1 Tricks for trees: Having reconstructed Tricks for trees: Having reconstructed phylogenies phylogenies what can we do with them? what can we do with them? DIMACS, June 2006 Mike Steel Mike Steel Allan Wilson Centre for Allan Wilson Centre for Molecular Ecology and Evolution Molecular Ecology and Evolution Biomathematics Research Centre Biomathematics Research Centre University of Canterbury, University of Canterbury, Christchurch, New Zealand Christchurch, New Zealand

1 Tricks for trees: Having reconstructed phylogenies what can we do with them? DIMACS, June 2006 Mike Steel Allan Wilson Centre for Molecular Ecology and.
Dec 22, 2015
Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript
1
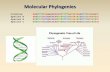
Tricks for trees: Having reconstructed Tricks for trees: Having reconstructed phylogenies phylogenies what can we do with them?what can we do with them?
DIMACS, June 2006
Mike Steel Mike Steel
Allan Wilson Centre for Allan Wilson Centre for Molecular Ecology and EvolutionMolecular Ecology and EvolutionBiomathematics Research CentreBiomathematics Research Centre
University of Canterbury, University of Canterbury, Christchurch, New ZealandChristchurch, New Zealand
2
Where are phylogenetic trees used? Evolutionary biology – species
relationships, dating divergences, speciation processes, molecular evolution.
Ecology – classifying new species; biodiversity, co-phylogeny, migration of populations.
Epidemiology – systematics, processes, dynamics
Extras - linguistics, stematology, psychology.
3
Phylogenetic trees[Definition] A phylogenetic X-tree is a tree T=(V,E) with a set X of labelled leaves, and all other vertices unlabelled and of degree >3.
If all non-leaf vertices have degree 3 then T is binary
4
Trees and splits
}:|{)( EeBAT ee
ee BAe |
),( XP
1
2
3
45
6
Partial order:
)'()(' TTTT
Buneman’s Theorem
5
Quartet trees• A quartet tree is a binary phylogenetic tree on 4 leaves (say, x,y,w,z) written xy|wz.
• A phylogenetic X-tree displays xy|wz if there is an edge in T whose deletion separates {x,y} from {w,z}
x
y
w
z
ry
zu
x
s
w
7
How are trees useful in epidemiology?
Systematics and reconstruction
How are different types/strains of a virus related?
When, where, and how did they arise?
What is their likely future evolution?
What was the ancestral sequence?
8
How are trees useful in epidemiology?
Processes and dynamics (“Phylodynamics”)
How do viruses change with time in a population? Population size etc
What is their rate of mutation, recombination, selection?
Within-host dynamcs
How do viruses evolve in a single patient?
How is this related to the progression of the disease?
How much compartmental variation exists?
10
What do the shapes of these trees tell us about the processes governing their evolution?
Eg. Population dynamics, selection
Coalescent prediction
13
Why do trees on the same taxa disagree?
1. Model violation1. “true model” differs from “assumed model”2. “true model = assumed model” but estimation method
not appropriate to model 3. model true but too parameter rich (non-identifyability)
2. Sampling error (and factors that make it worse!)3. Alignment error4. Evolutionary processes
1. Lineage sorting 2. Recombination3. Horizontal gene transfer; hybrid taxa4. Gene duplication and loss
15
Example: Deep divergence in the Metazoan
phylogeny
Fungi
Choanoflagellates
Arthropods
Nematodes
Deuterostomes
Platyhelminthes
ActinopterMammaliaCnidaria
Monosiga ovata
CryptococcusPhanerochaete
Ustilago
Schizosaccharomyces
Saccharomyces
Candida
Paracooccidioides
Gibberella
MagnaporthNeurospora
Glomus
Neocallimastix
Schistosoma mansoniSchistosoma japonicum
FasciolaEchinococcus
Dugesia
Strongyloides
Caenorhabditis briggsaeCaenorhabditis elegans
AncylostomaPristionchus
Brugia
Ascaris
Heterodera
Trichinella
Glossina
DrosophilaAnopheles
Monosiga brevicollis
Urochordata
Echinodermata
Ctenophora
Meloidogyne
Tardigrades
Chelicerata
HemipteraHymenoptera
Coleoptera
SiphonapteraLepidoptera
Crustacea
AnnelidaMolluscaCephalochordata
From Huson and Bryant, 2006
17
Models
1
2
3
4
vs
1
2
3
4
•“site saturation”
• subdividing long edges only offers a partial remedy (trade-off).
18
Why do trees on the same taxa disagree?
1. Model violation1. “true model” differs from “assumed model”2. “true model = assumed model” but estimation method
not appropriate to model3. model true but too parameter rich (non-identifyability)
2. Sampling error (and factors that make it worse!)3. Alignment4. Evolutionary processes
1. Lineage sorting 2. Recombination3. Horizontal gene transfer; hybrid taxa4. Gene duplication and loss
19
Gene trees vs species trees
Theorem J. H. Degnan and N.A. Rosenberg, 2006.
For n>5, for any tree, there are branch lengths and population sizes for which the most likely gene tree is different from the species tree.
Discordance of species trees with their most likely gene trees. PLoS Genetics, 2(5), e68 May, 2006
a b c a b c
20
Example
Orangutan Gorilla Chimpanzee Human
Adapted From the Tree of the Life Website,University of Arizona
?
22
Why do trees on the same taxa disagree?
1. Model violation1. “true model” differs from “assumed model”2. “true model = assumed model” but estimation method
not appropriate to model3. model true but too parameter rich (non-identifyability)
2. Sampling error (and factors that make it worse!)3. Alignment4. Evolutionary processes
1. Lineage sorting 2. Recombination3. Horizontal gene transfer; hybrid taxa4. Gene duplication and loss
23
Given a tree what questions might we want to answer?
How reliable is a split? Where is the root of the tree? Relative ranking of vertices?
Dating? How well supported is some ‘deep divergence’ resolved? What model best describes the evolution of the sequences
(molecular clock? dS/dN ratio constant? etc)
Statistical approaches: Non-parametric bootstrap Parametric bootstrap Likelihood ratio tests Bayesian posterior probabilities Tests (KH, SH, SOWH)Goldman, N., J. P. Anderson, and A. G. Rodrigo. 2000. Likelihood-based tests of topologies in phylogenetics. Systematic Biology 49: 652-670.
29
Consensus networks
Take the splits that are in at least x% of the trees and represent them by a graph
Splits Graph (G()) – Dress and HusonEach split is represented by a class of ‘parallel’ edges
Simplest example (n=4).
30
R.nivicola
(NS)
(SS)
(SS)
(NS)
(SS)
(NS)
(NS)
(SS)
chloroplast
JSA tree
(A)(NS)
(SS)
(N,NS)
(A)
(SS)
(SS)
(SS)
(SS) (NS)
(C,S)
(NS)
(N)
(NS, N)
31
R.nivicola
nuclear
ITS tree
(SS)
(NS)
(NS)
(SS)
(NS) (SS)
(SS)
(NS)
(SS)
(SS) (SS)
(SS)
(NS)
(SS)
(NS,N) (NS)
(NS) (SS)
(NS,N)
(A)
(A)
(N)
(SS,NS)
34
Comparing trees
Splits metric (Robinson-Foulds)
Statistical aspects.
Tree rearrangement operations – the graph of
trees (rSPR).
Cophylogeny
36
Supertrees
Compatibility concept Compatibility of rooted trees (BUILD) Why do we want to do this? Extension – higher order taxa, dates Methods for handling incompatible trees(MRP; mincut variants; minflip)
37
Compatibility
Example: Q={12|34, 13|45, 14|26}
1
2
3
4
5
6
A set Q of quartets is compatible if there is a phylogenetic X-tree T that displays each quartet of Q
Complexity?
38
Supertrees
Compatibility concept Compatibility of rooted trees (BUILD) Why do we want to do this? Extension – higher order taxa, dates Methods for handling incompatible trees(MRP; mincut variants; minflip)
39
Phylogenetic networks
Consensus setting: consensus networks Minimizing hybrid/reticulate vertices Supernetworks – Z closure, filtering
40
a
Networks can represent: Reticulate evolution (eg. hybrid species) Phylogenetic uncertainty (i.e. possible alternative trees)
Z-closure Given T1,…, Tk on overlapping sets of species,
let construct spcl2() and construct the ‘splits graph’ of the resulting splits that are ‘full’.
cb d a bc d a cbd
)()( 1 kTT
41
AA22
BB22
Split closure operation (Meacham 1986)
AA11
BB11
AA22
BB22
AA11
BB11UUBB22
AA11UUAA22
BB22
,,
AA11
BB11
44
Reconstructing ancestral sequences
Methods (MP, Likelihood, Bayesian)
Quiz. MP for a balanced tree = majority state?
Information-theoretic considerations
Related Documents