tropis, Iranolacerta, Lacerta, Latastia, Meroles, Mesalina,Nucras, Omanosaura, Ophisops, Parvilacerta, Pedioplanis,Philochortus, Phoenicolacerta, Podarcis, Poromera, Pseude-remias, Scelarcis, Takydromus, Teira, Timon, Tropidosaura,Zootoca); Rhineuridae (Rhineura); Teiidae, Teiinae(Ameiva, Aspidoscelis, Cnemidophorus, Dicrodon, Ken-tropyx, Teius), Tupinambinae (Callopistes, Crocodilurus,Dracaena, Tupinambis); Trogonophiidae (Agamodon,Diplometopon, Pachycalamus,Trogonophis)
IguaniaAgamidae, Agaminae (Acanthocercus, Agama, Brachysaura,Bufoniceps, Laudakia, Phrynocephalus, Pseudotrapelus,Trapelus, Xenagama), Amphibolurinae (Amphibolurus,Chelosania, Chlamydosaurus, Cryptagama, Ctenophorus,Diporiphora, Hypsilurus, Intellagama, Lophognathus,Moloch, Physignathus, Pogona, Rankinia, Tympanocryptis),Draconinae (Acanthosaura, Aphaniotis, Bronchocela, Ca-lotes, Ceratophora, Complicitus, Cophotis, Coryphophylax,Dendragama, Draco, Gonocephalus, Harpesaurus, Hypsica-lotes, Japalura, Lophocalotes, Lyriocephalus, Mantheyus,Oriocalotes, Otocryptis, Phoxophrys, Psammophilus,Pseudocalotes, Pseudocophotis, Ptyctolaemus, Salea, Sitana,Thaumatorhynchus), Hydrosaurinae (Hydrosaurus), Leiole-pidinae (Leiolepis), Uromastycinae (Uromastyx); Cha-maeleonidae, Brookesiinae (Brookesia), Chamaeleoninae(Archaius, Bradypodion, Calumma, Chamaeleo, Furcifer,Kinyongia, Nadzikambia, Rhampholeon, Rieppeleon, Trioceros);Corytophanidae (Basiliscus, Corytophanes, Laemanctus);Crotaphytidae (Crotaphytus, Gambelia); Dactyloidae (Ano-lis); Hoplocercidae (Enyalioides, Hoplocercus, Morunasaurus);Iguanidae (Amblyrhynchus, Brachylophus, Conolophus,Ctenosaura, Cyclura, Dipsosaurus, Iguana, Sauromalus);
Leiocephalidae (Leiocephalus); Leiosauridae, Enyaliinae(Anisolepis, Enyalius, Urostrophus), Leiosaurinae (Diplolaemus,Leiosaurus, Pristidactylus); Liolaemidae (Ctenoblepharys,Liolaemus, Phymaturus); Opluridae (Chalarodon, Oplurus);Phrynosomatidae (Callisaurus, Cophosaurus, Holbrookia,Petrosaurus, Phrynosoma, Sceloporus, Uma, Urosaurus, Uta);Polychrotidae (Polychrus); Tropiduridae (Eurolophosaurus,Microlophus, Plica, Stenocercus, Strobilurus, Tropidurus,Uracentron,Uranoscodon)
AnguimorphaAnguidae, Anguinae (Anguis, Dopasia, Ophisaurus,Pseudopus), Diploglossinae (Celestus, Diploglossus,Ophiodes), Gerrhonotinae (Abronia, Barisia, Colo-ptychon, Elgaria, Gerrhonotus, Mesaspis); Anniellidae(Anniella); Helodermatidae (Heloderma); Lanthano-tidae (Lanthanotus); Shinisauridae (Shinisaurus);Varanidae (Varanus); Xenosauridae (Xenosaurus)
SerpentesAcrochordidae (Acrochordus); Aniliidae (Anilius);Anomalepididae (Anomalepis, Helminthophis, Liotyphlops,Typhlophis); Anomochilidae (Anomochilus); Boidae,Boinae (Boa, Corallus, Epicrates, Eunectes), Candoiinae(Candoia), Erycinae (Eryx), Sanziniinae (Acrantophis,Sanzinia), Ungaliophiinae (Charina, Exiliboa, Lichanura,Ungaliophis); Bolyeriidae (Bolyeria, Casarea); Calaba-riidae (Calabaria); Colubridae incertae sedis (Blythia,Cyclocorus, Elapoidis, Gongylosoma, Helophis, Myersophis,Oreocalamus, Poecilopholis, Rhabdops, Tetralepis),Calamariinae (Calamaria, Calamorhabdium, Collor-habdium, Etheridgeum, Macrocalamus, Pseudorabdion,Rabdion), Colubrinae (Aeluroglena, Ahaetulla, Apros-doketophis, Archelaphe, Argyrogena, Arizona, Bamanophis,Bogertophis, Boiga, Cemophora, Chilomeniscus, Chionactis,Chironius, Chrysopelea, Coelognathus, Coluber, Colubroe-laps, Conopsis, Coronella, Crotaphopeltis, Cyclophiops,Dasypeltis, Dendrelaphis, Dendrophidion, Dipsadoboa,Dispholidus, Dolichophis, Drymarchon, Drymobius, Drymo-luber, Dryocalamus, Dryophiops, Eirenis, Elachistodon,Elaphe, Euprepiophis, Ficimia, Geagras, Gonyophis, Gonyo-soma, Gyalopion, Hapsidophrys, Hemerophis, Hemorrhois,Hierophis, Lampropeltis, Leptodrymus, Leptophis, Leptu-rophis, Limnophis, Liopeltis, Lycodon, Lytorhynchus, Macro-protodon, Mastigodryas, Meizodon, Oligodon, Oocatochus,Opheodrys, Oreocryptophis, Orthriophis, Oxybelis, Panthe-rophis, Philothamnus, Phyllorhynchus, Pituophis, Platyceps,Pseudelaphe, Pseudoficimia, Pseustes, Ptyas, Rhadinophis,Rhamnophis, Rhinechis, Rhinobothryum, Rhinocheilus, Rhy-nchocalamus, Rhynchophis, Salvadora, Scaphiophis, Scole-cophis, Senticolis, Simophis, Sonora, Spalerosophis, Spilotes,Stegonotus, Stenorrhina, Symphimus, Sympholis, Tantilla,Tantillita, Telescopus, Thelotornis, Thrasops, Toxicodryas,Trimorphodon, Xenelaphis, Xyelodontophis, Zamenis),

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 48 of 53http://www.biomedcentral.com/1471-2148/13/93
Dipsadinae (Adelphicos, Alsophis, Amastridium, Amnes-teophis, Antillophis, Apostolepis, Arrhyton, Atractus, Boiru-na, Borikenophis, Caaeteboia, Calamodontophis, Caraiba,Carphophis, Cercophis, Chapinophis, Chersodromus, Clelia,Coniophanes, Conophis, Contia, Coronelaps, Crisantophis,Cryophis, Cubophis, Darlingtonia, Diadophis, Diaphoro-lepis, Dipsas, Ditaxodon, Drepanoides, Echinanthera,Elapomorphus, Emmochliophis, Enuliophis, Enulius, Eryth-rolamprus, Farancia, Geophis, Gomesophis, Haitiophis,Helicops, Heterodon, Hydrodynastes, Hydromorphus, Hy-drops, Hypsiglena, Hypsirhynchus, Ialtris, Imantodes, Lepto-deira, Lioheterophis, Lygophis, Magliophis, Manolepis,Mussurana, Ninia, Nothopsis, Ocyophis, Omoadiphas, Oxy-rhopus, Paraphimophis, Phalotris, Philodryas, Phimophis,Plesiodipsas, Pliocercus, Pseudalsophis, Pseudoboa, Pseu-doeryx, Pseudoleptodeira, Pseudotomodon, Psomophis, Pty-chophis, Rhachidelus, Rhadinaea, Rhadinella, Rhadinophanes,Rodriguesophis, Saphenophis, Schwartzophis, Sibon, Sibyno-morphus, Siphlophis, Sordellina, Synophis, Tachymenis,Taeniophallus, Tantalophis, Thamnodynastes, Thermophis,Tomodon, Tretanorhinus, Trimetopon, Tropidodipsas, Tropi-dodryas, Uromacer, Uromacerina, Urotheca, Xenodon,Xenopholis), Grayiinae (Grayia), Natricinae (Adelophis,Afronatrix, Amphiesma, Amphiesmoides, Anoplohydrus,Aspidura, Atretium, Balanophis, Clonophis, Hologerrhum,Hydrablabes, Hydraethiops, Iguanognathus, Lycognathophis,Macropisthodon, Natriciteres, Natrix, Nerodia, Opistho-tropis, Parahelicops, Pararhabdophis, Paratapinophis,Regina, Rhabdophis, Seminatrix, Sinonatrix, Storeria,Thamnophis, Trachischium, Tropidoclonion, Tropidonophis,Virginia, Xenochrophis), Pseudoxenodontinae (Plagiopholis,Pseudoxenodon), Sibynophiinae (Scaphiodontophis, Siby-nophis); Cylindrophiidae (Cylindrophis); Elapidae (Acan-thophis, Aipysurus, Aspidelaps, Aspidomorphus, Austrelaps,Bungarus, Cacophis, Calliophis, Cryptophis, Demansia,Dendroaspis, Denisonia, Drysdalia, Echiopsis, Elapo-gnathus, Elapsoidea, Emydocephalus, Ephalophis, Furina,Hemachatus, Hemiaspis, Hemibungarus, Hoplocephalus,Hydrelaps, Hydrophis, Kolpophis, Laticauda, Loveridgelaps,Maticora, Micropechis, Micruroides, Micrurus, Naja,Notechis, Ogmodon, Ophiophagus, Oxyuranus, Parahy-drophis, Parapistocalamus, Parasuta, Pseudechis, Pseu-dohaje, Pseudolaticauda, Pseudonaja, Rhinoplocephalus,Salomonelaps, Simoselaps, Sinomicrurus, Suta, Thalas-sophis, Toxicocalamus, Tropidechis, Vermicella, Walterin-nesia); Gerrhopilidae (Gerrhopilus); Homalopsidae (Bitia,Brachyorrhos, Cantoria, Cerberus, Djokoiskandarus,Enhydris, Erpeton, Fordonia, Gerarda, Heurnia, Homa-lopsis, Myron, Pseudoferania); Lamprophiidae incertaesedis (Micrelaps, Montaspis, Oxyrhabdium), Aparallactinae(Amblyodipsas, Aparallactus, Brachyophis, Chilorhinophis,Elapotinus, Hypoptophis, Macrelaps, Polemon, Xenocala-mus), Atractaspidinae (Atractaspis, Homoroselaps), Lam-prophiinae (Boaedon, Bothrophthalmus, Chamaelycus,
Dendrolycus, Gonionotophis, Hormonotus, Inyoka, Lam-prophis, Lycodonomorphus, Lycophidion, Pseudoboodon),Prosymninae (Prosymna), Psammophiinae (Dipsina, Hemi-rhagerrhis, Malpolon, Mimophis, Psammophis, Psam-mophylax, Rhagerhis, Rhamphiophis), Pseudaspidinae(Buhoma, Psammodynastes, Pseudaspis, Pythonodipsas),Pseudoxyrhophiine (Alluaudina, Amplorhinus, Bothrolycus,Brygophis, Compsophis, Ditypophis, Dromicodryas, Duber-ria, Exallodontophis, Heteroliodon, Ithycyphus, Langaha,Leioheterodon, Liophidium, Liopholidophis, Lycodryas,Madagascarophis, Micropisthodon, Pararhadinaea, Paras-tenophis, Phisalixella, Pseudoxyrhopus, Thamnosophis);Leptotyphlopidae (Epacrophis, Epictia, Leptotyphlops,Mitophis, Myriopholis, Namibiana, Rena, Rhinoleptus,Siagonodon, Tetracheilostoma, Tricheilostoma, Trilepida);Loxocemidae (Loxocemus); Pareatidae (Aplopeltura,Asthenodipsas, Pareas); Pythonidae (Antaresia, Apodora,Aspidites, Bothrochilus, Broghammerus, Leiopython, Liasis,Morelia, Python); Tropidophiidae (Trachyboa, Tropido-phis); Typhlopidae (Acutotyphlops, Afrotyphlops, Aus-trotyphlops, Cyclotyphlops, Grypotyphlops, Letheobia,Megatyphlops, Ramphotyphlops, Rhinotyphlops, Typhlops);Uropeltidae (Brachyophidium, Melanophidium, Platyp-lectrurus, Plectrurus, Pseudotyphlops, Rhinophis,Teretrurus,Uropeltis); Viperidae, Azemiopinae (Azemiops), Crotalinae(Agkistrodon, Atropoides, Bothriechis, Bothriopsis, Bothro-cophias, Bothropoides, Bothrops, Calloselasma, Cerrophi-dion, Crotalus, Deinagkistrodon, Garthius, Gloydius,Hypnale, Lachesis, Mixcoatlus, Ophryacus, Ovophis, Porthi-dium, Protobothrops, Rhinocerophis, Sistrurus, Trimere-surus, Tropidolaemus), Viperinae (Atheris, Bitis, Causus,Cerastes, Daboia, Echis, Eristicophis, Macrovipera, Monta-theris, Montivipera, Proatheris, Pseudocerastes, Vipera);Xenodermatidae (Achalinus, Fimbrios, Stoliczkia, Xeno-dermus, Xylophis); Xenopeltidae (Xenopeltis); Xeno-phidiidae (Xenophidion); Xenotyphlopidae (Xenotyphlops)
Additional files
Additional file 1: Data File S1. The 4162-species ML phylogeny inNewick format; taxonomic changes are given in Additional file 2: Table S1.
Additional file 2: Table S1. GenBank accession numbers for all taxaincluded in this analysis.
Competing interestsThe authors declare that they have no competing interests.
Authors' contributionsRAP and JJW conceived the study. RAP, FTB, and JJW conducted analysesand checked results. RAP, FTB, and JJW wrote the MS. All authors read andapproved the final manuscript.
AcknowledgementsWe thank the many researchers who made this study possible through theirdetailed studies of squamate phylogeny with a (mostly) shared set ofmolecular markers, and uploading their sequence data to GenBank. Wethank E. Paradis, J. Lombardo T. Guiher, R. Walsh, and P. Muzio for

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 49 of 53http://www.biomedcentral.com/1471-2148/13/93
computational assistance, T. Gamble, D. Frost, D. Cannatella, H. Zaher, F.Grazziotin, P. Uetz, T. Jackman, and A. Bauer for taxonomic advice, and A.Larson, M. Vences, and five anonymous reviewers for comments on thismanuscript. The research was supported in part by the U.S. National ScienceFoundation, including a Bioinformatics Postdoctoral grant to R.A.P.(DBI-0905765), an AToL grant to J.J.W. (EF-0334923), and grants to theCUNY HPCC (CNS-0958379 and CNS-0855217).
Author details1Department of Biological Sciences, The George Washington University,2023 G St. NW, Washington, DC 20052, USA. 2Department of Biology, TheGraduate School and University Center, The City University of New York, 3655th Ave., New York, NY 10016, USA. 3Department of Biology, The College ofStaten Island, The City University of New York, 2800 Victory Blvd., StatenIsland, NY 10314, USA. 4Department of Ecology and Evolutionary Biology,University of Arizona, Tucson, AZ 85721-0088, USA.
Received: 30 January 2013 Accepted: 19 March 2013Published: 29 April 2013
References1. Uetz P: The Reptile Database. http://www.reptile-database.org/ Accessed
December, 2012.2. Greene HW: Snakes: the Evolution of Mystery in Nature. Berkeley: University of
California Press; 1997.3. Vitt LJ, Caldwell JP: Herpetology. 4th edition. Burlington: Elsevier; 2009.4. Pianka ER, Vitt LJ: Lizards: Windows to the Evolution of Diversity. Berkeley:
University of California Press; 2003.5. Kasturiratne A, Wickremasinghe AR, de Silva N, Gunawardena NK,
Pathmeswaran A, Premaratna R, Savioli L, Lalloo DG, de Silva HJ: Theglobal burden of snakebite: a literature analysis and modelling basedon regional estimates of envenoming and deaths. PLoS Med 2008,5:1591–1604.
6. Mahdavi A, Ferreira L, Sundback C, Nichol JW, Chan EP, Carter DJD,Bettinger CJ, Patanavanich S, Chignozha L, Ben-Joseph E, et al: Abiodegradable and biocompatible gecko-inspired tissue adhesive. ProcNatl Acad Sci USA 2008, 105:2307–2312.
7. Geim AK, Dubonos SV, Grigorieva IV, Novoselov KS, Zhukov AA, Shapoval SY:Microfabricated adhesive mimicking gecko foot-hair. Nat Mater 2003,2:461–463.
8. Huey RB, Bennett AF: Phylogenetic studies of coadaptation: preferredtemperatures versus optimal performance temperatures of lizards.Evolution 1987, 41:1098–1115.
9. Losos JB: The evolution of form and function: morphology andlocomotor performance in West-Indian Anolis lizards. Evolution 1990,44:1189–1203.
10. Wiens JJ, Brandley MC, Reeder TW: Why does a trait evolve multiple timeswithin a clade? Repeated evolution of snakelike body form in squamatereptiles. Evolution 2006, 60:123–141.
11. Bergmann PJ, Irschick DJ: Vertebral evolution and the diversification ofsquamate reptiles. Evolution 2012, 66:1044–1058.
12. Losos JB, Hillis DM, Greene HW: Who speaks with a forked tongue? Science2012, 338:1428–1429.
13. Estes R, de Queiroz K, Gauthier J: Phylogenetic relationships withinSquamata. In Phylogenetic Relationships of the Lizard Families. Edited byEstes R, Pregill G. Stanford: Stanford University Press; 1988:119–281.
14. Gauthier JA, Kearney M, Maisano JA, Rieppel O, Behike ADB: Assemblingthe Squamate Tree of Life: perspectives from the phenotype and thefossil record. Bull Peabody Mus Nat Hist 2012, 53:3–308.
15. Conrad JL: Phylogeny and systematics of Squamata (Reptilia) based onmorphology. Bull Am Mus Nat Hist 2008, 310:1–182.
16. Mulcahy DG, Noonan BP, Moss T, Townsend TM, Reeder TW, Sites JW Jr,Wiens JJ: Estimating divergence times and evaluating dating methodsusing phylogenomic and mitochondrial data in squamate reptiles. MolPhylogenet Evol 2012, 65:974–991.
17. Townsend TM, Larson A, Louis E, Macey JR: Molecular phylogenetics ofSquamata: the position of snakes, amphisbaenians, and dibamids, andthe root of the squamate tree. Syst Biol 2004, 53:735–757.
18. Vidal N, Hedges SB: The phylogeny of squamate reptiles (lizards, snakes,and amphisbaenians) inferred from nine nuclear protein-coding genes.CR Biol 2005, 328:1000–1008.
19. Wiens JJ, Kuczynski CA, Townsend T, Reeder TW, Mulcahy DG, Sites JW:Combining phylogenomics and fossils in higher-level squamate reptilephylogeny: molecular data change the placement of fossil taxa. Syst Biol2010, 59:674–688.
20. Wiens JJ, Hutter CR, Mulcahy DG, Noonan BP, Townsend TM, Sites JW,Reeder TW: Resolving the phylogeny of lizards and snakes(Squamata) with extensive sampling of genes and species. Biol Lett2012, 8:1043–1046.
21. Camp CL: Classification of the lizards. Bull Am Mus Nat Hist 1923,48:289–480.
22. Lee MSY: Convergent evolution and character correlation in burrowingreptiles: towards a resolution of squamate relationships. Biol J Linn Soc1998, 65:369–453.
23. Vidal N, Hedges SB: Molecular evidence for a terrestrial origin of snakes.Proc R Soc B 2004, 271:S226–S229.
24. Fry BG, Vidal N, Norman JA, Vonk FJ, Scheib H, Ramjan SFR, Kuruppu S,Fung K, Hedges SB, Richardson MK, et al: Early evolution of the venomsystem in lizards and snakes. Nature 2006, 439:584–588.
25. Zwickl DJ, Hillis DM: Increased taxon sampling greatly reducesphylogenetic error. Syst Biol 2002, 51:588–598.
26. Poe S: Evaluation of the strategy of long-branch subdivision to improvethe accuracy of phylogenetic methods. Syst Biol 2003, 52:423–428.
27. Heath TA, Zwickl DJ, Kim J, Hillis DM: Taxon sampling affects inferences ofmacroevolutionary processes from phylogenetic trees. Syst Biol 2008,57:160–166.
28. Graybeal A: Is it better to add taxa or characters to a difficultphylogenetic problem? Syst Biol 1998, 47:9–17.
29. Blankers T, Townsend TM, Pepe K, Reeder TW, Wiens JJ: Contrastingglobal-scale evolutionary radiations: phylogeny, diversification, andmorphological evolution in the major clades of iguanian lizards. Biol JLinn Soc 2013, 108:127–143.
30. Schulte JA, Moreno-Roark F: Live birth among iguanian lizards predatesPliocene-Pleistocene glaciations. Biol Lett 2010, 6:216–218.
31. Frost DR, Etheridge RE, Janies D, Titus TA: Total evidence, sequencealignment, evolution of polychrotid lizards, and a reclassification of theIguania (Squamata: Iguania). Am Mus Novit 2001, 3343:38.
32. Macey JR, Schulte JA, Larson A, Ananjeva NB, Wang YZ, Pethiyagoda R,Rastegar-Pouyani N, Papenfuss TJ: Evaluating trans-Tethys migration: anexample using acrodont lizard phylogenetics. Syst Biol 2000, 49:233–256.
33. Schulte JA, Valladares JP, Larson A: Phylogenetic relationships withinIguanidae inferred using molecular and morphological data and aphylogenetic taxonomy of iguanian lizards. Herpetologica 2003,59:399–419.
34. Townsend TM, Mulcahy DG, Noonan BP, Sites JW, Kuczynski CA, Wiens JJ,Reeder TW: Phylogeny of iguanian lizards inferred from 29 nuclear loci,and a comparison of concatenated and species-tree approaches for anancient, rapid radiation. Mol Phylogenet Evol 2011, 61:363–380.
35. Slowinski JB, Lawson R: Snake phylogeny: evidence from nuclear andmitochondrial genes. Mol Phylogenet Evol 2002, 24:194–202.
36. Wiens JJ, Kuczynski CA, Smith SA, Mulcahy DG, Sites JW, Townsend TM,Reeder TW: Branch lengths, support, and congruence: testing thephylogenomic approach with 20 nuclear loci in snakes. Syst Biol 2008,57:420–431.
37. Lawson R, Slowinski JB, Burbrink FT: A molecular approach to discerningthe phylogenetic placement of the enigmatic snake Xenophidionschaeferi among the Alethinophidia. J Zool 2004, 263:285–294.
38. Vidal N, Marin J, Morini M, Donnellan S, Branch WR, Thomas R, Vences M,Wynn A, Cruaud C, Hedges SB: Blindsnake evolutionary tree reveals longhistory on Gondwana. Biol Lett 2010, 6:558–561.
39. Adalsteinsson SA, Branch WR, Trape S, Vitt LJ, Hedges SB: Molecularphylogeny, classification, and biogeography of snakes of the familyLeptotyphlopidae (Reptilia, Squamata). Zootaxa 2009, 2244:1–50.
40. Kelly CMR, Barker NP, Villet MH, Broadley DG: Phylogeny, biogeographyand classification of the snake superfamily Elapoidea: a rapid radiationin the late Eocene. Cladistics 2009, 25:38–63.
41. Pyron RA, Burbrink FT, Colli GR, de Oca ANM, Vitt LJ, Kuczynski CA, Wiens JJ:The phylogeny of advanced snakes (Colubroidea), with discovery of anew subfamily and comparison of support methods for likelihood trees.Mol Phylogenet Evol 2011, 58:329–342.
42. Zaher H, Grazziotin FG, Cadle JE, Murphy RW, de Moura JC, Bonatto SL:Molecular phylogeny of advanced snakes (Serpentes, Caenophidia) with

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 50 of 53http://www.biomedcentral.com/1471-2148/13/93
an emphasis on South American xenodontines: a revised classificationand descriptions of new taxa. Pap Av Zool 2009, 49:115–153.
43. Grazziotin FG, Zaher H, Murphy RW, Scrocchi G, Benavides MA, ZhangY-P, Bonatto SL: Molecular phylogeny of the New World Dipsadidae(Serpentes: Colubroidea): a reappraisal. Cladistics 2012,28:437–459.
44. Pyron RA, Kandambi HKDK, Hendry CR, Pushpamal V, Burbrink FT,Somaweera R: Genus-level phylogeny of snakes reveals the origins ofspecies richness in Sri Lanka. Mol Phylogenet Evol 2013,66:969–978.
45. Vidal N, Delmas AS, David P, Cruaud C, Coujoux A, Hedges SB: Thephylogeny and classification of caenophidian snakes inferred from sevennuclear protein-coding genes. CR Biol 2007, 330:182–187.
46. Sanders KL, Lee MSY, Bertozzi T, Rasmussen AR: Multilocus phylogeny andrecent rapid radiation of the viviparous sea snakes (Elapidae:Hydrophiinae). Mol Phylogenet Evol 2012, 66:575–591.
47. Noonan BP, Chippindale PT: Dispersal and vicariance: the complexevolutionary history of boid snakes. Mol Phylogenet Evol 2006, 40:347–358.
48. Lynch VJ, Wagner GP: Did egg-laying boas break Dollo's law?Phylogenetic evidence for reversal to oviparity in sand boas (Eryx:Boidae). Evolution 2010, 64:207–216.
49. Schmitz A, Brandley MC, Mausfeld P, Vences M, Glaw F, Nussbaum RA,Reeder TW: Opening the black box: phylogenetics and morphologicalevolution of the Malagasy fossorial lizards of the subfamily "Scincinae".Mol Phylogenet Evol 2005, 34:118–133.
50. Brandley MC, Schmitz A, Reeder TW: Partitioned Bayesian analyses,partition choice, and the phylogenetic relationships of scincid lizards.Syst Biol 2005, 54:373–390.
51. Whiting AS, Bauer AM, Sites JW: Phylogenetic relationships and limb lossin sub-Saharan African scincine lizards (Squamata: Scincidae). MolPhylogenet Evol 2003, 29:582–598.
52. Skinner A, Lee MSY, Hutchinson MN: Rapid and repeated limb loss in aclade of scincid lizards. BMC Evol Biol 2008, 8:310.
53. Gamble T, Colli GR, Rodrigues MT, Werneck FP, Simons AM: Phylogeny andcryptic diversity in geckos (Phyllopezus; Phyllodactylidae; Gekkota) fromSouth America's open biomes. Mol Phylogenet Evol 2012, 62:943–953.
54. Gamble T, Daza JD, Colli GR, Vitt LJ, Bauer AM: A new genus ofminiaturized and pug-nosed gecko from South America(Sphaerodactylidae: Gekkota). Zool J Linn Soc 2011, 163:1244–1266.
55. Gamble T, Bauer AM, Colli GR, Greenbaum E, Jackman TR, Vitt LJ, SimonsAM: Coming to America: multiple origins of New World geckos. J EvolBiol 2011, 24:231–244.
56. Gamble T, Bauer AM, Greenbaum W, Jackman TR: Out of the blue: a novel,trans-Atlantic clade of geckos (Gekkota, Squamata). Zool Scripta 2008,37:355–366.
57. Oliver PM, Bauer AM, Greenbaum E, Jackman T, Hobbie T: Molecularphylogenetics of the arboreal Australian gecko genus Oedura Gray 1842(Gekkota: Diplodactylidae): another plesiomorphic grade? Mol PhylogenetEvol 2012, 63:255–264.
58. Nielsen SV, Bauer AM, Jackman TR, Hitchmough RA, Daugherty CH: NewZealand geckos (Diplodactylidae): cryptic diversity in a post-Gondwananlineage with trans-Tasman affinities. Mol Phylogenet Evol 2011, 59:1–22.
59. Gamble T, Greenbaum E, Jackman TR, Russell AP, Bauer AM: Repeatedorigin and loss of adhesive toepads in geckos. PLoS One 2012, 7:e39429.
60. Wood PL Jr, Heinicke MP, Jackman TR, Bauer AM: Phylogeny of bent-toedgeckos (Cyrtodactylus) reveals a west to east pattern of diversification.Mol Phylogenet Evol 2012, 65:992–1003.
61. Giugliano LG, Collevatti RG, Colli GR: Molecular dating and phylogeneticrelationships among Teiidae (Squamata) inferred by molecular andmorphological data. Mol Phylogenet Evol 2007, 45:168–179.
62. Reeder TW, Cole CJ, Dessauer HC: Phylogenetic relationships of whiptaillizards of the genus Cnemidophorus (Squamata: Teiidae): a test ofmonophyly, reevalution of karyotypic evolution, and review of hybridorigins. Am Mus Novit 2002, 3365:1–61.
63. Pellegrino KCM, Rodrigues MT, Yonenaga-Yassuda Y, Sites JW: A molecularperspective on the evolution of microteiid lizards (Squamata,Gymnophthalmidae), and a new classification for the family. Biol J LinnSoc 2001, 74:315–338.
64. Castoe TA, Doan TM, Parkinson CL: Data partitions and complex models inBayesian analysis: the phylogeny of gymnophthalmid lizards. Syst Biol2004, 53:448–469.
65. Pavlicev M, Mayer W: Fast radiation of the subfamily Lacertinae (Reptilia:Lacertidae): history or methodical artefact? Mol Phylogenet Evol 2009,52:727–734.
66. Mayer W, Pavilcev M: The phylogeny of the family Lacertidae (Reptilia)based on nuclear DNA sequences: convergent adaptations to aridhabitats within the subfamily Eremiainae. Mol Phylogenet Evol 2007,44:1155–1163.
67. Fu JZ: Toward the phylogeny of the family Lacertidae: why 4708 basepairs of mtDNA sequences cannot draw the picture. Biol J Linn Soc 2000,71:203–217.
68. Arnold EN, Arribas O, Carranza S: Systematics of the Palaearctic andOriental lizard tribe Lacertini (Squamata: Lacertidae: Lacertinae), withdescriptions of eight new genera. Zootaxa 2007, 1430:1–86.
69. Greenbaum E, Villanueva CO, Kusamba C, Aristote MM, Branch WR: Amolecular phylogeny of equatorial African Lacertidae, with thedescription of a new genus and species from eastern DemocraticRepublic of the Congo. Zool J Linn Soc 2011, 163:913–942.
70. Mott T, Vieites DR: Molecular phylogenetics reveals extrememorphological homoplasy in Brazilian worm lizards challenging currenttaxonomy. Mol Phylogenet Evol 2009, 51:190–200.
71. Kearney M, Stuart BL: Repeated evolution of limblessness and diggingheads in worm lizards revealed by DNA from old bones. Proc R Soc B2004, 271:1677–1683.
72. Driskell AC, Ane C, Burleigh JG, McMahon MM, O'Meara BC, Sanderson MJ:Prospects for building the tree of life from large sequence databases.Science 2004, 306:1172–1174.
73. Wiens JJ, Fetzner JW, Parkinson CL, Reeder TW: Hylid frog phylogeny andsampling strategies for speciose clades. Syst Biol 2005, 54:719–748.
74. de Queiroz A, Gatesy J: The supermatrix approach to systematics. TrendsEcol Evol 2007, 22:34–41.
75. Thomson RC, Shaffer HB: Sparse supermatrices for phylogenetic inference:taxonomy, alignment, rogue taxa, and the phylogeny of living turtles.Syst Biol 2010, 59:42–58.
76. Pyron RA, Wiens JJ: A large-scale phylogeny of Amphibia including over2,800 species, and a revised classification of extant frogs, salamanders,and caecilians. Mol Phylogenet Evol 2011, 61:543–583.
77. Hinchcliff CE, Roalson EH: Using supermatrices for phylogenetic inquiry:an example using the sedges. Syst Biol 2013, 62:205–219.
78. Guo P, Liu Q, Xu Y, Jiang K, Hou M, Ding L, Pyron RA, Burbrink FT: Out ofAsia: natricine snakes support the Cenozoic Beringian DispersalHypothesis. Mol Phylogenet Evol 2012, 63:825–833.
79. Pyron RA, Burbrink FT: Neogene diversification and taxonomic stability inthe snake tribe Lampropeltini (Serpentes: Colubridae). Mol PhylogenetEvol 2009, 52:524–529.
80. Burbrink FT, Lawson R: How and when did Old World ratsnakes disperseinto the New World? Mol Phylogenet Evol 2007, 43:173–189.
81. Lawson R, Slowinski JB, Crother BI, Burbrink FT: Phylogeny of theColubroidea (Serpentes): new evidence from mitochondrial and nucleargenes. Mol Phylogenet Evol 2005, 37:581–601.
82. Wiens JJ, Kuczynski CA, Arif S, Reeder TW: Phylogenetic relationships ofphrynosomatid lizards based on nuclear and mitochondrial data, anda revised phylogeny for Sceloporus. Mol Phylogenet Evol 2010,54:150–161.
83. Lemmon AR, Brown JM, Stanger-Hall K, Lemmon EM: The effect ofambiguous data on phylogenetic estimates obtained by maximumlikelihood and Bayesian inference. Syst Biol 2009, 58:130–145.
84. Roure B, Baurain D, Philippe H: Impact of missing data on phylogenies inferredfrom empirical phylogenomic datasets. Mol Biol Evol 2013, 30:197–214.
85. Wiens JJ, Morrill MC: Missing data in phylogenetic analysis: reconcilingresults from simulations and empirical data. Syst Biol 2011, 60:719–731.
86. Pyron RA: Divergence time estimation using fossils as terminal taxa andthe origins of Lissamphibia. Syst Biol 2011, 60:466–481.
87. Wiens JJ, Tiu J: Highly incomplete taxa can rescue phylogenetic analysesfrom the negative impacts of limited taxon sampling. PLoS One 2012,7:e42925.
88. Scanlon JD, Lee MSY: The Pleistocene serpent Wonambi and the earlyevolution of snakes. Nature 2000, 403:416–420.
89. Tchernov E, Rieppel O, Zaher H, Polcyn MJ, Jacobs LL: A fossil snake withlimbs. Science 2000, 287:2010–2012.
90. Rage J-C: Serpentes. Stuttgart: Gustav Fischer Verlag; 1984.91. Estes R: Sauria terrestria, Amphisbaenia. Stuttgart: Gustav Fischer Verlag; 1983.

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 51 of 53http://www.biomedcentral.com/1471-2148/13/93
92. Holman JA: Fossil Snakes of North America: Origin, Evolution, Distribution,Paleoecology. Bloomington: Indiana University Press; 2000.
93. Rage J-C: Fossil history. In Snakes: Ecology and Evolutionary Biology. Editedby Seigel RA, Collins JT, Novak SS. New York: McMillan; 1987:51–76.
94. Anisimova M, Gascuel O: Approximate likelihood-ratio test for branches: afast, accurate, and powerful alternative. Syst Biol 2006, 55:539–552.
95. Lee MSY: Squamate phylogeny, taxon sampling, and data congruence.Org Div Evol 2005, 5:25–45.
96. Townsend TM, Leavitt DH, Reeder TW: Intercontinental dispersal by amicroendemic burrowing reptile (Dibamidae). Proc R Soc B 2011, 278:2568–2574.
97. Kluge AG: Cladistic relationships in the Gekkonoidea (Squamata, Sauria).Misc Pub Mus Zool Univ Mich 1987, 173:1–54.
98. Gamble T, Bauer AM, Greenbaum E, Jackman TR: Evidence for Gondwananvicariance in an ancient clade of gecko lizards. J Biogeogr 2008, 35:88–104.
99. Melville J, Schulte JA, Larson A: A molecular study of phylogeneticrelationships and evolution of antipredator strategies in AustralianDiplodactylus geckos, subgenus Strophurus. Biol J Linn Soc 2004, 82:123–138.
100. Jennings WB, Pianka ER, Donnellan S: Systematics of the lizard familyPygopodidae with implications for the diversification of Australiantemperate biotas. Syst Biol 2003, 52:757–780.
101. Bauer AM, Jackman TR, Sadlier RA, Whitaker AH: Revision of the giantgeckos of New Caledonia (Reptilia: Diplodactylidae: Rhacodactylus).Zootaxa 2012, 3404:1–52.
102. Brown RM, Siler CD, Das I, Min Y: Testing the phylogenetic affinities ofSoutheast Asia's rarest geckos: flap-legged geckos (Luperosaurus), flyinggeckos (Ptychozoon) and their relationship to the pan-Asian genusGekko. Mol Phylogenet Evol 2012, 63:915–921.
103. Vicario S, Caccone A, Gauthier J: Xantusiid "night" lizards: a puzzlingphylogenetic problem revisited using likelihood-based Bayesianmethods on mtDNA sequences. Mol Phylogenet Evol 2003, 26:243–261.
104. Hedges SB, Bezy RL, Maxson LR: Phylogenetic relationships andbiogeography of xantusiid lizards, inferred from mitochondrial DNAsequences. Mol Biol Evol 1991, 8:767–780.
105. Smith HM: The name of the non-nominotypical subfamily of the lizardfamily Xantusiidae. Syst Zool 1987, 36:326–328.
106. Savage JM: The Amphibians and Reptiles of Costa Rica: a Herpetofaunabetween Two Continents, between Two Seas. Chicago: University of ChicagoPress; 2002.
107. Crother BI, Miyamoto MM, Presch WF: Phylogeny and biogeography of thelizard family Xantusiidae. Syst Zool 1986, 35:37–45.
108. Stanley EL, Bauer AM, Jackman TR, Branch WR, Mouton PLFN: Between arock and a hard polytomy: rapid radiation in the rupicolous girdledlizards (Squamata: Cordylidae). Mol Phylogenet Evol 2011, 58:53–70.
109. Frost DR, Janies D, Mouton PIN, Titus TA: A molecular perspective on thephylogeny of the girdled lizards (Cordylidae, Squamata). Am Mus Novit2001, 3310:1–10.
110. Greer AE: A subfamilial classification of scincid lizards. Bull Mus Comp Zool1970, 139:151–183.
111. Smith SA, Sadlier RA, Bauer AM, Austin CC, Jackman T: Molecularphylogeny of the scincid lizards of New Caledonia and adjacent areas:evidence for a single origin of the endemic skinks of Tasmantis. MolPhylogenet Evol 2007, 43:1151–1166.
112. Hedges SB, Conn CE: A new skink fauna from Caribbean islands(Squamata, Mabuyidae, Mabuyinae). Zootaxa 2012, 3288:1–244.
113. Vences M, Guayasamin JM, Miralles A, de La Riva I: To name or not toname: criteria to promote economy of change in supraspecific Linnaeanclassification schemes. Zootaxa 2013, 3636:201–244.
114. Bauer AM: On the identity of Lacerta punctata Linnaeus, 1758, the typespecies of the genus Euprepis Wagler, 1830, and the generic assignmentof Afro-Malagasy skinks. Afr J Herpetol 2003, 52:1–7.
115. Austin JJ, Arnold EN: Using ancient and recent DNA to explorerelationships of extinct and endangered Leiolopisma skinks (Reptilia:Scincidae) in the Mascarene islands. Mol Phylogenet Evol 2006,39:503–511.
116. Skinner A: Phylogenetic relationships and rate of early diversification ofAustralian Sphenomorphus group scincids (Scincoidea, Squamata). Biol JLinn Soc 2007, 92:347–366.
117. Linkem CW, Diesmos AC, Brown RM: Molecular systematics of thePhilippine forest skinks (Squamata: Scincidae: Sphenomorphus): testingmorphological hypotheses of interspecific relationships. Zool J Linn Soc2011, 163:1217–1243.
118. Siler CD, Diesmos AC, Alcala AC, Brown RM: Phylogeny of Philippineslender skinks (Scincidae: Brachymeles) reveals underestimated speciesdiversity, complex biogeographical relationships, and cryptic patterns oflineage diversification. Mol Phylogenet Evol 2011, 59:53–65.
119. Brandley MC, Ota H, Hikida T, de Oca ANM, Feria-Ortiz M, Guo XG, Wang YZ:The phylogenetic systematics of blue-tailed skinks (Plestiodon) and thefamily Scincidae. Zool J Linn Soc 2012, 165:163–189.
120. Crottini A, Dordel J, Kohler J, Glaw F, Schmitz A, Vences M: A multilocusphylogeny of Malagasy scincid lizards elucidates the relationships of thefossorial genera Androngo and Cryptoscincus. Mol Phylogenet Evol 2009,53:345–350.
121. Sindaco R, Metallinou M, Pupin F, Fasola M, Carranza S: Forgotten in theocean: systematics, biogeography and evolution of the Trachylepis skinksof the Socotra Archipelago. Zool Scripta 2012, 41:346–362.
122. Miralles A, Fuenmayor GR, Bonillo C, Schargel WE, Barros T, Garcia-Perez JE,Barrio-Amoros CL: Molecular systematics of Caribbean skinks of thegenus Mabuya (Reptilia, Scincidae), with descriptions of two new speciesfrom Venezuela. Zool J Linn Soc 2009, 156:598–616.
123. Harvey MB, Ugueto GN, Gutberlet RL: Review of teiid morphology with arevised taxonomy and phylogeny of the Teiidae (Lepidosauria:Squamata). Zootaxa 2012, 3459:1–156.
124. Doan TM, Castoe TA: Phylogenetic taxonomy of the Cercosaurini(Squamata: Gymnophthalmidae), with new genera for species ofNeusticurus and Proctoporus. Zool J Linn Soc 2005, 143:405–416.
125. Vidal N, Azvolinsky A, Cruaud C, Hedges SB: Origin of tropical Americanburrowing reptiles by transatlantic rafting. Biol Lett 2008, 4:115–118.
126. Vidal N, Hedges SB: The molecular evolutionary tree of lizards, snakes,and amphisbaenians. CR Biol 2009, 332:129–139.
127. Lee MSY: Hidden support from unpromising data sets strongly unitessnakes with anguimorph 'lizards'. J Evol Biol 2009, 22:1308–1316.
128. Hugall AF, Foster R, Lee MSY: Calibration choice, rate smoothing, and thepattern of tetrapod diversification according to the long nuclear geneRAG-1. Syst Biol 2007, 56:543–563.
129. Macey JR, Schulte JA, Larson A, Tuniyev BS, Orlov N, Papenfuss TJ:Molecular phylogenetics, tRNA evolution, and historical biogeography inanguid lizards and related taxonomic families. Mol Phylogenet Evol 1999,12:250–272.
130. Conrad JL, Ast JC, Montanari S, Norell MA: A combined evidencephylogenetic analysis of Anguimorpha (Reptilia: Squamata). Cladistics2011, 27:230–277.
131. Collar DC, Schulte JA, Losos JB: Evolution of extreme body size disparityin monitor lizards (Varanus). Evolution 2011, 65:2664–2680.
132. Chippindale PT, Ammerman LK, Campbell JA: Molecular approaches tophylogeny of Abronia (Anguidae: Gerrhonotinae), with emphasis onrelationships in subgenus Auriculabronia. Copeia 1998, 4:883–892.
133. Conroy CJ, Bryson RW, Lazcano D, Knight A: Phylogenetic placement of thepygmy alligator lizard based on mitochondrial DNA. J Herp 2005, 39:142–147.
134. Okajima Y, Kumazawa Y: Mitochondrial genomes of acrodont lizards:timing of gene rearrangements and phylogenetic and biogeographicimplications. BMC Evol Biol 2010, 10:141.
135. Castoe TA, de Koning APJ, Kim HM, Gu WJ, Noonan BP, Naylor G, Jiang ZJ,Parkinson CL, Pollock DD: Evidence for an ancient adaptive episode ofconvergent molecular evolution. Proc Natl Acad Sci USA 2009, 106:8986–8991.
136. Raxworthy CJ, Forstner MRJ, Nussbaum RA: Chameleon radiation byoceanic dispersal. Nature 2002, 415:784–787.
137. Townsend TM, Vieites DR, Glaw F, Vences M: Testing species-leveldiversification hypotheses in Madagascar: the case of microendemicBrookesia leaf chameleons. Syst Biol 2009, 58:641–656.
138. Townsend TM, Tolley KA, Glaw F, Bohme W, Vences M: Eastward fromAfrica: palaeocurrent-mediated chameleon dispersal to the Seychellesislands. Biol Lett 2011, 7:225–228.
139. Tolley KA, Townsend TM, Vences M: Large-scale phylogeny of chameleonssuggests African origins and Eocene diversification. Proc R Soc B 2013,280:20130184.
140. Honda M, Ota H, Kobayashi M, Nabhitabhata J, Yong H-S, Sengoku S, HikidaT: Phylogenetic relationships of the family Agamidae (Reptilia: Iguania)inferred from mitochondrial DNA sequences. Zool Sci 2000, 17:527–537.
141. Hugall AF, Foster R, Hutchinson M, Lee MSY: Phylogeny of Australasianagamid lizards based on nuclear and mitochondrial genes: implicationsfor morphological evolution and biogeography. Biol J Linn Soc 2008,93:343–358.

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 52 of 53http://www.biomedcentral.com/1471-2148/13/93
142. Honda M, Ota H, Sengoku S, Yong HS, Hikida T: Molecular evaluation ofphylogenetic significances in the highly divergent karyotypes of thegenus Gonocephalus (Reptilia: Agamidae) from tropical Asia. Zool Sci2002, 19:129–133.
143. Baig KJ, Wagner P, Ananjeva NB, Bohme W: A morphology-basedtaxonomic revision of Laudakia Gray, 1845 (Squamata: Agamidae).Vertebr Zool 2012, 62:213–260.
144. Wilms TM, Bohme W, Wagner P, Lutzmann N, Schmitz A: On thephylogeny and taxonomy of the genus Uromastyx Merrem, 1820(Reptilia: Squamata: Agamidae: Uromastycinae): resurrection of thegenus Saara Gray, 1845. Bonner zoologische Beiträge 2009, 56:55–99.
145. Wagner P, Melville J, Wilms TM, Schmitz A: Opening a box of cryptic taxa:the first review of the North African desert lizards in the Trapelusmutabilis Merrem, 1820 complex (Squamata: Agamidae) withdescriptions of new taxa. Zool J Linn Soc 2011, 163:884–912.
146. Etheridge R, de Queiroz K: A phylogeny of Iguanidae. In PhylogeneticRelationships of the Lizard Families: Essays Commemorating Charles L Camp. Editedby Estes RD, Pregill GK. Stanford: Stanford University Press; 1988:283–367.
147. Frost DR, Etheridge R: A phylogenetic analysis and taxonomy of iguanianlizards (Reptilia: Squamata). Misc Publ Univ Kansas 1989, 81:1–65.
148. Macey JR, Larson A, Ananjeva NB, Papenfuss TJ: Evolutionary shifts in threemajor structural features of the mitochondrial genome among iguanianlizards. J Mol Evol 1997, 44:660–674.
149. Noonan BP, Chippindale PT: Vicariant origin of Malagasy reptiles supportslate Cretaceous antarctic land bridge. Am Nat 2006, 168:730–741.
150. Noonan BP, Sites JW: Tracing the origins of iguanid lizards and boinesnakes of the Pacific. Am Nat 2010, 175:61–72.
151. Schulte JA, Cartwright EM: Phylogenetic relationships among iguanianlizards using alternative partitioning methods and TSHZ1: a newphylogenetic marker for reptiles. Mol Phylogenet Evol 2009, 50:391–396.
152. Nicholson KE, Crother BI, Guyer C, Savage JM: It is time for a newclassification of anoles (Squamata: Dactyloidae). Zootaxa 2012, 3477:1–108.
153. Poe S: Phylogeny of anoles. Herp Monogr 2004, 18:37–89.154. Guyer C, Savage JM: Cladistic relationships among anoles (Sauria,
Iguanidae). Syst Zool 1986, 35:509–531.155. Losos JB: Lizards in an Evolutionary Tree: the Ecology of Adaptive Radiation in
Anoles. Berkeley: University of California Press; 2009.156. Cannatella DC, de Queiroz K: Phylogenetic systematics of the anoles: is a
new taxonomy warranted? Syst Zool 1989, 38:57–69.157. Poe S: 1986 redux: new genera of anoles (Squamata: Dactyloidae) are
unwarranted. Zootaxa 2013, 3626:295–299.158. Heise PJ, Maxson LR, Dowling HG, Hedges SB: Higher-level snake
phylogeny inferred from mitochondrial DNA sequences of 12S rRNA and16S rRNA genes. Mol Biol Evol 1995, 12:259–265.
159. Burbrink FT, Pyron RA: The taming of the skew: estimating properconfidence intervals for divergence dates. Syst Biol 2008, 57:317–328.
160. Pyron RA, Burbrink FT: Extinction, ecological opportunity, and the originsof global snake diversity. Evolution 2012, 66:163–178.
161. Gower DJ, Vidal N, Spinks JN, McCarthy CJ: The phylogenetic position ofAnomochilidae (Reptilia: Serpentes): first evidence from DNA sequences.J Zool Syst Evol Res 2005, 43:315–320.
162. Cadle JE, Dessauer HC, Gans C, Gartside DF: Phylogenetic relationships andmolecular evolution in uropeltid snakes (Serpentes, Uropeltidae):allozymes and albumin immunology. Biol J Linn Soc 1990, 40:293–320.
163. Wallach V, Wuster W, Broadley DG: In praise of subgenera: taxonomicstatus of cobras of the genus Naja Laurenti (Serpentes: Elapidae).Zootaxa 2009, 2236:26–36.
164. Williams D, Wuster W, Fry BG: The good, the bad and the ugly: Australiansnake taxonomists and a history of the taxonomy of Australia'svenomous snakes. Toxicon 2006, 48:919–930.
165. Rawlings LH, Rabosky DL, Donnellan SC, Hutchinson MN: Pythonphylogenetics: inference from morphology and mitochondrial DNA. BiolJ Linn Soc 2008, 93:603–619.
166. Burbrink FT: Inferring the phylogenetic position of Boa constrictor amongthe Boinae. Mol Phylogenet Evol 2005, 34:167–180.
167. Kluge AG: Boine snake phylogeny and research cycles. Misc Publ Mus ZoolUniv Mich 1991, 178:1–58.
168. Romer AS: Osteology of the Reptiles. Chicago: University of Chicago Press;1956.
169. Kelly CMR, Barker NP, Villet MH: Phylogenetics of advanced snakes(Caenophidia) based on four mitochondrial genes. Syst Biol 2003, 52:439–459.
170. Kraus F, Brown WM: Phylogenetic relationships of colubroid snakes basedon mitochondrial DNA sequences. Zool J Linn Soc 1998, 122:455–487.
171. Boulenger GA: Catalogue of snakes in the British Museum. London: BritishMuseum of Natural History; 1894.
172. Guo P, Wu Y, He S, Shi H, Zhao E: Systematics and molecular phylogeneticsof Asian snail-eating snakes (Pareatidae). Zootaxa 2011, 3001:57–64.
173. Garrigues T, Dauga C, Ferquel E, Choumet V, Failloux AB: Molecularphylogeny of Vipera Laurenti, 1768 and the related genera Macrovipera(Reuss, 1927) and Daboia (Gray, 1842), with comments about neurotoxicVipera aspis aspis populations. Mol Phylogenet Evol 2005, 35:35–47.
174. Lenk P, Kalyabina S, Wink M, Joger U: Evolutionary relationships amongthe true vipers (Reptilia: Viperidae) inferred from mitochondrial DNAsequences. Mol Phylogenet Evol 2001, 19:94–104.
175. Castoe TA, Parkinson CL: Bayesian mixed models and the phylogeny ofpitvipers (Viperidae: Serpentes). Mol Phylogenet Evol 2006, 39:91–110.
176. Fenwick AM, Gutberlet RL, Evans JA, Parkinson CL: Morphological andmolecular evidence for phylogeny and classification of South Americanpitvipers, genera Bothrops, Bothriopsis, and Bothrocophias (Serpentes:Viperidae). Zool J Linn Soc 2009, 156:617–640.
177. Kelly CMR, Branch WR, Broadley DG, Barker NP, Villet MH: Molecularsystematics of the African snake family Lamprophiidae Fitzinger,1843 (Serpentes: Elapoidea), with particular focus on the generaLamprophis Fitzinger 1843 and Mehelya Csiki 1903. Mol PhylogenetEvol 2011, 58:415–426.
178. Vidal N, Branch WR, Pauwels OSG, Hedges SB, Broadley DG, Wink M, CruaudC, Joger U, Nagy ZT: Dissecting the major African snake radiation: amolecular phylogeny of the Lamprophiidae Fitzinger (Serpentes,Caenophidia). Zootaxa 2008, 1945:51–66.
179. Sanders KL, Lee MSY, Leys R, Foster R, Keogh JS: Molecular phylogeny anddivergence dates for Australasian elapids and sea snakes (Hydrophiinae):evidence from seven genes for rapid evolutionary radiations. J Evol Biol2008, 21:682–695.
180. Sanders KL, Lee MSY: Molecular evidence for a rapid late-Mioceneradiation of Australasian venomous snakes (Elapidae, Colubroidea). MolPhylogenet Evol 2008, 46:1165–1173.
181. Chen X, Huang S, Guo P, Colli GR, de Oca AN M, Vitt LJ, Pyron RA, BurbrinkFT: Understanding the formation of ancient intertropical disjunctdistributions using Asian and Neotropical hinged-teeth snakes(Sibynophis and Scaphiodontophis: Serpentes: Colubridae). Mol PhylogenetEvol 2012, 66:254–261.
182. Huang S, Liu SY, Guo P, Zhang YP, Zhao EM: What are the closest relativesof the hot-spring snakes (Colubridae, Thermophis), the relict speciesendemic to the Tibetan Plateau? Mol Phylogenet Evol 2009, 51:438–446.
183. He M, Feng JC, Liu SY, Guo P, Zhao EM: The phylogenetic position ofThermophis (Serpentes: Colubridae), an endemic snake from theQinghai-Xizang Plateau, China. J Nat Hist 2009, 43:479–488.
184. Socha JJ: Gliding flight in Chrysopelea: turning a snake into a wing.Integr Comp Biol 2011, 51:969–982.
185. Helfenberger N: Phylogenetic relationship of Old World ratsnakes basedon visceral organ topography, osteology, and allozyme variation. Russ JHerp 2001, 8:1–56.
186. Alfaro ME, Arnold SJ: Molecular systematics and evolution of Regina andthe Thamnophiine snakes. Mol Phylogenet Evol 2001, 21:408–423.
187. de Queiroz A, Lawson R, Lemos-Espinal JA: Phylogenetic relationships ofNorth American garter snakes (Thamnophis) based on fourmitochondrial genes: How much DNA sequence is enough?Mol Phylogenet Evol 2002, 22:315–329.
188. Cope ED: Eleventh contribution to the herpetology of tropical America:Dugés, A. Proc Amer Philos Soc 1879, 18:261–277.
189. Fitzinger L: Systema Reptilium. Fasciculus Primus, Amblyglossae. Vienna: ApudBraumüller and Seidel Bibliopolas; 1843.
190. Vidal N, Dewynter M, Gower DJ: Dissecting the major American snakeradiation: a molecular phylogeny of the Dipsadidae Bonaparte(Serpentes, Caenophidia). CR Biol 2010, 333:48–55.
191. Caldwell MW: Squamate phylogeny and the relationships of snakes andmosasauroids. Zool J Linn Soc 1999, 125:115–147.
192. Heath TA, Hedtke SM, Hillis DM: Taxon sampling and the accuracy ofphylogenetic analyses. J Syst Evol 2008, 46:239–257.
193. Hedtke SM, Townsend TM, Hillis DM: Resolution of phylogenetic conflictin large data sets by increased taxon sampling. Syst Biol 2006,55:522–529.

Pyron et al. BMC Evolutionary Biology 2013, 13:93 Page 53 of 53http://www.biomedcentral.com/1471-2148/13/93
194. Rannala B, Huelsenbeck JP, Yang ZH, Nielsen R: Taxon sampling and theaccuracy of large phylogenies. Syst Biol 1998, 47:702–710.
195. Poe S, Swofford DL: Taxon sampling revisited. Nature 1999, 398:299–300.196. Fisher-Reid MC, Wiens JJ: What are the consequences of combining
nuclear and mitochondrial data for phylogenetic analysis? Lessons fromPlethodon salamanders and 13 other vertebrate clades. BMC Evol Biol2011, 11:300.
197. McMahon MM, Sanderson MJ: Phylogenetic supermatrix analysis ofGenBank sequences from 2228 papilionoid legumes. Syst Biol 2006,55:818–836.
198. Lemmon AR, Emme SA, Lemmon EM: Anchored hybrid enrichment formassively high-throughput phylogenomics. Syst Biol 2012, 61:727–744.
199. Faircloth BC, McCormack JE, Crawford NG, Harvey MG, Brumfield RT, GlennTC: Ultraconserved elements anchor thousands of genetic markersspanning multiple evolutionary timescales. Syst Biol 2012, 61:717–726.
200. Crawford NG, Faircloth BC, McCormack JE, Brumfield D, Winker K, Glenn TC:More than 1000 ultraconserved elements provide evidence that turtlesare the sister group of archosaurs. Biol Lett 2012, 8:783–786.
201. Edwards SV, Liu L, Pearl DK: High-resolution species trees withoutconcatenation. Proc Natl Acad Sci USA 2007, 104:5936–5941.
202. Heled J, Drummond AJ: Bayesian inference of species trees frommultilocus data. Mol Biol Evol 2010, 27:570–580.
203. Ricklefs RE, Losos JB, Townsend TM: Evolutionary diversification of cladesof squamate reptiles. J Evol Biol 2007, 20:1751–1762.
204. Brandley MC, Huelsenbeck JP, Wiens JJ: Rates and patterns in theevolution of snake-like body form in squamate reptiles: evidence forrepeated re-evolution of lost digits and long-term persistence ofintermediate body forms. Evolution 2008, 62:2042–2064.
205. Pyron RA, Burbrink FT: Systematics of the Common Kingsnake(Lampropeltis getula; Serpentes: Colubridae) and the burden of heritagein taxonomy. Zootaxa 2009, 2241:22–32.
206. Pyron RA: A likelihood method for assessing molecular divergence timeestimates and the placement of fossil calibrations. Syst Biol 2010, 59:185–194.
207. Edgar RC: MUSCLE: multiple sequence alignment with high accuracy andhigh throughput. Nucleic Acids Res 2004, 32:1792–1797.
208. Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliamH, Valentin F, Wallace IM, Wilm A, Lopez R, et al: Clustal W and Clustal Xversion 2.0. Bioinformatics 2007, 23:2947–2948.
209. Stamatakis A, Aberer AJ, Goll C, Smith SA, Berger SA, Izquierdo-Carrasco F:RAxML-Light: a tool for computing terabyte phylogenies. Bioinformatics2012, 28:2064–2066.
210. Stamatakis A: RAxML-VI-HPC: maximum likelihood-based phylogeneticanalyses with thousands of taxa and mixed models. Bioinformatics 2006,22:2688–2690.
211. Felsenstein J: Inferring Phylogenies. Sunderland: Sinauer Associates; 2004.212. Anisimova M, Gil M, Dufayard JF, Dessimoz C, Gascuel O: Survey of branch
support methods demonstrates accuracy, power, and robustness of fastlikelihood-based approximation schemes. Syst Biol 2011, 60:685–699.
213. Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O: Newalgorithms and methods to estimate maximum-likelihood phylogenies:assessing the performance of PhyML 3.0. Syst Biol 2010, 59:307–321.
214. Schneider JG: Historiae Amphibiorum Naturalis et Literariae. FasciculusSecundus continens Crocodilos, Scincos, Chamaesauras, Boas, Pseudoboas,Elapes, Angues, Amphisbaenas et Caecilias. Jena: Frommani; 1801.
215. McDowell SB: A catalog of the snakes of New Guinea and the Solomons,with special reference to those in the Bernice P. Bishop museum. Part III.Boinae and Acrochordoidea (Reptilia, Serpentes). J Herp 1979, 13:1–92.
doi:10.1186/1471-2148-13-93Cite this article as: Pyron et al.: A phylogeny and revised classification ofSquamata, including 4161 species of lizardsand snakes. BMC Evolutionary Biology 2013 13:93.
Submit your next manuscript to BioMed Centraland take full advantage of:
• Convenient online submission
• Thorough peer review
• No space constraints or color figure charges
• Immediate publication on acceptance
• Inclusion in PubMed, CAS, Scopus and Google Scholar
• Research which is freely available for redistribution
Submit your manuscript at www.biomedcentral.com/submit
Related Documents