RESEARCH POSTER PRESENTATION DESIGN © 2012 www.PosterPresentation s.com Ventricular septal defects (VSDs) are the most common congenital heart defects reported in horses. VSDs are characterized by an abnormal communication between the left and right ventricles that varies in size, location, and clinical relevance. Even small, high velocity defects known as restrictive VSDs can lead to impaired cardiac function if the cusps of the aortic valve are drawn into the defect, resulting in aortic insufficiency. When severe, these lesions result in volume overload in the left heart and ultimately congestive heart failure. Arabian horses are overrepresented for incidence of congenital heart defects including VSDs, and a genetic basis for VSDs in the breed is considered likely. Identification of genetic markers associated with VSDs would facilitate the isolation of a VSD causative mutation and ultimately enable the development of a genetic test for Arabian breeding managers to promote the health of the breed and limit the prevalence of this defect. Introduction Cardiac auscultation and echocardiographic evaluations were used to phenotype 13 affected and 38 normal Arabian horses. DNA was extracted from whole blood samples and genotyped using the Axiom® Equine Genotyping Array that contains 670,000 SNPs across the equine genome. Materials and Methods Ventricular septal defects (VSDs) in Arabian horses are associated with a single genetic locus identifiable through a genome-wide association study (GWAS). Hypothesis Figure 3. Manhattan plot illustrating the most significant SNP association on chromosome 25 in the EMMAX analysis. Bonferroni adjusted and raw P values are displayed. Results Discussion Despite the suspected heritability of VSDs in Arabian horses, no prior studies have undertaken to identify SNPs, haplotypes, and genes associated with VSDs in this breed. Case vs. control GWAS was successful in identifying a chromosomal region of interest. Chromosome 25 has the strongest association with VSDs and withstood Bonferroni correction. Phenotypic diversity is a well-known feature of VSDs in many species. A specific form of VSD as a component of a more severe congenital malformation known as tetralogy of Fallot is rarely diagnosed in Arabian horses and other horse breeds. Our data suggests that tetralogy of Fallot is genotypically distinct from the VSD variety tested in this study (perimembranous). Further defining the identified region of interest and investigating variants in candidate genes through whole genome sequencing will ideally guide future translational research and aid in the understanding of VSD pathogenesis. References 1. Leroux, A. A., et al. "Prevalence and Risk Factors for Cardiac Diseases in a Hospital-‐-Based Population of 3,434 Horses (1994–2011)." Journal of Veterinary Internal Medicine 27.6 (2013): 1563-1570. 2. Hall, T. L., K. G. Magdesian, and M. D. Kittleson. "Congenital cardiac defects in neonatal foals: 18 cases (1992–2007)." Journal of veterinary internal medicine 24.1 (2010): 206-212. 3. Thomas, Bill. "The Equine Heart." CEH Horse Report: A Publication of the Center for Equine Health, UC Davis School of Veterinary Medicine 24.4 (2006): 1-12. Center for Equine Health. UC Davis School of Veterinary Medicine. Web. 10 Jan. 2015. <http://www.vetmed.ucdavis.edu/ceh/>. 4. Kahn, Cynthia M. "Congenital and Inherited Disorders of the Cardiovascular System in Horses." The Merck/Merial Manual for Pet Health. Home ed. Whitehouse Station, NJ: Merck, 2007. Print. 5. Norton, Elaine. "Genome Wide Association Study for Equine Exertional Rhabdomyolysis in Standardbred Horses." Plant and Animal Genome XXII Conference. Plant and Animal Genome, 2014. Acknowledgements Department of Medicine & Epidemiology, School of Veterinary Medicine, University of California Davis E Morgan, E Ontiveros, K Estell, J Stern Identification of Genetic Markers for Ventricular Septal Defects in Arabian Horses Location Unadjusted P Bonferroni P Chr25.18647558 4.14E-011 1.30E-05 Chr25.18658851 4.70E-09 0.0015 Chr25.18659166 4.70E-09 0.0015 Mixed linear model analysis (EMMAX) was performed to further reduce the effects of population stratification. QQ plot improved: results displayed. Chromosome 25 significance persisted after 330,000 permutations. Figure 1. A 2D and color Doppler echocardiogram still image is provided of a 2 year old Arabian gelding. The image is obtained from the right thorax. The VSD (arrow) is visualized with turbulent flow passing into the right ventricle, just below the aortic valve (asterisk). Table 1. The SNPs most strongly associated with VSDs from the EMMAX analysis are detailed by location in EquCab2 and test statistics. Figure 4. Additive association model with 330,000 permutations. The pink line denotes a genome-wide significance value of P=0.05. Phenotype Auscultation: A VSD is associated with two possible heart murmurs. The shunt itself causes a loud, holosytolic murmur with a point of maximal intensity over the right thorax. A murmur of relative pulmonic stenosis is often auscultated as well. Though the right ventricular outflow tract may be anatomically normal, the increased volume of blood leaving the right ventricle results in a holosystolic crescendo- decrescendo murmur most audible over the pulmonary valve region on the left thorax. Echocardiogram: Echocardiography is used to definitively diagnose a VSD and classify its location. With cardiac ultrasound, the defect is visualized as a communication between the two ventricles. The velocity and direction of flow through the shunt is measured to determine if the defect is restrictive (hemodynamically insignificant) or non-restrictive. The downstream effects of volume overload, such as dilation of the pulmonary artery and left atrium and eccentric hypertrophy of the left ventricle, are assessed. This study targets perimembranous VSDs. Extracted DNA from whole EDTA blood and formalin-fixed, paraffin embedded tissue Hybridized DNA of 12 cases and 39 controls to Axiom® Equine Genotyping Arrays Conducted Genome- Wide Association Study (GWAS) and quality control using Golden Helix Software 37,414 unmapped SNPs were excluded for analysis Quality control removed an additional 257,795 SNPs (minor allele frequency inclusion threshold of 0.08, maximum per individual missing genotype rate of 5%, and maximum Hardy Weinberg Equilibrium P-value of 0.001) 375,587 SNPS were retained and analyzed after quality control and exclusion of unmapped SNPs. 5 cases and 8 controls were removed based on MDS plot indicating population stratification. Case vs. control association with 8 cases and 30 controls yielded a genome inflation factor of 1.28 and identified a tightly clustered region of association on Chromosome 25. The top SNP identified on Chromosome 25 yielded a Bonferroni P value of 1.30E-05 Figure 5. QQ Plot demonstrating that population stratification has been corrected after adjustment by MDS and EMMAX. Figure 7. Healthy 2 month old Arabian colt. Figure 4. QQ Plot prior to adjustment for population stratification (MDS and EMMAX; Genome inflation factor=3.43). (Praw=4.14E-011) (Padj=1.30E-05) Figure 2. Spectral Doppler echocardiographic image depicts blood flow across the VSD. The flow measures 5.5 m/sec toward the transducer in systole. This confirms the shunt direction is left to right and the velocity confirms that this VSD is hemodynamically insignificant (restrictive). P=0.026 Figure 6. Haplotype analysis. Complex VSD represents tetralogy of Fallot- a genetically distinct disease complex in humans.

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

RESEARCH POSTER PRESENTATION DESIGN © 2012
www.PosterPresentations.com
QU ICK START ( con t . )
How to change the template color
theme You can easily change the color theme of your poster
by going to the DESIGN menu, click on COLORS, and
choose the color theme of your choice. You can also
create your own color theme.
You can also manually change the color of your
background by going to VIEW > SLIDE MASTER. After
you finish working on the master be sure to go to
VIEW > NORMAL to continue working on your poster.
How to add Text The template comes with a
number of pre-formatted
placeholders for headers and
text blocks. You can add
more blocks by copying and
pasting the existing ones or
by adding a text box from the
HOME menu.
Text size Adjust the size of your text based on how much
content you have to present. The default template
text offers a good starting point. Follow the
conference requirements.
How to add Tables To add a table from scratch go to the
INSERT menu and
click on TABLE. A drop-down box will
help you select rows and columns.
You can also copy and a paste a table from Word or
another PowerPoint document. A pasted table may
need to be re-formatted by RIGHT-CLICK > FORMAT
SHAPE, TEXT BOX, Margins.
Graphs / Charts You can simply copy and paste charts and graphs
from Excel or Word. Some reformatting may be
required depending on how the original document
has been created.
How to change the column
configuration RIGHT-CLICK on the poster background and select
LAYOUT to see the column options available for this
template. The poster columns can also be
customized on the Master. VIEW > MASTER.
How to remove the info bars If you are working in PowerPoint for Windows and
have finished your poster, save as PDF and the bars
will not be included. You can also delete them by
going to VIEW > MASTER. On the Mac adjust the Page-
Setup to match the Page-Setup in PowerPoint before
you create a PDF. You can also delete them from the
Slide Master.
Save your work Save your template as a PowerPoint document. For
printing, save as PowerPoint of “Print-quality” PDF.
Print your poster When you are ready to have your poster printed go
online to PosterPresentations.com and click on the
“Order Your Poster” button. Choose the poster type
the best suits your needs and submit your order. If
you submit a PowerPoint document you will be
receiving a PDF proof for your approval prior to
printing. If your order is placed and paid for before
noon, Pacific, Monday through Friday, your order will
ship out that same day. Next day, Second day, Third
day, and Free Ground services are offered. Go to
PosterPresentations.com for more information.
Student discounts are available on our Facebook
page.
Go to PosterPresentations.com and click on the
FB icon.
© 2013 PosterPresentations.com 2117 Fourth Street , Unit C Berkeley CA 94710
(—THIS SIDEBAR DOES NOT PRINT—)
DES IG N G U IDE
This PowerPoint 2007 template produces a
36”x48” presentation poster. You can use it
to create your research poster and save
valuable time placing titles, subtitles, text,
and graphics.
We provide a series of online tutorials that
will guide you through the poster design
process and answer your poster production
questions. To view our template tutorials, go
online to PosterPresentations.com and click
on HELP DESK.
When you are ready to print your poster, go
online to PosterPresentations.com
Need assistance? Call us at 1.510.649.3001
QU ICK START
Zoom in and out As you work on your poster zoom in
and out to the level that is more
comfortable to you.
Go to VIEW > ZOOM.
Title, Authors, and Affiliations Start designing your poster by adding the title, the
names of the authors, and the affiliated institutions.
You can type or paste text into the provided boxes.
The template will automatically adjust the size of
your text to fit the title box. You can manually
override this feature and change the size of your
text.
TIP: The font size of your title should be bigger than
your name(s) and institution name(s).
Adding Logos / Seals Most often, logos are added on each side of the title.
You can insert a logo by dragging and dropping it
from your desktop, copy and paste or by going to
INSERT > PICTURES. Logos taken from web sites are
likely to be low quality when printed. Zoom it at
100% to see what the logo will look like on the final
poster and make any necessary adjustments.
TIP: See if your school’s logo is available on our
free poster templates page.
Photographs / Graphics You can add images by dragging and dropping from
your desktop, copy and paste, or by going to INSERT
> PICTURES. Resize images proportionally by holding
down the SHIFT key and dragging one of the corner
handles. For a professional-looking poster, do not
distort your images by enlarging them
disproportionally.
Image Quality Check Zoom in and look at your images at 100%
magnification. If they look good they will print well.
ORIGINAL
DISTORTED
Corner handles
Go
od
pri
nti
ng
qu
alit
y
Bad
pri
nti
ng
qu
alit
y
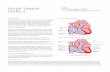
Ventricular septal defects (VSDs) are the most common congenital heart defects reported in horses. VSDs are characterized by an abnormal communication between the left and right ventricles that varies in size, location, and clinical relevance. Even small, high velocity defects known as restrictive VSDs can lead to impaired cardiac function if the cusps of the aortic valve are drawn into the defect, resulting in aortic insufficiency. When severe, these lesions result in volume overload in the left heart and ultimately congestive heart failure.
Arabian horses are overrepresented for incidence of congenital heart defects including VSDs, and a genetic basis for VSDs in the breed is considered likely. Identification of genetic markers associated with VSDs would facilitate the isolation of a VSD causative mutation and ultimately enable the development of a genetic test for Arabian breeding managers to promote the health of the breed and limit the prevalence of this defect.
Introduction
Cardiac auscultation and echocardiographic evaluations were used to phenotype 13 affected and 38 normal Arabian horses. DNA was extracted from whole blood samples and genotyped using the Axiom® Equine Genotyping Array that contains 670,000 SNPs across the equine genome.
Materials and Methods
Ventricular septal defects (VSDs) in Arabian horses are associated with a single genetic locus identifiable through a genome-wide association study (GWAS).
Hypothesis
Figure 3. Manhattan plot illustrating the most significant SNP association on chromosome 25 in the EMMAX analysis. Bonferroni adjusted and raw P values are displayed.
Results
Discussion Despite the suspected heritability of VSDs in Arabian horses, no prior studies have undertaken to identify SNPs, haplotypes, and genes associated with VSDs in this breed. Case vs. control GWAS was successful in identifying a chromosomal region of interest.
Chromosome 25 has the strongest association with VSDs and withstood Bonferroni correction. Phenotypic diversity is a well-known feature of VSDs in many species. A specific form of VSD as a component of a more severe congenital malformation known as tetralogy of Fallot is rarely diagnosed in Arabian horses and other horse breeds. Our data suggests that tetralogy of Fallot is genotypically distinct from the VSD variety tested in this study (perimembranous).
Further defining the identified region of interest and investigating variants in candidate genes through whole genome sequencing will ideally guide future translational research and aid in the understanding of VSD pathogenesis.
References 1. Leroux, A. A., et al. "Prevalence and Risk Factors for Cardiac Diseases in a Hospital-‐-Based Population of 3,434 Horses (1994–2011)." Journal of Veterinary Internal Medicine 27.6 (2013): 1563-1570.
2. Hall, T. L., K. G. Magdesian, and M. D. Kittleson. "Congenital cardiac defects in neonatal foals: 18
cases (1992–2007)." Journal of veterinary internal medicine 24.1 (2010): 206-212.
3. Thomas, Bill. "The Equine Heart." CEH Horse Report: A Publication of the Center for Equine Health,
UC Davis School of Veterinary Medicine 24.4 (2006): 1-12. Center for Equine Health. UC Davis School
of Veterinary Medicine. Web. 10 Jan. 2015. <http://www.vetmed.ucdavis.edu/ceh/>.
4. Kahn, Cynthia M. "Congenital and Inherited Disorders of the Cardiovascular System in Horses." The
Merck/Merial Manual for Pet Health. Home ed. Whitehouse Station, NJ: Merck, 2007. Print.
5. Norton, Elaine. "Genome Wide Association Study for Equine Exertional Rhabdomyolysis in Standardbred Horses." Plant and Animal Genome XXII Conference. Plant and Animal Genome, 2014.
Acknowledgements
Department of Medicine & Epidemiology, School of Veterinary Medicine, University of California Davis
E Morgan, E Ontiveros, K Estell, J Stern
Identification of Genetic Markers for Ventricular Septal Defects in Arabian Horses
Location Unadjusted P Bonferroni P
Chr25.18647558 4.14E-011 1.30E-05
Chr25.18658851 4.70E-09 0.0015
Chr25.18659166 4.70E-09 0.0015
Mixed linear model analysis (EMMAX) was performed to further reduce the effects of population stratification. QQ plot improved: results displayed. Chromosome 25 significance persisted after 330,000 permutations.
Figure 1. A 2D and color Doppler echocardiogram still image is provided of a 2 year old Arabian gelding. The image is obtained from the right thorax. The VSD (arrow) is visualized with turbulent flow passing into the right ventricle, just below the aortic valve (asterisk).
Table 1. The SNPs most strongly associated with VSDs from the EMMAX analysis are detailed by location in EquCab2 and test statistics.
Figure 4. Additive association model with 330,000 permutations. The pink line denotes a genome-wide significance value of P=0.05.
Phenotype Auscultation: A VSD is associated with two possible heart murmurs. The shunt itself causes a loud, holosytolic murmur with a point of maximal intensity over the right thorax. A murmur of relative pulmonic stenosis is often auscultated as well. Though the right ventricular outflow tract may be anatomically normal, the increased volume of blood leaving the right ventricle results in a holosystolic crescendo-decrescendo murmur most audible over the pulmonary valve region on the left thorax.
Echocardiogram: Echocardiography is used to definitively diagnose a VSD and classify its location. With cardiac ultrasound, the defect is visualized as a communication between the two ventricles. The velocity and direction of flow through the shunt is measured to determine if the defect is restrictive (hemodynamically insignificant) or non-restrictive. The downstream effects of volume overload, such as dilation of the pulmonary artery and left atrium and eccentric hypertrophy of the left ventricle, are assessed. This study targets perimembranous VSDs.
Extracted DNA from whole EDTA
blood and formalin-fixed,
paraffin embedded tissue
Hybridized DNA of 12 cases and 39 controls to
Axiom® Equine Genotyping
Arrays
Conducted Genome-Wide Association Study (GWAS) and
quality control using Golden Helix
Software
37,414 unmapped SNPs were
excluded for analysis
Quality control removed an additional 257,795 SNPs (minor allele frequency inclusion threshold of 0.08, maximum
per individual missing genotype rate of 5%, and maximum Hardy Weinberg
Equilibrium P-value of 0.001)
375,587 SNPS were retained and analyzed after quality control
and exclusion of unmapped SNPs.
5 cases and 8 controls were
removed based on MDS plot indicating
population stratification.
Case vs. control association with 8 cases
and 30 controls yielded a genome inflation factor of
1.28 and identified a tightly clustered region of
association on Chromosome 25.
The top SNP identified on
Chromosome 25 yielded a
Bonferroni P value of 1.30E-05
Figure 5. QQ Plot demonstrating that population stratification has been corrected after adjustment by MDS and EMMAX.
Figure 7. Healthy 2 month old Arabian colt.
Figure 4. QQ Plot prior to adjustment for population stratification (MDS and EMMAX; Genome inflation factor=3.43).
(Praw=4.14E-011) (Padj=1.30E-05)
Figure 2. Spectral Doppler echocardiographic image depicts blood flow across the VSD. The flow measures 5.5 m/sec toward the transducer in systole. This confirms the shunt direction is left to right and the velocity confirms that this VSD is hemodynamically insignificant (restrictive).
P=0.026
Figure 6. Haplotype analysis. Complex VSD represents tetralogy of Fallot- a genetically distinct disease complex in humans.
Related Documents