SPM MATLAB TOOLBOX J. Tauchmanov´ a, M. Hromˇ c´ ık, R. Jech Czech Technical University, Faculty of Electrical Engineering, Department of Control Engineering, Department of Neurology, First Faculty of Medicine, Prague Abstract The SPM toolbox is a noncommercial software for processing fMRI data. The toolbox is developed at the Department of Imaging Neuroscience, University College London. The common application of the SPM toolbox is the first level analysis consisting in the detection of active brain regions for one patient. The second level analysis, also covered by the SPM toolbox, is related to processing of fMRI data across the whole patients group. Besides these functions, the toolbox offers tools for DCM analysis, such as the parameters estimation routines, tools for comparison of resulting models, and a function for averaging models across the whole patients group. 1 Introduction SPM toolbox serves for data processing, especially for fMRI, EEG and MEG data sets. It in- cludes preprocessing steps, statistical analysis and DCM analysis. There exist several versions of this toolbox. The oldest available version was called SPM99, then SPM2 followed and nowadays version SPM5 is used. fMRI abbreviates functional Magnetic Resonance Imaging. This method is able to create map of brain activity. The results of fMRI experiment are functional images and structural anatomical images which are useful as a base for localization of activated regions. The functional images are put through the statistical analysis and the result is statistically significant regions where the so-called haemodynamic responses were detected. But there appear some complications during this procedure for example some statistical discrepancies. The SPM toolbox try to correct them and to produce regular results. 2 fMRI analysis 2.1 Preprocessing The first stage of fMRI data analysis is data preprocessing - preparation of data for statistical analysis. SPM toolbox offers these adjustments. Realigning corrects movement artifacts in data. Least squares approach and spatial transformation are used. Subsequently slice timing correction shifts the data (each voxel’s time series) as if whole volume was acquired at exactly the same time. It is accomplished by a simple shift of the phase of the sines that make up signal [9]. Then there is spatial normalization for transformation of scans to standard space defined by template images (for instance Talairach standard space). This procedure employees the affine transformation and nonlinear deformation. It is very important step for the group statistics. Finally there is implemented smoothing procedure for noise suppression. The Gaussian filter is usually used. 2.2 1st level analysis After the preprocessing steps the statistical analysis is carried on. Brain activity is mapped by detection of voxels activated by stimulation. This procedure is called the first level analysis. It can be carried out by correlation analysis or by predetermined haemodynamic response. This

Welcome message from author
This document is posted to help you gain knowledge. Please leave a comment to let me know what you think about it! Share it to your friends and learn new things together.
Transcript

SPM MATLAB TOOLBOX
J. Tauchmanova, M. Hromcık, R. Jech
Czech Technical University, Faculty of Electrical Engineering, Department of ControlEngineering, Department of Neurology, First Faculty of Medicine, Prague
Abstract
The SPM toolbox is a noncommercial software for processing fMRI data. Thetoolbox is developed at the Department of Imaging Neuroscience, UniversityCollege London. The common application of the SPM toolbox is the first levelanalysis consisting in the detection of active brain regions for one patient. Thesecond level analysis, also covered by the SPM toolbox, is related to processing offMRI data across the whole patients group. Besides these functions, the toolboxoffers tools for DCM analysis, such as the parameters estimation routines, toolsfor comparison of resulting models, and a function for averaging models acrossthe whole patients group.
1 Introduction
SPM toolbox serves for data processing, especially for fMRI, EEG and MEG data sets. It in-cludes preprocessing steps, statistical analysis and DCM analysis. There exist several versions ofthis toolbox. The oldest available version was called SPM99, then SPM2 followed and nowadaysversion SPM5 is used.fMRI abbreviates functional Magnetic Resonance Imaging. This method is able to create map ofbrain activity. The results of fMRI experiment are functional images and structural anatomicalimages which are useful as a base for localization of activated regions. The functional images areput through the statistical analysis and the result is statistically significant regions where theso-called haemodynamic responses were detected. But there appear some complications duringthis procedure for example some statistical discrepancies. The SPM toolbox try to correct themand to produce regular results.
2 fMRI analysis
2.1 Preprocessing
The first stage of fMRI data analysis is data preprocessing - preparation of data for statisticalanalysis. SPM toolbox offers these adjustments. Realigning corrects movement artifacts indata. Least squares approach and spatial transformation are used. Subsequently slice timingcorrection shifts the data (each voxel’s time series) as if whole volume was acquired at exactlythe same time. It is accomplished by a simple shift of the phase of the sines that make up signal[9]. Then there is spatial normalization for transformation of scans to standard space defined bytemplate images (for instance Talairach standard space). This procedure employees the affinetransformation and nonlinear deformation. It is very important step for the group statistics.Finally there is implemented smoothing procedure for noise suppression. The Gaussian filter isusually used.
2.2 1st level analysis
After the preprocessing steps the statistical analysis is carried on. Brain activity is mapped bydetection of voxels activated by stimulation. This procedure is called the first level analysis. Itcan be carried out by correlation analysis or by predetermined haemodynamic response. This

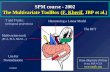
statistical analysis is based on GLM - General Linear Model. It means to define so-called designmatrix which embodies the fMRI experiment. Then the estimation of GLM parameters followsand finally the statistical parametric map is produced. It indicates the likeliest voxels associatedwith haemodynamic responses. The Fig.1 presents the result of the first level analysis. In thiscase the stimulus was left hand motion. The activated voxels are colorfully marked.
Figure 1: The statistical map - the first level analysis
2.3 2nd level analysis
The next type of analysis allows generalization of some conclusions to the whole group of patients[5]. There exist statistical methods for combining results across the subjects. The result isstatistical map as well and it is valid for all patients from investigated group. These methodsinclude ”fixed-effects” and ”random-effects” procedure. Fixed-effects allow inferences to bemade about the particular subject in the experiment, while random-effects allows inferencesto be made about the population. Random effects analysis is considered more appropriate forfMRI research because it deals with making inferences on the population [6]. The second levelanalysis can be useful for instance for selection of brain areas during DCM analysis.

Figure 2: The statistical map - the second level analysis
3 DCM analysis
The next type of fMRI processing is DCM analysis. It is a part of SPM toolbox as well.Dynamic Casual Modelling (DCM) is a statistical technique for detection of connections amongselected brain areas [3]. The DCM procedure treats the brain as a deterministic nonlineardynamic system. This system has some inputs and produces some outputs, measured by fMRIas the BOLD (Blood Oxygen Level Dependent) signals. The inputs to the system are thesignals defining a certain fMRI experiment, i.e. time series representing some stimuli such asfinger movement commands, projection of emotional pictures to the patients etc. Furthermore amodel structure must be predefined before the DCM analysis is applied seeing Fig.3. Thereforecertain special knowledge in brain organization is necessary. In addition, there are typicallyseveral structure candidates that must be processed and evaluated separately which can becometime consuming.

Figure 3: The predefined models
The DCM models are estimated using Bayesian estimators [8]. The inferences aboutconnections are made using the posterior or conditional density [3]. The DCM result is thelikeliest model accompanied by strength values of significant connections Fig.4. This figure wascreated by means of SPM toolbox [1].
3.0.1 DCM drawbacks
DCM is undoubtedly an established and commonly used method for identification of functionalbrain organization from fMRI data. DCM processing however can have some drawbacks fromthe user’s point of view. They are explained in detail in the paper [2] and are based on experienceof the authors with processing one particular fMRI experiment by DCM.The predefinition of models is a complication not only for a user who does not have deepexperience with fMRI, even educated experts have sometimes problems to establish the mostappropriate model structure. As a result, a tedious trial-and-error loop must be performed toarrive at acceptable findings.The DCM analysis of one model itself takes some time and the computation for a complete setof models can take a few days easily. Sometimes there is a specific additional uncertainty suchas an alternative selection of a brain area coordinates which leads to additional structures tobe considered. In the case of analysis of a therapy influence all these demands are doubled inprinciple (by data received after the therapy and their processing). These issues are discussedin detail in [2]. These troubles could be considerably reduced if an alternative identificationprocedure were used in place of DCM that would automatically detect the internal structure ofthe most appropriate dynamical model.Methods for fitting measured data sets that do not require any particular assumptions on thesystem structure (apart of the order and linearity) are however commonly used in systemsand signals identification routines, to estimate communication channels behavior, mechanicalstructures models, chemical processes etc. [4]. Our aim is therefore to try applicability of thosemethods in the fMRI case.We expect some problems here related to small amount of data samples; the fMRI investigationdoes not allow fast sampling rates and the patients can be put subject to the experiment for alimited time period. The DCM features we present above as drawbacks certainly help on the

other hand to eliminate the lack of data (by incorporation of some a-priori information in fact).
LS1a
RSM1
LSM1
LS2a
SMAa
0.90 0.21
1.00 0.39
Intrinsic connectionsP(|connection| > 0.00)connection strength
LS1aRSM1
LSM1LS2a
SMAaLS1aRSM1LSM1LS2aSMAa
−1
−0.5
0
0.5
from
A − Intrinsic connections
to LS1aRSM1
LSM1LS2a
SMAaLS1aRSM1LSM1LS2aSMAa
0
0.2
0.4
0.6
0.8
1
from
A {probabilities}
to
P(A > 0.00)
Figure 4: The significant connections
4 SPM simulator
The important part of SPM toolbox is data simulator which generates the fMRI DCM data onbasis of our requirements. The simulator is started by means of function named spm dcm create.The function is able to generates simulated signals of selected brain regions with required param-eters such as the signal-to-noise ratio, the number of regions, interscan interval (TR, samplingperiod in principle), number of scans (samples), number of conditions (stimulating inputs) andthe vectors of onsets (starting instants) and durations for input signals (stimulations). Thechoice of the connectivity matrix A, the input matrix C and the modulatory matrix B follows.
5 Conclusion
The SPM toolbox is very useful software for fMRI data processing. It is suitable for doctors,for instance the version SPM5 permits batch data processing. And statistical analysis is nottime-consuming for certain group of patients. The toolbox has also SPM simulator which isable to generate the typical fMRI data. It is very important for some experiments with fMRIdata because we can define for instance very high quality of data or on the other hand very badquality of data.

6 ACKNOWLEDGMENTS
The work of Jana Tauchmanova has been supported by the Ministry of Education of the CzechRepublic under Research Program No. MSM6840770038 and under the project Center of AppliedCybernetics (1M0567).The work of Martin Hromcık has been supported by the Ministry of Education of the CzechRepublic under the project Center for Applied Cybernetics (1M0567).The work of Robert Jech has been supported by IGA MZ CR 1A/8629-5, NR8937-4 and ResearchProgram MSM0021620849.
References
[1] Web site of University College London http://www.fil.ion.ucl.ac.uk/spm/
[2] J. Tauchmanova, R. Jech, P. Havrankova, M. Hromcik, DCM analysis of fMRI data inpatients with writer’s cramp, submitted to IEEE EMBC 2008, Vancouver.
[3] K.J. Friston, L. Harrison, W. Penny, Dynamic casual modelling, NeuroImage, 2003.
[4] L. Ljung, System Identification - Theory For the User, 2nd ed, PTR Prentice Hall, UpperSaddle River, N.J., 1999.
[5] S. M. Smith, Overview of fMRI analysis, The British Journal of Radiology, 2004.
[6] Human Brain Imaging, http://fmri.pbwiki.com.
[7] K. E. Stephan, L. M. Harrison, W. D. Penny, K. J. Friston Dynamic casual models of brainresponses, Computational Neuroscience Course, 2004.
[8] W. D. Penny, K. E. Stephan, A. Mechelli, K. J. Friston Comparing dynamic casual models,NeuroImage, 2004.
[9] SPM5 manual, http://www.fil.ion.ucl.ac.uk/spm/doc/manual.pdf.
Jana TauchmanovaDepartment of Control Engineering, FEE, CTU, [email protected]
Martin HromcıkDepartment of Control Engineering, FEE, CTU, [email protected]
Robert JechDepartment of Neurology, First Faculty of Medicine and General Faculty Hospital, CharlesUniversity, [email protected]
Related Documents